LOVE: Fast Leave-One-Out Cross-Validation
Leave-one-out cross-validation (LOO-CV) is a gold standard for GP model selection: it evaluates how well each training point is predicted by the model trained on the remaining points. Unlike the marginal likelihood, LOO-CV directly measures predictive quality and is less sensitive to model misspecification.
The catch is that naive LOO-CV requires separate GP solves, each costing after the initial factorization -- total. The LOVE method (Pleiss et al., 2018) sidesteps this by using a Lanczos decomposition to approximate cheaply, enabling fast predictive variance at arbitrary test points from a single cached factorization.
What you’ll learn:
- How to build a LOVE cache via Lanczos factorization
- How to compute fast predictive variances from the cache
- How to use the cache for LOO-CV scoring
- Kronecker GP: exact MLL and posterior predictions on grids
- Comparing LOO-CV vs MLL for hyperparameter selection
1. Background¶
Given a GP with kernel matrix and noise variance , the noisy kernel matrix is . The predictive variance at a test point is
where is the cross-covariance vector and is the prior variance.
Computing exactly requires an factorization upfront, and each subsequent solve costs . LOVE approximates using a rank- Lanczos decomposition (), so that each variance computation costs only for the matrix-vector product , followed by an diagonal scaling.
For LOO-CV, the key identities are:
where . The diagonal entries can be approximated efficiently using the LOVE cache, since .
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import jax
import jax.numpy as jnp
import lineax as lx
import matplotlib.pyplot as plt
import gaussx
jax.config.update("jax_enable_x64", True)2. Setup: 1D GP regression¶
We generate 100 noisy observations from a smooth function and build the RBF kernel matrix.
key = jax.random.PRNGKey(42)
n_train = 100
n_test = 200
# Training data
key, subkey = jax.random.split(key)
X_train = jnp.sort(jax.random.uniform(subkey, (n_train,), minval=-5.0, maxval=5.0))
f_true = jnp.sin(X_train) + 0.3 * jnp.cos(3.0 * X_train)
key, subkey = jax.random.split(key)
y_train = f_true + 0.2 * jax.random.normal(subkey, (n_train,))
# Test data
X_test = jnp.linspace(-5.0, 5.0, n_test)
# Plot
fig, ax = plt.subplots(figsize=(10, 4))
ax.scatter(
X_train,
y_train,
s=30,
edgecolors="k",
linewidths=0.5,
alpha=0.6,
label="Training data",
zorder=5,
)
ax.plot(
X_test,
jnp.sin(X_test) + 0.3 * jnp.cos(3.0 * X_test),
"k--",
lw=1.5,
label="True function",
zorder=4,
)
ax.set(xlabel="x", ylabel="y")
ax.legend(fontsize=9)
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.show()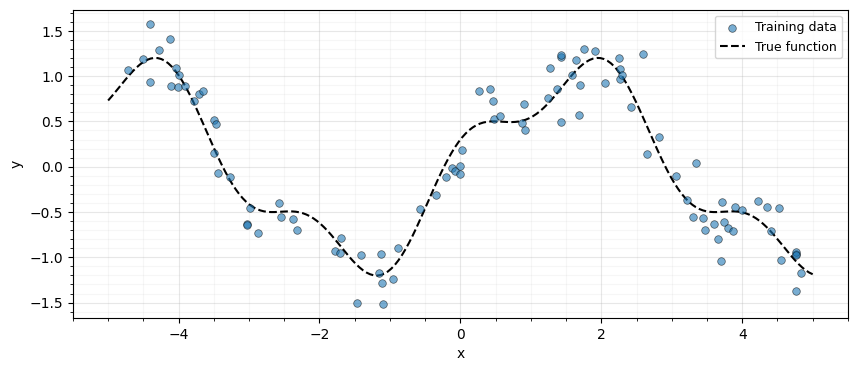
Kernel and noise parameters¶
def rbf_kernel(x1, x2, lengthscale, variance):
"""RBF kernel matrix between two 1D input arrays."""
sq_dist = (x1[:, None] - x2[None, :]) ** 2
return variance * jnp.exp(-0.5 * sq_dist / lengthscale**2)
# Kernel hyperparameters
lengthscale = 1.0
variance = 1.0
noise_var = 0.04 # sigma^2
# Build the noisy kernel matrix K + sigma^2 I
K_train = rbf_kernel(X_train, X_train, lengthscale, variance)
K_noisy = K_train + noise_var * jnp.eye(n_train)
# Wrap as a lineax operator (PSD tag enables symmetric eigensolver)
K_op = lx.MatrixLinearOperator(K_noisy, lx.positive_semidefinite_tag)
print(f"Training points: {n_train}")
print(f"Kernel matrix: {K_noisy.shape}")Training points: 100
Kernel matrix: (100, 100)
3. Build LOVE cache¶
The gaussx.love_cache function runs a Lanczos iteration on the
kernel operator, producing a low-rank approximation of .
The result is a LOVECache containing:
- Q — orthonormal Lanczos basis, shape
(n, m) - inv_eigvals — reciprocal eigenvalues of the tridiagonal matrix, shape
(m,)
cache = gaussx.love_cache(K_op, lanczos_order=50, key=jax.random.PRNGKey(0))
print("LOVECache fields:")
print(f" Q shape: {cache.Q.shape}")
print(f" inv_eigvals shape: {cache.inv_eigvals.shape}")LOVECache fields:
Q shape: (100, 50)
inv_eigvals shape: (50,)
The Lanczos order is half the matrix size here, but in practice suffices. The cache can be reused for any number of test points.
4. Fast predictive variance¶
For each test point , we need the cross-covariance vector and the prior variance . LOVE gives us the quadratic form cheaply.
# Cross-covariance for all test points: shape (n_test, n_train)
K_cross = rbf_kernel(X_test, X_train, lengthscale, variance)
# Prior variance at test points (diagonal of K_**)
K_test_diag = variance * jnp.ones(n_test)
# LOVE variance: vmap over test points
love_quad = jax.vmap(lambda k_row: gaussx.love_variance(cache, k_row))(K_cross)
love_var = K_test_diag - love_quad
print(f"LOVE predictive variance: shape {love_var.shape}")LOVE predictive variance: shape (200,)
Compare against exact variance¶
We compute the exact predictive variance via a dense solve for validation.
# Exact: k_** - k_*^T K^{-1} k_* via gaussx.solve
alpha_cross = jax.vmap(
lambda k_row: gaussx.solve(K_op, k_row),
in_axes=0,
)(K_cross)
exact_quad = jnp.sum(K_cross * alpha_cross, axis=1)
exact_var = K_test_diag - exact_quad
# Compare
max_err = jnp.max(jnp.abs(love_var - exact_var))
rel_err = jnp.max(jnp.abs(love_var - exact_var) / jnp.abs(exact_var + 1e-12))
print(f"Max absolute error: {max_err:.2e}")
print(f"Max relative error: {rel_err:.2e}")Max absolute error: 1.89e-15
Max relative error: 4.88e-13
fig, axes = plt.subplots(1, 2, figsize=(12, 3))
axes[0].plot(X_test, exact_var, label="Exact", linewidth=2, zorder=3)
axes[0].plot(X_test, love_var, "--", label="LOVE (m=50)", linewidth=2, zorder=3)
axes[0].set(
xlabel="x", ylabel="Predictive variance", title="Predictive variance comparison"
)
axes[0].legend(fontsize=9)
axes[0].grid(True, which="major", alpha=0.3)
axes[0].grid(True, which="minor", alpha=0.1)
axes[0].minorticks_on()
axes[1].plot(X_test, jnp.abs(love_var - exact_var), color="tab:red", lw=2)
axes[1].set(xlabel="x", ylabel="Absolute error", title="LOVE approximation error")
axes[1].set_yscale("log")
axes[1].grid(True, which="major", alpha=0.3)
axes[1].grid(True, which="minor", alpha=0.1)
axes[1].minorticks_on()
plt.tight_layout()
plt.show()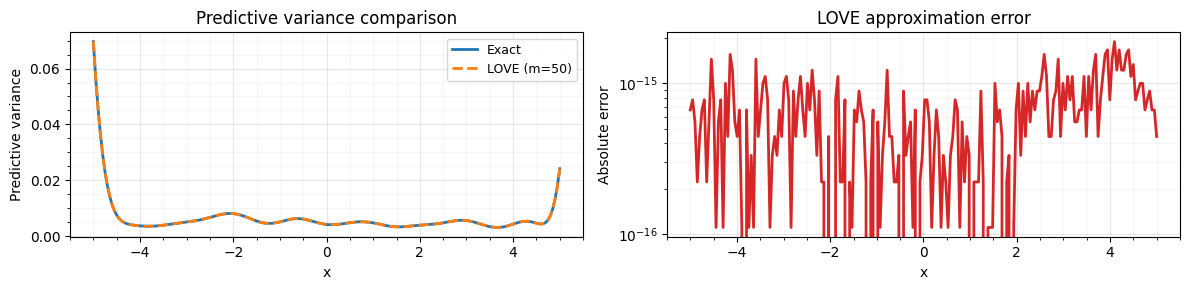
Speedup: vmap over many test points¶
The LOVE cache makes it trivial to compute variance for a batch
of test points via jax.vmap.
love_variance_batch = jax.jit(
jax.vmap(lambda k_row: gaussx.love_variance(cache, k_row))
)
# Warm up JIT
_ = love_variance_batch(K_cross).block_until_ready()
# Time it
import timeit
n_repeats = 100
love_time = (
timeit.timeit(
lambda: love_variance_batch(K_cross).block_until_ready(), number=n_repeats
)
/ n_repeats
)
# Compare with exact solve
exact_fn = jax.jit(jax.vmap(lambda k_row: gaussx.solve(K_op, k_row), in_axes=0))
_ = exact_fn(K_cross).block_until_ready()
exact_time = (
timeit.timeit(lambda: exact_fn(K_cross).block_until_ready(), number=n_repeats)
/ n_repeats
)
print(f"LOVE variance ({n_test} points): {love_time * 1e3:.2f} ms")
print(f"Exact solve ({n_test} points): {exact_time * 1e3:.2f} ms")
print(f"Speedup: {exact_time / love_time:.1f}x")LOVE variance (200 points): 0.09 ms
Exact solve (200 points): 0.59 ms
Speedup: 6.2x
5. LOO-CV via LOVE¶
Leave-one-out cross-validation computes, for each training point , the predictive mean and variance when that point is held out. The classical identities (Rasmussen & Williams, 2006, Eq. 5.12) give:
where . The diagonal of is the expensive part; LOVE approximates it cheaply.
# Compute alpha = K^{-1} y (single solve, not LOO-specific)
alpha = gaussx.solve(K_op, y_train)
# Approximate K^{-1}_{ii} using LOVE cache:
# K^{-1}_{ii} = e_i^T Q diag(inv_eigvals) Q^T e_i
# = sum_j (Q_{ij}^2 * inv_eigvals_j)
K_inv_diag_love = jnp.sum(cache.Q**2 * cache.inv_eigvals[None, :], axis=1)
# Clamp diagonal to positive values to avoid negative variance
K_inv_diag_love = jnp.clip(K_inv_diag_love, a_min=1e-12)
# LOO predictive mean and variance
mu_loo = y_train - alpha / K_inv_diag_love
var_loo = 1.0 / K_inv_diag_love
print(f"LOO predictive mean: shape {mu_loo.shape}")
print(f"LOO predictive variance: shape {var_loo.shape}")LOO predictive mean: shape (100,)
LOO predictive variance: shape (100,)
LOO log-predictive density¶
The LOO log-predictive density (LPD) is a standard model selection criterion:
loo_lpd = -0.5 * n_train * jnp.log(2 * jnp.pi) - 0.5 * jnp.sum(
jnp.log(var_loo) + (y_train - mu_loo) ** 2 / var_loo
)
print(f"LOO log-predictive density: {loo_lpd:.4f}")LOO log-predictive density: -170.9582
# Visualize LOO predictions
fig, ax = plt.subplots(figsize=(10, 4))
ax.scatter(
X_train,
y_train,
s=30,
edgecolors="k",
linewidths=0.5,
alpha=0.4,
label="Training data",
zorder=5,
)
ax.errorbar(
X_train,
mu_loo,
yerr=2 * jnp.sqrt(var_loo),
fmt=".",
markersize=4,
alpha=0.6,
capsize=2,
label=r"LOO mean $\pm$ 2$\sigma$",
zorder=3,
)
ax.set(xlabel="x", ylabel="y", title="Leave-one-out predictions (LOVE)")
ax.legend(fontsize=9)
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.show()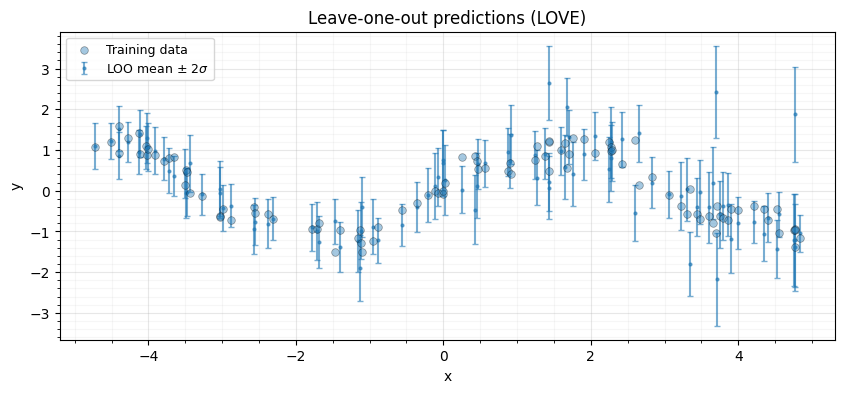
6. Kronecker GP¶
When data lies on a 2D grid, the kernel matrix factorizes as a Kronecker product , reducing to where .
gaussx provides kronecker_mll for the marginal log-likelihood and
kronecker_posterior_predictive for predictions.
2D grid problem¶
# Grid dimensions
n1, n2 = 10, 10
grid_shape = (n1, n2)
N = n1 * n2
# 1D grids
x1 = jnp.linspace(-3.0, 3.0, n1)
x2 = jnp.linspace(-3.0, 3.0, n2)
# True function on grid
X1, X2 = jnp.meshgrid(x1, x2, indexing="ij")
f_grid = jnp.sin(X1) * jnp.cos(X2)
y_grid = f_grid.ravel()
# Add noise
key, subkey = jax.random.split(key)
y_grid_noisy = y_grid + 0.1 * jax.random.normal(subkey, (N,))
# Build per-dimension kernel matrices
ls1, ls2 = 1.0, 1.0
var1, var2 = 1.0, 1.0
noise_var_grid = 0.01
def rbf_kernel_1d(x, lengthscale, var):
"""1D RBF kernel matrix."""
sq_dist = (x[:, None] - x[None, :]) ** 2
return var * jnp.exp(-0.5 * sq_dist / lengthscale**2)
K1 = rbf_kernel_1d(x1, ls1, var1)
K2 = rbf_kernel_1d(x2, ls2, var2)
# Wrap as lineax operators (PSD tag for symmetric eigensolvers)
K_factors = [
lx.MatrixLinearOperator(K1, lx.positive_semidefinite_tag),
lx.MatrixLinearOperator(K2, lx.positive_semidefinite_tag),
]
print(f"Grid: {n1} x {n2} = {N} points")
print(f"K1: {K1.shape}, K2: {K2.shape}")Grid: 10 x 10 = 100 points
K1: (10, 10), K2: (10, 10)
Marginal log-likelihood¶
mll = gaussx.kronecker_mll(K_factors, y_grid_noisy, noise_var_grid, grid_shape)
print(f"Kronecker MLL: {mll:.4f}")Kronecker MLL: -4.0269
Verify against dense GP¶
For this small grid we can also compute the dense MLL directly.
K_full = jnp.kron(K1, K2) + noise_var_grid * jnp.eye(N)
K_full_op = lx.MatrixLinearOperator(K_full, lx.positive_semidefinite_tag)
alpha_full = gaussx.solve(K_full_op, y_grid_noisy)
sign, logdet_full = jnp.linalg.slogdet(K_full)
mll_dense = -0.5 * (y_grid_noisy @ alpha_full + logdet_full + N * jnp.log(2 * jnp.pi))
print(f"Kronecker MLL: {mll:.4f}")
print(f"Dense MLL: {mll_dense:.4f}")
print(f"Difference: {jnp.abs(mll - mll_dense):.2e}")Kronecker MLL: -4.0269
Dense MLL: -4.0269
Difference: 1.71e-13
Posterior predictions¶
We predict at a finer test grid using kronecker_posterior_predictive.
The function accepts per-dimension cross-covariance arrays and
test diagonal arrays.
# Test grid (finer)
n1_test, n2_test = 20, 20
x1_test = jnp.linspace(-3.0, 3.0, n1_test)
x2_test = jnp.linspace(-3.0, 3.0, n2_test)
# Per-dimension cross-covariances: (n_test_d, n_train_d)
K_cross_1 = rbf_kernel_1d(x1_test, ls1, var1)[:, :n1] # (n1_test, n1)
K_cross_2 = rbf_kernel_1d(x2_test, ls2, var2)[:, :n2] # (n2_test, n2)
# Fix: compute proper cross-covariance between test and train grids
K_cross_1 = var1 * jnp.exp(-0.5 * (x1_test[:, None] - x1[None, :]) ** 2 / ls1**2)
K_cross_2 = var2 * jnp.exp(-0.5 * (x2_test[:, None] - x2[None, :]) ** 2 / ls2**2)
K_cross_factors = [K_cross_1, K_cross_2]
# Per-dimension test diagonals: (n_test_d,)
K_test_diag_1 = var1 * jnp.ones(n1_test)
K_test_diag_2 = var2 * jnp.ones(n2_test)
K_test_diag_factors = [K_test_diag_1, K_test_diag_2]
# Posterior predictions
pred_mean, pred_var = gaussx.kronecker_posterior_predictive(
K_factors,
y_grid_noisy,
noise_var_grid,
grid_shape,
K_cross_factors,
K_test_diag_factors=K_test_diag_factors,
)
print(f"Posterior mean shape: {pred_mean.shape}")
print(f"Posterior var shape: {pred_var.shape}")Posterior mean shape: (400,)
Posterior var shape: (400,)
X1_test, X2_test = jnp.meshgrid(x1_test, x2_test, indexing="ij")
f_test_true = jnp.sin(X1_test) * jnp.cos(X2_test)
fig, axes = plt.subplots(1, 3, figsize=(14, 4))
im0 = axes[0].contourf(X1_test, X2_test, f_test_true, levels=20, cmap="RdBu_r")
axes[0].set_title("True function")
plt.colorbar(im0, ax=axes[0])
im1 = axes[1].contourf(
X1_test, X2_test, pred_mean.reshape(n1_test, n2_test), levels=20, cmap="RdBu_r"
)
axes[1].set_title("Posterior mean")
plt.colorbar(im1, ax=axes[1])
im2 = axes[2].contourf(
X1_test, X2_test, pred_var.reshape(n1_test, n2_test), levels=20, cmap="viridis"
)
axes[2].set_title("Posterior variance")
plt.colorbar(im2, ax=axes[2])
for ax in axes:
ax.set(xlabel="$x_1$", ylabel="$x_2$")
ax.scatter(X1.ravel(), X2.ravel(), c="k", s=5, alpha=0.3, zorder=5)
plt.tight_layout()
plt.show()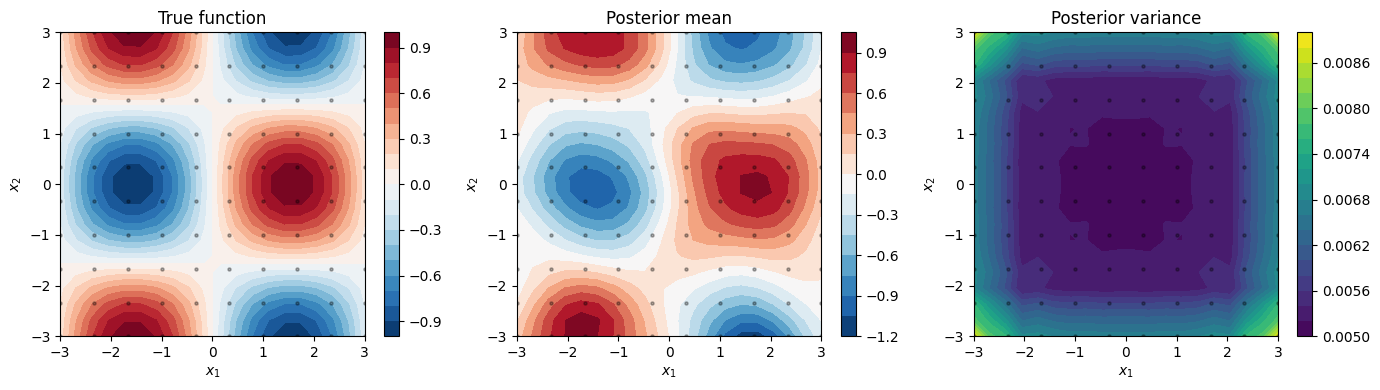
7. Hyperparameter selection: LOO-CV vs MLL¶
We sweep over a range of lengthscales and compare two model selection criteria:
- Marginal log-likelihood (MLL) — the standard Bayesian criterion
- LOO log-predictive density — approximated via LOVE
Both should favour similar lengthscales, but LOO-CV can be more robust when the model is misspecified.
lengthscales = jnp.linspace(0.3, 3.0, 20)
mll_scores = []
loo_scores = []
for ls_val in lengthscales:
ls_val = float(ls_val)
# Build kernel matrix
K_ls = rbf_kernel(X_train, X_train, ls_val, 1.0)
K_ls_noisy = K_ls + noise_var * jnp.eye(n_train)
K_ls_op = lx.MatrixLinearOperator(K_ls_noisy, lx.positive_semidefinite_tag)
# MLL
alpha_ls = gaussx.solve(K_ls_op, y_train)
sign_ls, logdet_ls = jnp.linalg.slogdet(K_ls_noisy)
mll_val = -0.5 * (y_train @ alpha_ls + logdet_ls + n_train * jnp.log(2 * jnp.pi))
mll_scores.append(float(mll_val))
# LOO-CV via LOVE
cache_ls = gaussx.love_cache(K_ls_op, lanczos_order=50, key=jax.random.PRNGKey(0))
K_inv_diag_ls = jnp.sum(cache_ls.Q**2 * cache_ls.inv_eigvals[None, :], axis=1)
# Clamp diagonal to positive values to avoid negative variance
K_inv_diag_ls = jnp.clip(K_inv_diag_ls, a_min=1e-12)
mu_loo_ls = y_train - alpha_ls / K_inv_diag_ls
var_loo_ls = 1.0 / K_inv_diag_ls
loo_val = -0.5 * n_train * jnp.log(2 * jnp.pi) - 0.5 * jnp.sum(
jnp.log(var_loo_ls) + (y_train - mu_loo_ls) ** 2 / var_loo_ls
)
loo_scores.append(float(loo_val))
mll_scores = jnp.array(mll_scores)
loo_scores = jnp.array(loo_scores)fig, axes = plt.subplots(1, 2, figsize=(12, 4))
axes[0].plot(lengthscales, mll_scores, "o-", label="MLL", lw=2)
axes[0].axvline(
lengthscales[jnp.argmax(mll_scores)], color="tab:blue", linestyle="--", alpha=0.5
)
axes[0].set(
xlabel="Lengthscale", ylabel="Log-likelihood", title="Marginal log-likelihood"
)
axes[0].legend(fontsize=9)
axes[0].grid(True, which="major", alpha=0.3)
axes[0].grid(True, which="minor", alpha=0.1)
axes[0].minorticks_on()
axes[1].plot(
lengthscales,
loo_scores,
"s-",
color="tab:orange",
label="LOO-LPD (LOVE)",
lw=2,
)
axes[1].axvline(
lengthscales[jnp.argmax(loo_scores)], color="tab:orange", linestyle="--", alpha=0.5
)
axes[1].set(
xlabel="Lengthscale", ylabel="Log-predictive density", title="LOO-CV score (LOVE)"
)
axes[1].legend(fontsize=9)
axes[1].grid(True, which="major", alpha=0.3)
axes[1].grid(True, which="minor", alpha=0.1)
axes[1].minorticks_on()
plt.tight_layout()
plt.show()
best_ls_mll = float(lengthscales[jnp.argmax(mll_scores)])
best_ls_loo = float(lengthscales[jnp.argmax(loo_scores)])
print(f"Best lengthscale (MLL): {best_ls_mll:.2f}")
print(f"Best lengthscale (LOO-CV): {best_ls_loo:.2f}")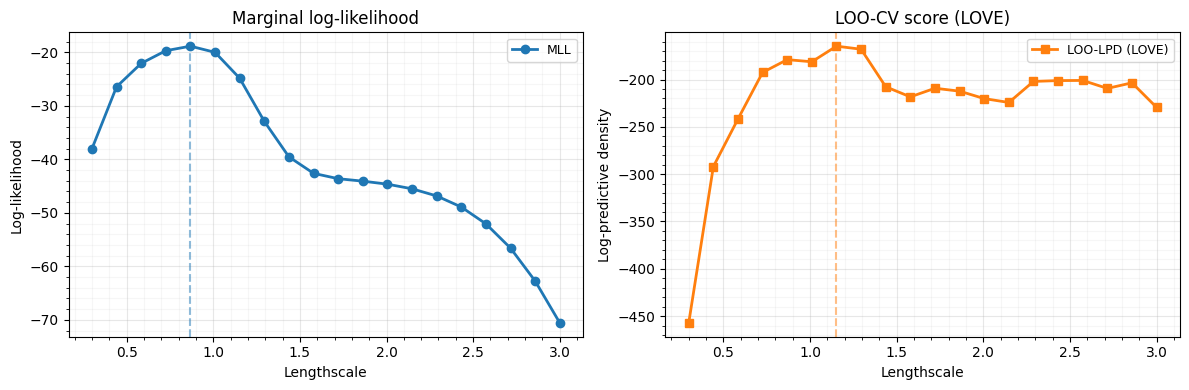
Best lengthscale (MLL): 0.87
Best lengthscale (LOO-CV): 1.15
8. Summary¶
| Method | Function | Cost |
|---|---|---|
| LOVE cache | love_cache | one-time |
| LOVE var | love_variance | /point |
| LOO-CV | Cache diagonal | all pts |
| Kron MLL | kronecker_mll | |
| Kron pred | kronecker_posterior_predictive |
Key takeaways:
love_cachecomputes a Lanczos factorization once;love_variancereuses it for fast predictive variance at any test point.- LOO-CV diagonals come for free from the LOVE cache -- no additional solves needed.
- Kronecker structure turns grid GPs from into .
- MLL and LOO-CV generally agree on hyperparameters, but LOO-CV provides a complementary, directly predictive criterion.
References:
- Pleiss, G., Gardner, J., Weinberger, K., & Wilson, A. G. (2018). Constant-time predictive distributions for Gaussian processes. ICML.
- Rasmussen, C. E. & Williams, C. K. I. (2006). Gaussian Processes for Machine Learning. MIT Press.