GP on a 2D Grid with Kronecker Structure
When data lives on a regular grid, the kernel matrix factorizes as . This enables GP inference on grids with thousands of points that would be intractable with a dense kernel.
This example fits a 2D GP to noisy observations on a grid using Kronecker structure for solve, logdet, and prediction.
Background¶
When inputs lie on a Cartesian product grid and the kernel is separable,
the kernel matrix factorizes as a Kronecker product where and . With , this reduces:
- Storage from to .
- Solve from to .
- Log-determinant via .
These savings make GP inference feasible on grids with thousands of points that would be intractable with a dense kernel matrix. See Saatci (2012) for a comprehensive treatment and Gilboa et al. (2015) for extensions to higher-dimensional grids.
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import jax
import jax.numpy as jnp
import lineax as lx
import matplotlib.pyplot as plt
import gaussx
jax.config.update("jax_enable_x64", True)Generate 2D grid data¶
# Grid
nx, ny = 15, 15
N = nx * ny
x = jnp.linspace(0, 4, nx)
y = jnp.linspace(0, 4, ny)
xx, yy = jnp.meshgrid(x, y, indexing="ij")
# True function: a smooth 2D surface
def f_true(x, y):
return jnp.sin(x) * jnp.cos(1.5 * y)
z_true = f_true(xx, yy)
# Noisy observations
noise_std = 0.15
key = jax.random.PRNGKey(7)
z_obs = z_true + noise_std * jax.random.normal(key, z_true.shape)
print(f"Grid: {nx}x{ny} = {N} points")Grid: 15x15 = 225 points
fig, axes = plt.subplots(1, 2, figsize=(10, 4))
im0 = axes[0].pcolormesh(xx, yy, z_true, cmap="RdBu_r", shading="auto")
axes[0].set_title("True function")
plt.colorbar(im0, ax=axes[0])
im1 = axes[1].pcolormesh(xx, yy, z_obs, cmap="RdBu_r", shading="auto")
axes[1].set_title(f"Noisy observations ($\\sigma$={noise_std})")
plt.colorbar(im1, ax=axes[1])
for ax in axes:
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.set_aspect("equal")
plt.tight_layout()
plt.show()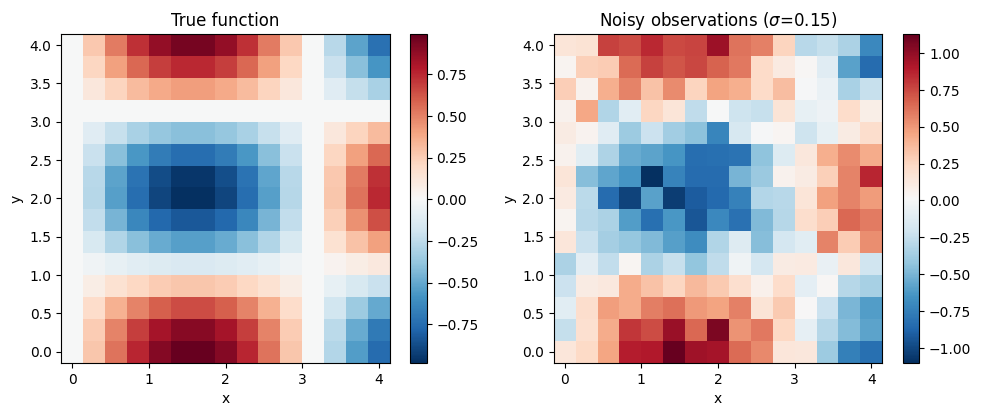
Build Kronecker kernel¶
Separable RBF:
Separability holds for any product of 1D stationary kernels (RBF, Matern, periodic). For non-separable kernels, sums of Kronecker products can be used as approximations.
def rbf_1d(coords, lengthscale, variance):
sq_dist = (coords[:, None] - coords[None, :]) ** 2
return variance * jnp.exp(-0.5 * sq_dist / lengthscale**2)
ls_x, ls_y = 1.0, 1.0
var_x, var_y = 1.0, 1.0
noise_var = noise_std**2
Kx = rbf_1d(x, ls_x, var_x)
Ky = rbf_1d(y, ls_y, var_y)
Kx_op = lx.MatrixLinearOperator(Kx, lx.positive_semidefinite_tag)
Ky_op = lx.MatrixLinearOperator(Ky, lx.positive_semidefinite_tag)
# Full kernel: Kx kron Ky + noise * I
K_kron = gaussx.Kronecker(Kx_op, Ky_op)
print(f"Kronecker kernel: {K_kron.in_size()}x{K_kron.out_size()}")
print(f"Storage: {nx}x{nx} + {ny}x{ny} = {nx**2 + ny**2}")
print(f"Dense would be: {N}x{N} = {N**2:,}")Kronecker kernel: 225x225
Storage: 15x15 + 15x15 = 450
Dense would be: 225x225 = 50,625
Add noise and solve¶
We need . For now we add noise to the dense matrix; a full implementation would keep this structured.
K_noisy = K_kron.as_matrix() + noise_var * jnp.eye(N)
op = lx.MatrixLinearOperator(K_noisy, lx.positive_semidefinite_tag)
# Flatten observations for solve
z_vec = z_obs.ravel()
alpha = gaussx.solve(op, z_vec)
# Posterior mean on the same grid
z_pred = (K_kron.as_matrix() @ alpha).reshape(nx, ny)
print(f"Solve residual: {jnp.max(jnp.abs(op.mv(alpha) - z_vec)):.2e}")Solve residual: 3.15e-14
Log-marginal likelihood¶
The log-marginal likelihood (LML) for GP regression is
Note that the noise term breaks the pure Kronecker structure of . However, eigendecomposition-based approaches (Saatci, 2012) or Krylov methods can handle efficiently by working with the per-factor eigenvalues.
def lml(z, op):
alpha = gaussx.solve(op, z)
n = len(z)
ld = gaussx.logdet(op)
return -0.5 * jnp.dot(z, alpha) - 0.5 * ld - 0.5 * n * jnp.log(2 * jnp.pi)
print(f"Log-marginal likelihood: {lml(z_vec, op):.2f}")Log-marginal likelihood: 55.04
Visualize results¶
fig, axes = plt.subplots(1, 3, figsize=(14, 4))
im0 = axes[0].pcolormesh(xx, yy, z_obs, cmap="RdBu_r", shading="auto")
axes[0].set_title("Observations")
plt.colorbar(im0, ax=axes[0])
im1 = axes[1].pcolormesh(xx, yy, z_pred, cmap="RdBu_r", shading="auto")
axes[1].set_title("GP posterior mean")
plt.colorbar(im1, ax=axes[1])
im2 = axes[2].pcolormesh(xx, yy, jnp.abs(z_true - z_pred), cmap="Reds", shading="auto")
axes[2].set_title("|True - Predicted|")
plt.colorbar(im2, ax=axes[2])
for ax in axes:
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.set_aspect("equal")
plt.tight_layout()
plt.show()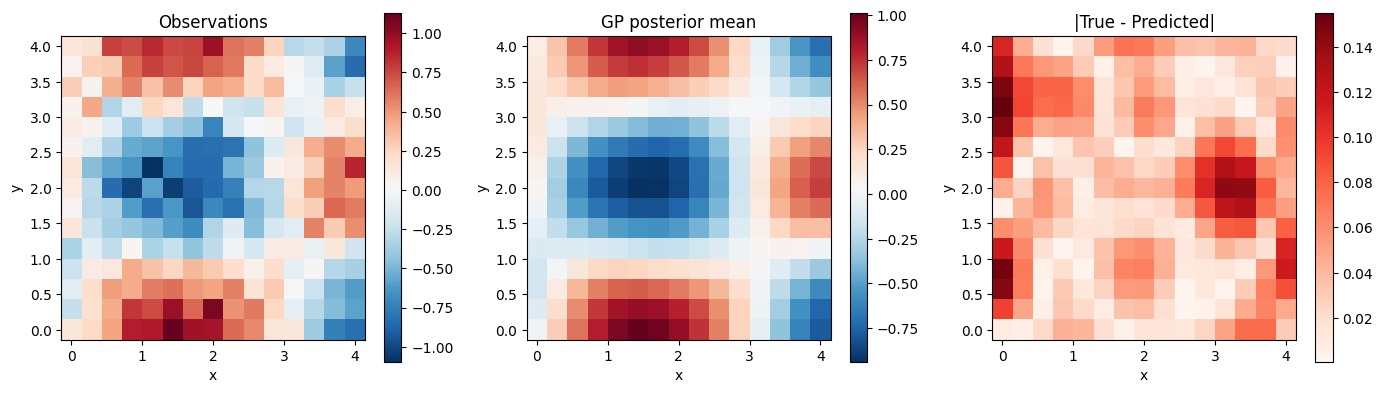
Kronecker-only operations¶
Even though we fell back to dense for the noisy solve, the Kronecker structure enables cheap operations on the kernel itself.
print("logdet(Kx kron Ky):")
print(f" Structured: {gaussx.logdet(K_kron):.4f}")
print(f" Dense: {jnp.linalg.slogdet(K_kron.as_matrix())[1]:.4f}")
print("\ntrace(Kx kron Ky):")
print(f" Structured: {gaussx.trace(K_kron):.4f}")
print(f" Dense: {jnp.trace(K_kron.as_matrix()):.4f}")
L_kron = gaussx.cholesky(K_kron)
print(f"\ncholesky type: {type(L_kron).__name__}")logdet(Kx kron Ky):
Structured: -4064.3464
Dense: -3904.5906
trace(Kx kron Ky):
Structured: 225.0000
Dense: 225.0000
cholesky type: Kronecker
References¶
- Gilboa, E., Saatci, Y., & Cunningham, J. P. (2015). Scaling multidimensional inference for structured Gaussian processes. IEEE TPAMI, 37(2), 424--436.
- Rasmussen, C. E. & Williams, C. K. I. (2006). Gaussian Processes for Machine Learning. MIT Press.
- Saatci, Y. (2012). Scalable Inference for Structured Gaussian Process Models. PhD thesis, University of Cambridge.