Kernel Ridge Regression with gaussx
A minimal kernel regression example showing how gaussx handles the core linear algebra: solving and computing the log-marginal likelihood.
This is the computational backbone of Gaussian process regression.
Background¶
Gaussian process (GP) regression is a Bayesian nonparametric method for function estimation. The key idea is to place a GP prior over the space of functions, observe noisy evaluations of the function at a finite set of inputs, and then condition on the data to obtain a posterior process. Because the GP prior is conjugate with Gaussian observation noise, the posterior is again a GP and can be computed in closed form.
Given training inputs with observations and test inputs , the posterior predictive distribution is
where is the kernel matrix evaluated at training points, is the cross-covariance between test and training points, is the kernel at test points, and is the observation noise variance.
The computational bottleneck is solving the kernel system and computing the log-determinant , both of which appear in the log-marginal likelihood. For training points, naive dense approaches cost ; structured kernels and iterative solvers can reduce this substantially. See Rasmussen & Williams (2006), Ch. 2 for a thorough treatment.
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import jax
import jax.numpy as jnp
import lineax as lx
import matplotlib.pyplot as plt
import gaussx
jax.config.update("jax_enable_x64", True)Generate data¶
A simple 1D regression problem: noisy samples from a smooth function.
key = jax.random.PRNGKey(42)
n_train = 50
n_test = 200
noise_std = 0.2
# True function
f_true = lambda x: jnp.sin(3 * x) * jnp.exp(-0.5 * x**2)
# Training data
x_train = jax.random.uniform(key, (n_train,), minval=-3.0, maxval=3.0)
x_train = jnp.sort(x_train)
y_train = f_true(x_train) + noise_std * jax.random.normal(
jax.random.PRNGKey(1), (n_train,)
)
# Test grid
x_test = jnp.linspace(-3.5, 3.5, n_test)fig, ax = plt.subplots(figsize=(10, 4))
ax.plot(x_test, f_true(x_test), "k--", lw=1.5, zorder=4, label="True function")
ax.scatter(
x_train,
y_train,
s=30,
c="C0",
edgecolors="k",
linewidths=0.5,
zorder=5,
label="Training data",
)
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.legend(fontsize=9)
ax.set_title("Kernel Regression: Data")
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.tight_layout()
plt.show()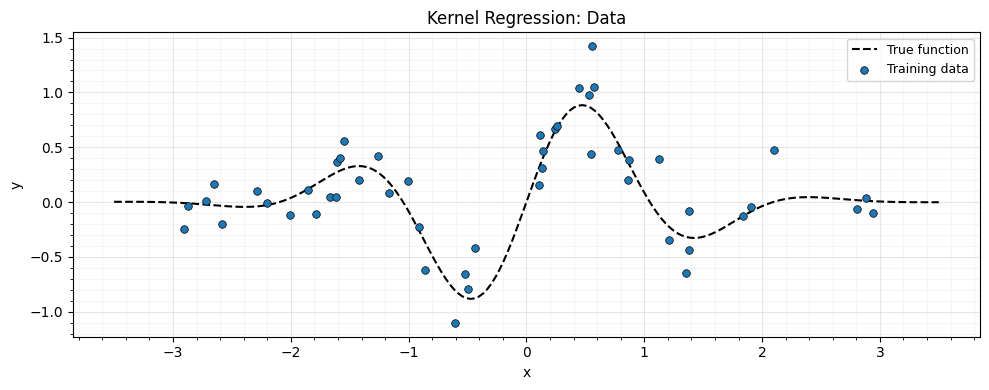
def rbf_kernel(x1, x2, lengthscale, variance):
"""Squared-exponential kernel matrix."""
sq_dist = (x1[:, None] - x2[None, :]) ** 2
return variance * jnp.exp(-0.5 * sq_dist / lengthscale**2)
# Hyperparameters
lengthscale = 0.8
variance = 1.0
noise_var = noise_std**2
# Kernel matrices
K_train = rbf_kernel(x_train, x_train, lengthscale, variance)
K_test_train = rbf_kernel(x_test, x_train, lengthscale, variance)Solve with gaussx¶
The key computation: .
We wrap the kernel matrix as a lineax operator and use
gaussx.solve which dispatches to the appropriate solver.
# K + noise * I as a LowRankUpdate? No -- this is dense.
# Wrap as a MatrixLinearOperator with PSD tag.
K_noisy = K_train + noise_var * jnp.eye(n_train)
op = lx.MatrixLinearOperator(K_noisy, lx.positive_semidefinite_tag)
# Solve for weights
alpha = gaussx.solve(op, y_train)
# Predict
y_pred = K_test_train @ alpha
print("alpha shape:", alpha.shape)
print("prediction shape:", y_pred.shape)alpha shape: (50,)
prediction shape: (200,)
def log_marginal_likelihood(y, op):
"""GP log-marginal likelihood."""
alpha = gaussx.solve(op, y)
data_fit = -0.5 * jnp.dot(y, alpha)
complexity = -0.5 * gaussx.logdet(op)
constant = -0.5 * y.shape[0] * jnp.log(2 * jnp.pi)
return data_fit + complexity + constant
lml = log_marginal_likelihood(y_train, op)
print(f"Log-marginal likelihood: {lml:.4f}")Log-marginal likelihood: -18.2919
Optimizing hyperparameters with grad¶
gaussx primitives are differentiable. We can optimize hyperparameters via gradient descent on the negative log-marginal likelihood.
The analytic gradient of the log-marginal likelihood with respect to a hyperparameter is
where and . The first term rewards data fit and the second penalizes model complexity (Rasmussen & Williams, 2006, Eq. 5.9).
Rather than implementing this expression manually, gaussx + JAX
computes the gradient via automatic differentiation through solve
and logdet. This is both simpler and less error-prone.
def neg_lml(log_params):
"""Negative log-marginal likelihood as a function of log-hyperparameters."""
ls = jnp.exp(log_params[0])
var = jnp.exp(log_params[1])
noise = jnp.exp(log_params[2])
K = rbf_kernel(x_train, x_train, ls, var) + noise * jnp.eye(n_train)
op = lx.MatrixLinearOperator(K, lx.positive_semidefinite_tag)
return -log_marginal_likelihood(y_train, op)
# Initial log-hyperparameters
log_params = jnp.log(jnp.array([lengthscale, variance, noise_var]))
# Gradient
grad_fn = jax.grad(neg_lml)
g = grad_fn(log_params)
print("Gradient of neg-LML:", g)
# Simple gradient descent (a few steps)
lr = 0.1
for _i in range(20):
g = grad_fn(log_params)
log_params = log_params - lr * g
optimized = jnp.exp(log_params)
print(
f"\nOptimized: lengthscale={optimized[0]:.3f}, "
f"variance={optimized[1]:.3f}, noise={optimized[2]:.4f}"
)Gradient of neg-LML: [36.22926626 -2.36412712 -6.9243587 ]
Optimized: lengthscale=0.388, variance=0.177, noise=0.0522
Prediction with optimized hyperparameters¶
ls_opt, var_opt, noise_opt = jnp.exp(log_params)
K_opt = rbf_kernel(x_train, x_train, ls_opt, var_opt) + noise_opt * jnp.eye(n_train)
op_opt = lx.MatrixLinearOperator(K_opt, lx.positive_semidefinite_tag)
alpha_opt = gaussx.solve(op_opt, y_train)
K_star = rbf_kernel(x_test, x_train, ls_opt, var_opt)
y_pred_opt = K_star @ alpha_opt
# Predictive variance
# solve is vector-valued, so use vmap to solve for each test point
K_ss_diag = var_opt * jnp.ones(n_test) # diagonal of K_**
V = jax.vmap(lambda k: gaussx.solve(op_opt, k))(K_star) # (n_test, n_train)
y_var = K_ss_diag - jnp.sum(K_star * V, axis=1)
y_std = jnp.sqrt(jnp.maximum(y_var, 0.0))
print(f"Mean prediction range: [{y_pred_opt.min():.2f}, {y_pred_opt.max():.2f}]")
print(f"Mean std range: [{y_std.min():.4f}, {y_std.max():.4f}]")Mean prediction range: [-0.72, 0.90]
Mean std range: [0.0855, 0.3984]
fig, ax = plt.subplots(figsize=(10, 4))
ax.plot(x_test, f_true(x_test), "k--", lw=1.5, zorder=4, label="True function")
ax.scatter(
x_train,
y_train,
s=30,
c="C0",
edgecolors="k",
linewidths=0.5,
zorder=5,
label="Training data",
)
ax.plot(x_test, y_pred_opt, "C1-", lw=2, zorder=3, label="GP mean")
ax.fill_between(
x_test,
y_pred_opt - 2 * y_std,
y_pred_opt + 2 * y_std,
alpha=0.2,
color="C1",
label="95% CI",
)
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.legend(fontsize=9)
ax.set_title("GP Regression (optimized hyperparameters)")
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.tight_layout()
plt.show()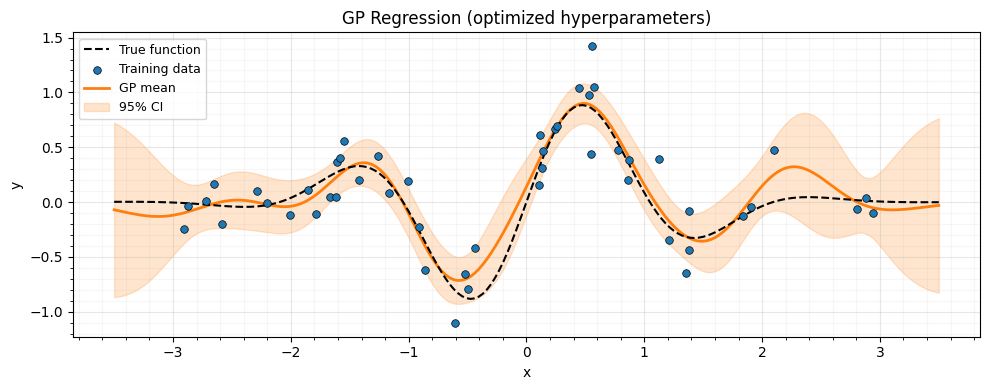
Summary¶
This example showed how gaussx provides the core linear algebra for GP regression:
gaussx.solve— solve the kernel system for weightsgaussx.logdet— log-determinant for the marginal likelihoodjax.grad— differentiate through both to optimize hyperparameters
For large-scale problems, the same code works with structured kernels (Kronecker for grid data, low-rank for inducing points) — gaussx automatically dispatches to the efficient algorithm.
References¶
- Rasmussen, C. E. & Williams, C. K. I. (2006). Gaussian Processes for Machine Learning. MIT Press.
- Williams, C. K. I. & Rasmussen, C. E. (1996). Gaussian processes for regression. Proc. NeurIPS 8.