Latent GP Classification — the Three Patterns
For non-conjugate likelihoods (Bernoulli, Poisson, StudentT, …) the GP latent function cannot be marginalized analytically — we need gp_sample inside the NumPyro model. This notebook walks the three pyrox patterns through a 2D binary-classification problem.
What you’ll learn:
- Use
gp_sampleto register the latent GP function as a sample site under an arbitrary observation likelihood. - Compose it with a
Bernoulli(logits=f)observation model. - Fit kernel hyperparameters with SVI across all three patterns.
Background¶
Joint model¶
For a non-conjugate likelihood , the hierarchical model is
where is the logistic link. The joint factorizes as
Because the likelihood is not Gaussian, the marginal has no closed form — we can’t collapse out the way we did for regression.
Variational inference + the whitening trick¶
We approximate by a factorized variational distribution and maximize the evidence lower bound
A naive parameterization places directly over the latent values at the training inputs. Under NumPyro’s AutoNormal this becomes a mean-field Gaussian — independent across the training latents. That is a terrible approximation here, because the prior has strong off-diagonal correlations and the optimum of the ELBO ends up dominated by that KL, not by the data fit. In practice the posterior over barely moves and the predictive decision boundary is essentially noise.
The standard fix is the whitening reparameterization (Murray & Adams, 2010; Hensman et al., 2015). Cholesky-factor the prior covariance once,
so the latent enters the likelihood through a deterministic transformation of an i.i.d. unit-Gaussian site. Now mean-field is well-conditioned — there are no a-priori correlations between the to begin with — and the GP correlations come back into for free through . Empirically this is the difference between a chaotic boundary and one that tracks the data.
gp_sample modes — when to reach for which¶
pyrox exposes gp_sample("f", prior, *, whitened, guide) as the NumPyro-aware registration for the latent function. The three modes are mutually exclusive:
- default (
whitened=False,guide=None) — registers a single sample site viagaussx.MultivariateNormal. The right call for MCMC (NUTS samples directly in the original -space and handles the prior covariance correctly) and for conjugate workflows where you marginalize out viagp_factor. - whitened (
whitened=True) — registers an -dimensional unit-Gaussian site and returns the deterministic shown above. The right call for SVI on non-conjugate likelihoods with NumPyro auto-guides such asAutoNormal. This is what every pattern in this notebook uses. - guide (
guide=...) — delegates to a concrete sparse variational guide (FullRankGuide,MeanFieldGuide, orWhitenedGuideover inducing values), which is the right call for the sparse SVGP workflow when is too large for an -dimensional latent.
Posterior predictive¶
For a test input the posterior predictive is
which we estimate by Monte Carlo + the logistic-Gaussian approximation further below.
The three patterns¶
| Pattern | Kernel hyperparameters live in | When to reach for it |
|---|---|---|
| A | Pure eqx.Module + numpyro.sample + eqx.tree_at | Lightweight; no base class required. |
| B | Custom PyroxModule kernel that calls pyrox_sample in __call__ | Self-contained probabilistic kernel. |
| C | Parameterized kernel (shipped RBF, Matern, …) with set_prior | Full registry, constraints, autoguides, modes. |
Only the kernel construction differs across patterns — the whitening boilerplate and the Bernoulli likelihood are shared via a small helper.
Setup¶
import subprocess
import sys
try:
import google.colab # noqa: F401
IN_COLAB = True
except ImportError:
IN_COLAB = False
if IN_COLAB:
subprocess.run(
[
sys.executable,
"-m",
"pip",
"install",
"-q",
"pyrox[colab] @ git+https://github.com/jejjohnson/pyrox@main",
],
check=True,
)import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import equinox as eqx
import jax
import jax.numpy as jnp
import jax.random as jr
import matplotlib.pyplot as plt
import numpyro
import numpyro.distributions as dist
from jaxtyping import Array, Float
from numpyro.infer import SVI, Trace_ELBO
from numpyro.infer.autoguide import AutoNormal
from numpyro.optim import Adam
from pyrox._core import PyroxModule, pyrox_method
from pyrox.gp import RBF, GPPrior, Kernel, gp_sample
from pyrox.gp._src.kernels import rbf_kernel
jax.config.update("jax_enable_x64", True)Reproducibility readout.
import importlib.util
try:
from IPython import get_ipython
ipython = get_ipython()
except ImportError:
ipython = None
if ipython is not None and importlib.util.find_spec("watermark") is not None:
ipython.run_line_magic("load_ext", "watermark")
ipython.run_line_magic(
"watermark",
"-v -m -p jax,equinox,numpyro,gaussx,pyrox,matplotlib",
)
else:
print("watermark extension not installed; skipping reproducibility readout.")Python implementation: CPython
Python version : 3.13.5
IPython version : 9.10.0
jax : 0.9.2
equinox : 0.13.6
numpyro : 0.20.1
gaussx : 0.0.10
pyrox : 0.0.2
matplotlib: 3.10.8
Compiler : GCC 11.2.0
OS : Linux
Release : 6.8.0-1044-azure
Machine : x86_64
Processor : x86_64
CPU cores : 16
Architecture: 64bit
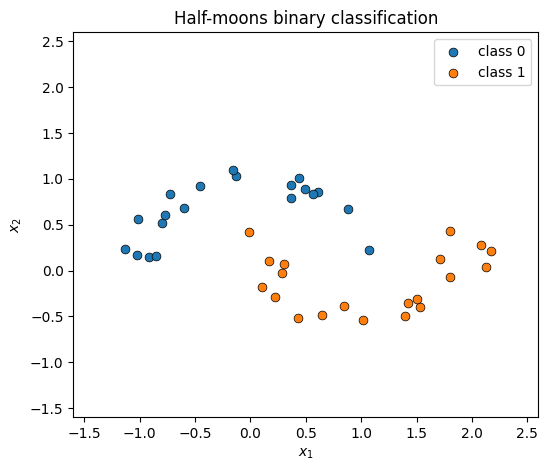
Toy 2D binary dataset¶
Two interleaved half-moons — the boundary between them is the function the latent GP has to learn.
key = jr.PRNGKey(0)
def make_half_moons(key, n_per_class=30):
k1, k2, k3, k4 = jr.split(key, 4)
theta0 = jnp.pi * jr.uniform(k1, (n_per_class,))
x0 = jnp.stack([jnp.cos(theta0), jnp.sin(theta0)], axis=-1)
x0 = x0 + 0.1 * jr.normal(k2, x0.shape)
theta1 = jnp.pi + jnp.pi * jr.uniform(k3, (n_per_class,))
x1 = jnp.stack([1.0 + jnp.cos(theta1), 0.5 + jnp.sin(theta1)], axis=-1)
x1 = x1 + 0.1 * jr.normal(k4, x1.shape)
X = jnp.concatenate([x0, x1], axis=0)
y = jnp.concatenate([jnp.zeros(n_per_class), jnp.ones(n_per_class)], axis=0).astype(
jnp.int32
)
return X, y
X_train, y_train = make_half_moons(key, n_per_class=20)
grid_lo, grid_hi = -1.6, 2.6
grid_steps = 40
xx, yy = jnp.meshgrid(
jnp.linspace(grid_lo, grid_hi, grid_steps),
jnp.linspace(grid_lo, grid_hi, grid_steps),
)
X_grid = jnp.stack([xx.ravel(), yy.ravel()], axis=-1)
print(f"Training points: {X_train.shape[0]}")
print(f"Grid points: {X_grid.shape[0]}")
def scatter_data(ax):
mask0 = y_train == 0
ax.scatter(
X_train[mask0, 0],
X_train[mask0, 1],
s=40,
c="C0",
edgecolors="k",
linewidths=0.5,
label="class 0",
zorder=5,
)
ax.scatter(
X_train[~mask0, 0],
X_train[~mask0, 1],
s=40,
c="C1",
edgecolors="k",
linewidths=0.5,
label="class 1",
zorder=5,
)
ax.set_xlim(grid_lo, grid_hi)
ax.set_ylim(grid_lo, grid_hi)
ax.set_xlabel(r"$x_1$")
ax.set_ylabel(r"$x_2$")
fig, ax = plt.subplots(figsize=(6, 5))
scatter_data(ax)
ax.set_title("Half-moons binary classification")
ax.legend()
plt.show()Training points: 40
Grid points: 1600
The common model skeleton¶
All three patterns share the same whitened latent + Bernoulli likelihood — the only thing that changes is how kernel is built (which is the whole point of comparing the patterns):
prior = GPPrior(kernel, X, jitter=1e-4) # kernel built per-pattern
f = gp_sample("f", prior, whitened=True)
# samples u ~ N(0, I_N) and returns the deterministic
# f = mu(X) + L u with L = chol(K + jitter*I)
obs ~ Bernoulli(logits=f)gp_sample(..., whitened=True) is the model-layer entry point that registers the unit-Gaussian site as "f_u" and the unwhitened function value as the deterministic "f". Each pattern’s model function then reads as “build the kernel, then run the standard whitened-classification pipeline.”
Pattern A — pure Equinox + eqx.tree_at¶
class RBFLite(Kernel):
"""Minimal Equinox-native RBF kernel."""
variance: Float[Array, ""]
lengthscale: Float[Array, ""]
def __call__(
self, X1: Float[Array, "N1 D"], X2: Float[Array, "N2 D"]
) -> Float[Array, "N1 N2"]:
return rbf_kernel(X1, X2, self.variance, self.lengthscale)
def model_pattern_a(X, y):
variance = numpyro.sample("variance", dist.LogNormal(0.0, 1.0))
lengthscale = numpyro.sample("lengthscale", dist.LogNormal(0.0, 1.0))
kernel = RBFLite(variance=jnp.array(1.0), lengthscale=jnp.array(1.0))
kernel = eqx.tree_at(
lambda k: (k.variance, k.lengthscale),
kernel,
(variance, lengthscale),
)
prior = GPPrior(kernel=kernel, X=X, jitter=1e-4)
f = gp_sample("f", prior, whitened=True)
numpyro.sample("obs", dist.Bernoulli(logits=f), obs=y)Pattern B — PyroxModule kernel with pyrox_sample¶
class RBFPyrox(Kernel, PyroxModule):
"""PyroxModule kernel with inline prior registration."""
pyrox_name: str = "RBFPyrox"
@pyrox_method
def __call__(
self, X1: Float[Array, "N1 D"], X2: Float[Array, "N2 D"]
) -> Float[Array, "N1 N2"]:
variance = self.pyrox_sample("variance", dist.LogNormal(0.0, 1.0))
lengthscale = self.pyrox_sample("lengthscale", dist.LogNormal(0.0, 1.0))
return rbf_kernel(X1, X2, variance, lengthscale)
def model_pattern_b(X, y):
kernel = RBFPyrox()
prior = GPPrior(kernel=kernel, X=X, jitter=1e-4)
f = gp_sample("f", prior, whitened=True)
numpyro.sample("obs", dist.Bernoulli(logits=f), obs=y)Pattern C — Parameterized kernel with set_prior¶
def model_pattern_c(X, y):
kernel = RBF()
kernel.set_prior("variance", dist.LogNormal(0.0, 1.0))
kernel.set_prior("lengthscale", dist.LogNormal(0.0, 1.0))
prior = GPPrior(kernel=kernel, X=X, jitter=1e-4)
f = gp_sample("f", prior, whitened=True)
numpyro.sample("obs", dist.Bernoulli(logits=f), obs=y)Fit all three with the same SVI loop¶
def fit(model_fn, seed, n_steps=400):
guide = AutoNormal(model_fn)
svi = SVI(model_fn, guide, Adam(2e-2), Trace_ELBO())
state = svi.init(jr.PRNGKey(seed), X_train, y_train)
losses = []
for _ in range(n_steps):
state, loss = svi.update(state, X_train, y_train)
losses.append(float(loss))
return state, svi, guide, losses
state_a, svi_a, guide_a, losses_a = fit(model_pattern_a, 1)
state_b, svi_b, guide_b, losses_b = fit(model_pattern_b, 2)
state_c, svi_c, guide_c, losses_c = fit(model_pattern_c, 3)
def posterior_hyperparams(svi, state, variance_site, lengthscale_site):
params = svi.get_params(state)
variance = float(jnp.exp(params[f"{variance_site}_auto_loc"]))
lengthscale = float(jnp.exp(params[f"{lengthscale_site}_auto_loc"]))
return variance, lengthscale
v_a, ls_a = posterior_hyperparams(svi_a, state_a, "variance", "lengthscale")
v_b, ls_b = posterior_hyperparams(
svi_b, state_b, "RBFPyrox.variance", "RBFPyrox.lengthscale"
)
v_c, ls_c = posterior_hyperparams(svi_c, state_c, "RBF.variance", "RBF.lengthscale")
print(f"{'pattern':<10} {'variance':>10} {'lengthscale':>12}")
print("-" * 36)
print(f"{'A':<10} {v_a:>10.3f} {ls_a:>12.3f}")
print(f"{'B':<10} {v_b:>10.3f} {ls_b:>12.3f}")
print(f"{'C':<10} {v_c:>10.3f} {ls_c:>12.3f}")pattern variance lengthscale
------------------------------------
A 4.707 0.704
B 4.070 0.699
C 4.326 0.767
fig, ax = plt.subplots(figsize=(12, 5))
ax.plot(losses_a, "C0-", label="Pattern A", alpha=0.85)
ax.plot(losses_b, "C1-", label="Pattern B", alpha=0.85)
ax.plot(losses_c, "C2-", label="Pattern C", alpha=0.85)
ax.set_xlabel("SVI step")
ax.set_ylabel(r"$-$ELBO")
ax.set_title("SVI convergence — three patterns")
ax.legend()
plt.show()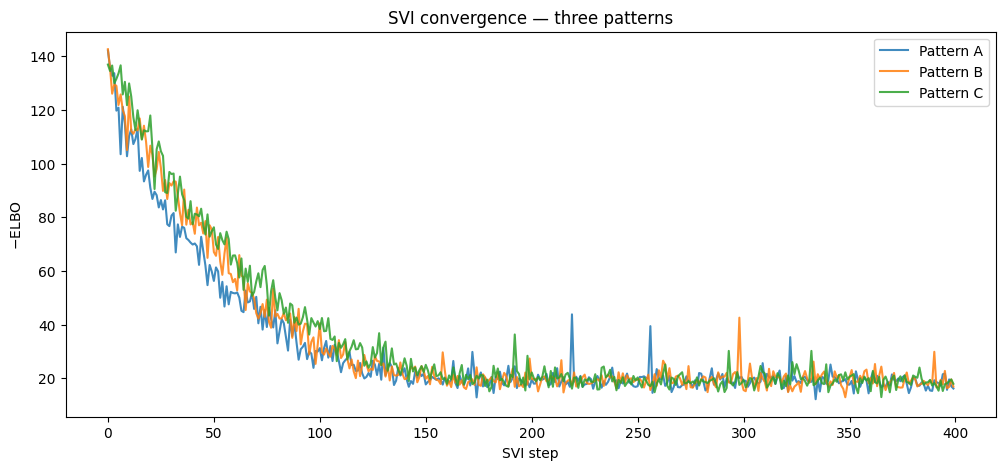
Decision boundaries — MC-averaged posterior predictive¶
We want the Bayesian posterior predictive
where is the SVI guide posterior over the kernel hyperparameters and the whitened latent, and is the standard noise-free GP conditional
with and . ε is the same jitter used inside latent_f.
We estimate the outer integral by Monte Carlo and the inner integral by the logistic-Gaussian approximation
which is exact for the probit link and within a percent for the logistic link that we actually use (MacKay, 1992).
Putting it together — for each MC sample :
- Draw from the SVI guide.
- Reconstruct the training-time latent with the matching Cholesky.
- Build the GP prior at and condition on to get at the grid points.
- Apply the logistic-Gaussian approximation to get per-sample class probabilities.
Average:
This uses the actual fitted Bernoulli model — no Gaussian-likelihood stand-in anywhere.
N_MC = 16
def predict_prob(guide, svi_obj, state, variance_site, lengthscale_site):
"""MC-averaged posterior predictive for the Bernoulli GP.
Each guide sample yields ``(theta_s, u_s)`` over the kernel
hyperparameters and the whitened latent. We reconstruct the
training-time latent ``f_s = L(theta_s) @ u_s``, condition the GP
on ``f_s`` as noise-free observations, and apply the
logistic-Gaussian approximation to get class probabilities at the
grid points before averaging across samples.
"""
params = svi_obj.get_params(state)
posterior = guide.sample_posterior(jr.PRNGKey(42), params, sample_shape=(N_MC,))
variances = posterior[variance_site]
lengthscales = posterior[lengthscale_site]
u_samples = posterior["f_u"] # (N_MC, N_train)
probs = []
for v, ls, u in zip(variances, lengthscales, u_samples):
kernel = RBF(init_variance=float(v), init_lengthscale=float(ls))
prior = GPPrior(kernel=kernel, X=X_train, jitter=1e-4)
L = jnp.linalg.cholesky(prior._prior_operator().as_matrix())
f_train = L @ u
# Condition on the reconstructed latent as noise-free observations.
cond = prior.condition(f_train, jnp.array(1e-4))
mean, var = cond.predict(X_grid)
kappa = 1.0 / jnp.sqrt(1.0 + jnp.pi * jnp.clip(var, min=0.0) / 8.0)
probs.append(jax.nn.sigmoid(kappa * mean))
return jnp.mean(jnp.stack(probs), axis=0).reshape(grid_steps, grid_steps)
prob_a = predict_prob(guide_a, svi_a, state_a, "variance", "lengthscale")
prob_b = predict_prob(
guide_b, svi_b, state_b, "RBFPyrox.variance", "RBFPyrox.lengthscale"
)
prob_c = predict_prob(guide_c, svi_c, state_c, "RBF.variance", "RBF.lengthscale")
fig, axes = plt.subplots(1, 3, figsize=(18, 5), sharey=True)
for ax, prob, title in zip(
axes, [prob_a, prob_b, prob_c], ["Pattern A", "Pattern B", "Pattern C"]
):
cf = ax.contourf(
xx, yy, prob, levels=10, cmap="RdBu_r", alpha=0.7, vmin=0.0, vmax=1.0
)
ax.contour(xx, yy, prob, levels=[0.5], colors="k", linewidths=1.5)
scatter_data(ax)
ax.set_title(title)
ax.legend(fontsize=9)
fig.colorbar(cf, ax=axes, shrink=0.9, label=r"$p(y=1 \mid x)$")
plt.show()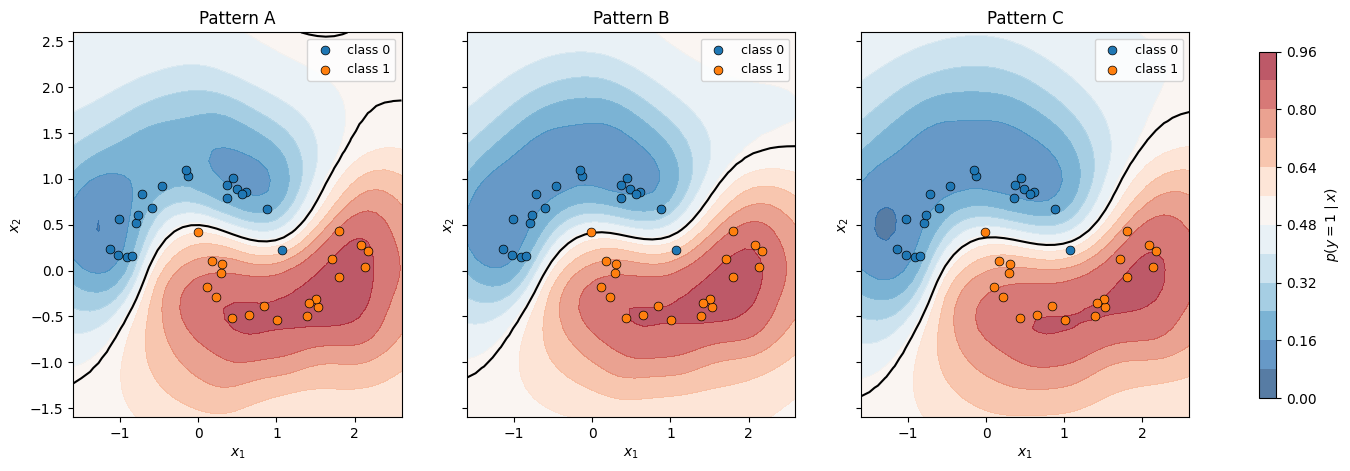
When to use which¶
The non-conjugate story is exactly the conjugate story with gp_factor swapped for gp_sample + an explicit likelihood. The pattern choice is still purely ergonomic:
- Pattern A — lightest; sample hyperparameters at the model level and splice into a pure Equinox kernel. Good for one-off Bayesian extensions.
- Pattern B — custom
PyroxModulekernel that owns its priors. Good when the kernel is a reusable probabilistic building block. - Pattern C — full registry via
Parameterized. Use the shippedRBF,Matern, etc., and attach priors / autoguides / modes declaratively.
Switch the likelihood, change nothing else — Bernoulli(logits=f) becomes Poisson(rate=jnp.exp(f)) for counts, StudentT(...) for robust regression, and so on.