Multi-Output Gaussian Processes — LMC, ICM, OILMM
Setup¶
A vector-valued GP assigns to every finite collection of inputs a joint Gaussian over . Stacking the columns by — observations of output 1 first, then output 2, etc. — the prior is
and the entire question of “what is a multi-output GP” reduces to: what structure does have? A plain Gram matrix is too expensive — to factor — and is unstructured: it doesn’t say how outputs are coupled. The three constructions here put successively more structure on :
| Kernel | Structure imposed on | Solve cost (Cholesky) | When to use |
|---|---|---|---|
LMCKernel | with rank-1 | generic; when latents tied | Outputs share some structure but each latent has its own kernel / lengthscale. |
ICMKernel | — a single Kronecker product | via | Outputs are smooth versions of one underlying field. |
OILMMKernel | with | via projection to independent scalar GPs | Many outputs, few latents (). |
pyrox.gp exposes all three as equinox.Module kernels whose cross_covariance_operator returns a lineax operator carrying the corresponding structure tag (is_kronecker, is_block_diagonal). Downstream gaussx solvers dispatch on those tags so the dense matrix is never materialised internally.
This notebook walks through:
- LMC on two coupled outputs — share information across outputs to predict where one is missing.
- ICM as a single-Kronecker special case — show the structure tag pyrox emits so solvers can pick the fast path.
- OILMM project / back-project workflow — many outputs, few latents, independent scalar GPs.
Setup¶
import subprocess
import sys
try:
import google.colab # noqa: F401
IN_COLAB = True
except ImportError:
IN_COLAB = False
if IN_COLAB:
subprocess.run(
[
sys.executable,
"-m",
"pip",
"install",
"-q",
"pyrox[colab] @ git+https://github.com/jejjohnson/pyrox@main",
],
check=True,
)import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import jax
import jax.numpy as jnp
import jax.random as jr
import matplotlib.pyplot as plt
import numpy as np
from gaussx import is_kronecker
from pyrox.gp import RBF, ICMKernel, LMCKernel, OILMMKernel
jax.config.update("jax_enable_x64", True)import importlib.util
try:
from IPython import get_ipython
ipython = get_ipython()
except ImportError:
ipython = None
if ipython is not None and importlib.util.find_spec("watermark") is not None:
ipython.run_line_magic("load_ext", "watermark")
ipython.run_line_magic(
"watermark",
"-v -m -p jax,equinox,gaussx,pyrox,lineax,matplotlib",
)
else:
print("watermark extension not installed; skipping reproducibility readout.")Python implementation: CPython
Python version : 3.13.5
IPython version : 9.10.0
jax : 0.9.2
equinox : 0.13.6
gaussx : 0.0.10
pyrox : 0.0.6
lineax : 0.1.0
matplotlib: 3.10.8
Compiler : GCC 11.2.0
OS : Linux
Release : 6.8.0-1044-azure
Machine : x86_64
Processor : x86_64
CPU cores : 16
Architecture: 64bit
1. LMC — borrowing strength across outputs¶
Definition¶
The Linear Model of Coregionalization assumes independent latent scalar GPs and obtains every output as a linear combination:
with a free mixing matrix. The cross-output covariance is, by independence of the ,
where is the rank-1 outer product of the -th column of with itself. Stacking observations as — output by output — the joint Gram matrix is
The Kronecker structure tells you that LMC adds no new freedom within each latent: is rank-1 by construction. Only the number of latents controls expressiveness. When different lengthscales are needed (e.g. one fast latent + one slow latent), LMC captures it; when one shared kernel suffices, ICM is the cheaper restriction.
Cross-output reconstruction¶
We hide a chunk of and reconstruct it from via the LMC posterior. Splitting the joint vector into observed indices and unobserved , the standard Gaussian conditional gives
An independent per-output GP would have block-diagonal , so would be zero between ’s gap entries and ’s observed entries — the conditional mean would collapse to 0 in the gap. LMC’s has off-block entries via , and that’s exactly the route by which informs .
n = 60
key = jr.PRNGKey(0)
key_g1, key_g2, key_eps1, key_eps2 = jr.split(key, 4)
x = jnp.linspace(-3.0, 3.0, n).reshape(-1, 1)
# Two latent GPs with *different* lengthscales — one short, one long. The
# LMC model below uses these same lengthscales, so the data is in-class.
true_kernel_short = RBF(init_variance=1.0, init_lengthscale=0.6)
true_kernel_long = RBF(init_variance=0.5, init_lengthscale=2.0)
K_short = true_kernel_short(x, x) + 1e-8 * jnp.eye(n)
K_long = true_kernel_long(x, x) + 1e-8 * jnp.eye(n)
L_short = jnp.linalg.cholesky(K_short)
L_long = jnp.linalg.cholesky(K_long)
g1 = L_short @ jr.normal(key_g1, (n,)) # short-lengthscale latent
g2 = L_long @ jr.normal(key_g2, (n,)) # long-lengthscale latent
W = jnp.array([[1.0, 0.4], [0.3, 1.1]])
noise_std = 0.05
y1_full = W[0, 0] * g1 + W[0, 1] * g2 + noise_std * jr.normal(key_eps1, (n,))
y2_full = W[1, 0] * g1 + W[1, 1] * g2 + noise_std * jr.normal(key_eps2, (n,))
# Hide the centre of y1 — the LMC must reconstruct it from y2 and the LMC structure.
mask_y1 = (x[:, 0] > -1.0) & (x[:, 0] < 1.0)
y1 = y1_full.at[mask_y1].set(jnp.nan)
y2 = y2_fullBuild an LMC kernel with two latent processes and compute the dense Gram for the joint GP. We do the mask handling by stripping the missing entries from vec(Y); the LMC posterior at those held-out indices is what we want.
# Two latent kernels (give them different lengthscales so LMC ≠ ICM).
lmc = LMCKernel(
kernels=(
RBF(pyrox_name="rbf_q0", init_variance=1.0, init_lengthscale=0.6),
RBF(pyrox_name="rbf_q1", init_variance=0.5, init_lengthscale=2.0),
),
mixing=W,
)
K_full = lmc.full_covariance(x) # (2N, 2N) — vec ordering: y1 then y2
print(f"LMC Gram shape: {K_full.shape} (P=2 outputs × N={n} inputs)")
# Build the joint observation vector and apply the missing-entry mask.
# vec(Y) ordering: lmc.full_covariance stacks rows as [output 0 inputs..., output 1 inputs...].
y_vec = jnp.concatenate([y1, y2])
mask_vec = jnp.concatenate([~mask_y1, jnp.ones_like(mask_y1, dtype=bool)])
y_obs = y_vec[mask_vec]
K_oo = K_full[mask_vec][:, mask_vec] # observed-observed
K_uo = K_full[~mask_vec][:, mask_vec] # unobserved-observed (the gap entries of y1)
K_uu = K_full[~mask_vec][:, ~mask_vec]
noise_var = noise_std**2
A = K_oo + noise_var * jnp.eye(K_oo.shape[0])
alpha = jnp.linalg.solve(A, y_obs)
mean_gap = K_uo @ alpha
var_gap = jnp.diag(K_uu - K_uo @ jnp.linalg.solve(A, K_uo.T))LMC Gram shape: (120, 120) (P=2 outputs × N=60 inputs)
fig, axes = plt.subplots(1, 2, figsize=(18, 5))
# y1 — has the gap.
axes[0].plot(x[:, 0], y1_full, "k--", alpha=0.5, label="true y1")
axes[0].scatter(
x[~mask_y1, 0],
y1_full[~mask_y1],
c="C1",
edgecolors="k",
linewidths=0.5,
zorder=5,
label="observed y1",
)
axes[0].scatter(
x[mask_y1, 0],
mean_gap,
c="C0",
edgecolors="k",
linewidths=0.5,
zorder=5,
label="LMC posterior mean (gap)",
)
axes[0].fill_between(
x[mask_y1, 0],
mean_gap - 2 * jnp.sqrt(var_gap),
mean_gap + 2 * jnp.sqrt(var_gap),
color="C0",
alpha=0.2,
label=r"$\pm 2\sigma$",
)
axes[0].axvspan(-1.0, 1.0, color="0.85", alpha=0.4)
axes[0].set_title("y1 — held-out gap reconstructed via LMC + y2")
axes[0].set_xlabel("x")
axes[0].set_ylabel("y1")
axes[0].legend(loc="upper right")
# y2 — fully observed; show LMC sees it.
axes[1].plot(x[:, 0], y2_full, "k--", alpha=0.5, label="true y2")
axes[1].scatter(
x[:, 0], y2, c="C1", edgecolors="k", linewidths=0.5, zorder=5, label="observed y2"
)
axes[1].axvspan(-1.0, 1.0, color="0.85", alpha=0.4, label="(corresponds to y1's gap)")
axes[1].set_title("y2 — fully observed, drives LMC's gap reconstruction")
axes[1].set_xlabel("x")
axes[1].set_ylabel("y2")
axes[1].legend(loc="upper right")
plt.show()
rmse = float(jnp.sqrt(jnp.mean((mean_gap - y1_full[mask_y1]) ** 2)))
print(f"LMC RMSE on held-out y1 gap: {rmse:.4f}")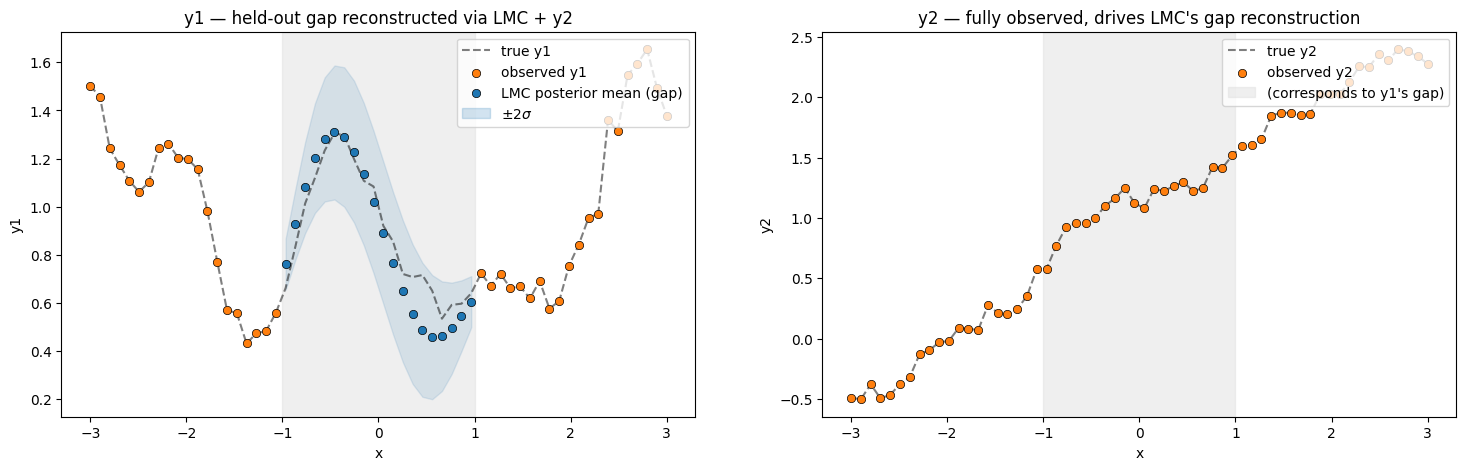
LMC RMSE on held-out y1 gap: 0.0958
The LMC posterior fills the gap in near the truth even though no values were observed in — the cross-output covariance routes information from through the latent processes back into .
2. ICM — the single-Kronecker special case¶
Definition and derivation¶
The Intrinsic Coregionalization Model is the LMC restriction to a single shared latent kernel: for all . The sum-of-Kroneckers collapses by linearity:
Adding per-output diagonal “nugget” terms — extra signal variance unique to each output — gives the standard ICM coregionalization matrix
Why the Kronecker tag matters¶
Cholesky of a matrix is ; with that is already flops. The Kronecker identity
lets gaussx.kronecker_mll evaluate the same log-marginal at — three orders of magnitude cheaper for . The tag is what tells the solver which path to take.
pyrox.gp.ICMKernel.cross_covariance_operator returns a lineax operator with the Kronecker tag set, so any downstream gaussx log-marginal / solve routine can pick the Kronecker-exact path automatically — even though as_matrix() exists for inspection.
icm = ICMKernel(
kernel=RBF(init_variance=1.0, init_lengthscale=0.6),
mixing=jnp.array([[1.0, 0.5], [0.5, 1.2], [0.2, 0.9]]), # P=3 outputs, Q=2 latents
kappa=jnp.array([0.05, 0.05, 0.05]), # per-output extra diagonal variance
)
K_op = icm.cross_covariance_operator(x, x)
print(f"ICM operator: {type(K_op).__name__}")
print(f"is_kronecker tag set: {is_kronecker(K_op)}")
print(f"Operator dense shape: {K_op.as_matrix().shape}")
print(f"Coregionalization B:\n{icm.coregionalization_matrix()}")ICM operator: Kronecker
is_kronecker tag set: True
Operator dense shape: (180, 180)
Coregionalization B:
[[1.3 1.1 0.65]
[1.1 1.74 1.18]
[0.65 1.18 0.9 ]]
is_kronecker(K_op) → True is the load-bearing fact: it tells gaussx that a single Cholesky on the block factor and a single Cholesky on the kernel factor are enough — the dense system is never materialised internally even though as_matrix() exists for inspection.
3. OILMM — project, run scalar GPs, back-project¶
The orthogonality assumption and what it buys¶
OILMM (Bruinsma et al. 2020) is the LMC restricted to a semi-orthogonal mixing matrix:
Geometrically, the columns of are an orthonormal basis of a -dimensional subspace of , and the projector onto that subspace is . Within the subspace, acts isometrically.
Under this constraint, with per-output Gaussian noise , , the likelihood factorises across latents. Multiplying both sides by :
Critically, the projected noise is diagonal in when is orthogonal: there is no off-diagonal coupling between latents after projection (Bruinsma et al. Theorem 1). So the projected problems are fully independent: each is a scalar GP regression on with kernel and noise variance .
The recipe¶
Project observations: .
Fit independent scalar GPs .
Predict in latent space: .
Back-project to output space:
Cost. Steps 2-3 are scalar GP solves at instead of one multi-output solve at . When — many outputs from a few latents — this is a polynomial-degree win. gaussx.oilmm_project and gaussx.oilmm_back_project are the implementation; OILMMKernel.project / back_project are thin wrappers.
P = 4
Q = 2
n_in = 80
key_y, key_w = jr.split(jr.PRNGKey(7), 2)
x_oi = jnp.linspace(-3.0, 3.0, n_in).reshape(-1, 1)
# Generate a random orthogonal mixing matrix via QR.
W_random = jr.normal(key_w, (P, Q))
W_oi, _ = jnp.linalg.qr(W_random) # (P, Q) with W^T W = I_Q
# Two ground-truth latent GPs (well-separated lengthscales).
true_kernels = [
RBF(init_variance=1.0, init_lengthscale=0.5),
RBF(init_variance=0.6, init_lengthscale=2.0),
]
G = jnp.stack(
[
jnp.linalg.cholesky(k(x_oi, x_oi) + 1e-8 * jnp.eye(n_in))
@ jr.normal(jr.fold_in(key_y, q), (n_in,))
for q, k in enumerate(true_kernels)
],
axis=-1,
) # (N, Q)
noise_var_per_output = 0.04 * jnp.ones(P)
Y = G @ W_oi.T + jnp.sqrt(noise_var_per_output) * jr.normal(
jr.fold_in(key_y, 99), (n_in, P)
)Project the -dim observations into the -dim latent space.
oilmm = OILMMKernel(
kernels=(
RBF(pyrox_name="rbf_lat0", init_variance=1.0, init_lengthscale=0.5),
RBF(pyrox_name="rbf_lat1", init_variance=0.6, init_lengthscale=2.0),
),
mixing=W_oi,
check_orthogonal=True,
)
Y_lat, noise_lat = oilmm.project(Y, noise_var_per_output)
print(f"Y shape: {Y.shape}")
print(f"Y_latent shape: {Y_lat.shape} (per-latent observations)")
print(f"per-latent noise variance: {np.asarray(noise_lat)}")Y shape: (80, 4)
Y_latent shape: (80, 2) (per-latent observations)
per-latent noise variance: [0.04 0.04]
Now run two independent scalar GP regressions — one per latent — using gaussx.log_marginal_likelihood for each, and do simple posterior-mean prediction in latent space before back-projecting.
def scalar_gp_predict(
kernel: RBF,
x_train: jax.Array,
y_train: jax.Array,
x_test: jax.Array,
noise_var: float,
):
K_xx = kernel(x_train, x_train) + noise_var * jnp.eye(x_train.shape[0])
K_sx = kernel(x_test, x_train)
K_ss_diag = kernel.diag(x_test)
alpha = jnp.linalg.solve(K_xx, y_train)
mean = K_sx @ alpha
v = jnp.linalg.solve(K_xx, K_sx.T)
var = K_ss_diag - jnp.sum(K_sx * v.T, axis=-1)
return mean, var
x_test = jnp.linspace(-3.5, 3.5, 200).reshape(-1, 1)
f_means = []
f_vars = []
for q, kernel in enumerate(oilmm.independent_gps()):
mean_q, var_q = scalar_gp_predict(
kernel, x_oi, Y_lat[:, q], x_test, float(noise_lat[q])
)
f_means.append(mean_q)
f_vars.append(var_q)
F_means = jnp.stack(f_means, axis=-1) # (M, Q)
F_vars = jnp.stack(f_vars, axis=-1)
# Back-project to the original P=4 output space.
y_means, y_vars = oilmm.back_project(F_means, F_vars)
print(
f"Back-projected predictive mean shape: {y_means.shape} (M={x_test.shape[0]}, P={P})"
)Back-projected predictive mean shape: (200, 4) (M=200, P=4)
fig, axes = plt.subplots(2, 2, figsize=(18, 9))
for p, ax in enumerate(axes.ravel()):
ax.scatter(
x_oi[:, 0],
Y[:, p],
c="C1",
edgecolors="k",
linewidths=0.5,
s=20,
zorder=5,
label="observed",
)
ax.plot(x_test[:, 0], y_means[:, p], color="C0", label="OILMM mean")
ax.fill_between(
x_test[:, 0],
y_means[:, p] - 2 * jnp.sqrt(y_vars[:, p]),
y_means[:, p] + 2 * jnp.sqrt(y_vars[:, p]),
color="C0",
alpha=0.2,
label=r"$\pm 2\sigma$",
)
ax.set_title(f"output {p}")
ax.set_xlabel("x")
ax.set_ylabel(f"y_{p}")
if p == 0:
ax.legend(loc="upper right")
plt.tight_layout()
plt.show()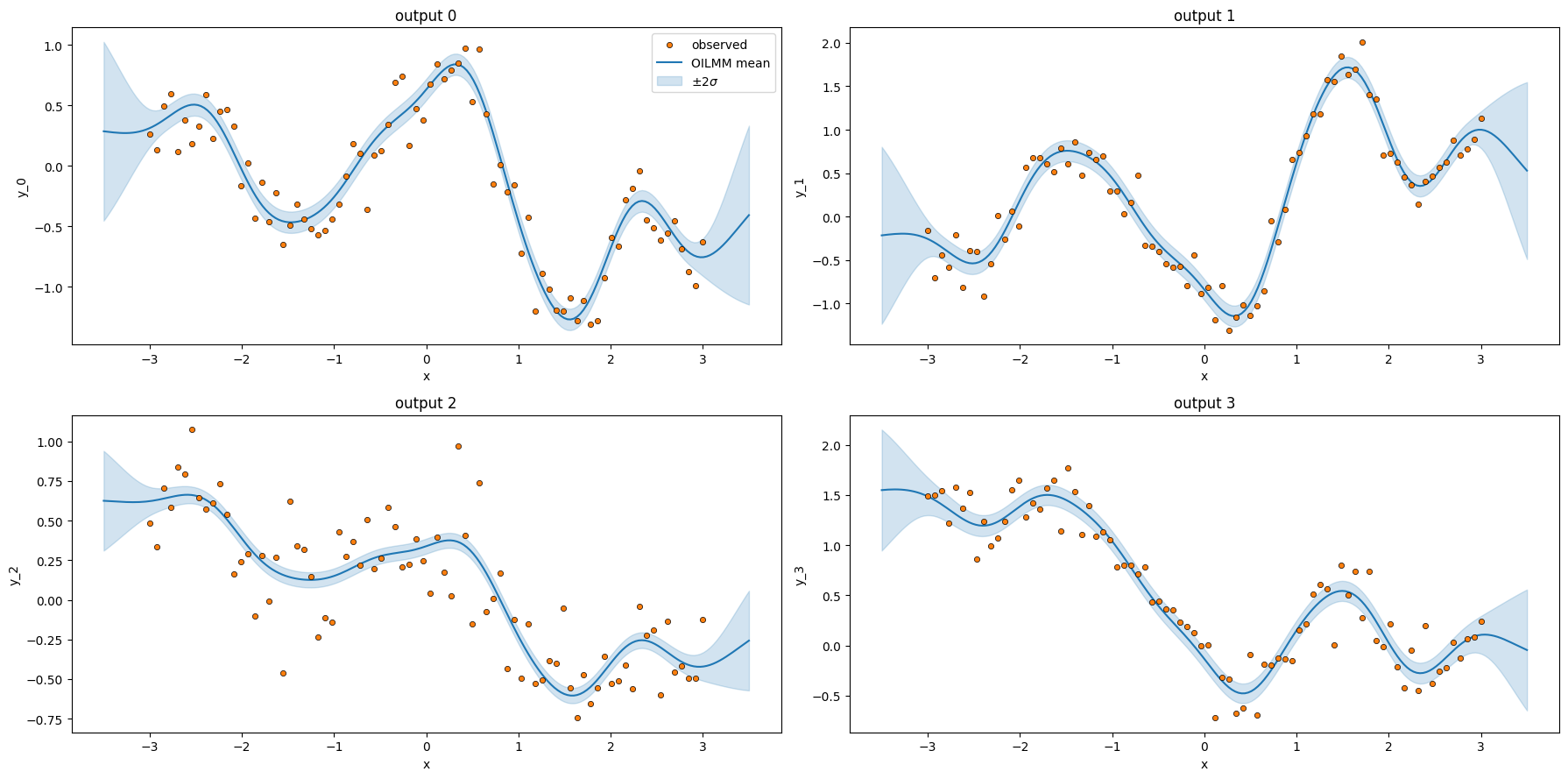
Two scalar GP fits — one per latent — produced predictive means and variances for all four outputs. This is the OILMM win at scale: when is large but the cross-output structure is low-rank (), you pay scalar GP costs instead of one monolithic -output cost.
Sanity check — oilmm_project round-trips¶
oilmm_back_project is the inverse of oilmm_project for the orthogonal mixing case. Verify directly:
F_back, _ = oilmm.back_project(Y_lat, jnp.zeros_like(Y_lat))
F_recon = F_back # back-projected mean
# Project Y_recon → latent and confirm it round-trips.
Y_lat_again, _ = oilmm.project(F_recon, 0.0)
err = float(jnp.max(jnp.abs(Y_lat - Y_lat_again)))
print(f"max round-trip error (project → back-project → project): {err:.2e}")max round-trip error (project → back-project → project): 1.11e-15
Takeaways¶
LMCKernel: with rank-1 — the most expressive of the three because every latent has its own kernel . Cross-output coupling enters via the off-block entries of , which is what lets the conditional posterior fill a gap in from observations of .ICMKernel: the LMC restriction , which by linearity collapses to with . Theis_kroneckertag on the returnedlineaxoperator letsgaussxuse for an exact solve instead of .OILMMKernel: the LMC restriction , which decouples the projected likelihood ( are independent across ) so becomes independent scalar GP problems. Cost drops from to — a polynomial win when . Round-trip:project(Y, σ²) → Q scalar GPs → back_project(μ, var).