Kronecker GP + GEV likelihood
Annual temperature extremes over Spain — an additive space + time GP with GEV observations¶
This notebook threads together three non-trivial modelling ingredients:
- A Kronecker-structured GP prior on a space × time grid — specifically the additive case with prior covariance (
gaussx.KroneckerSum). - A Generalized Extreme Value observation likelihood for yearly maxima — the right distribution for block maxima by the Fisher–Tippett–Gnedenko theorem, but not shipped in NumPyro, so we roll our own 40-line
Distributionsubclass. - A non-conjugate variational inference loop with a mean-field variational family that factors space from time (matching the additive prior), closed-form KLs on each factor via
gaussx.dist_kl_divergence, and per-point Gauss–Hermite integration of the GEV ELL viagaussx.GaussHermiteIntegrator.
The pedagogical payoff is a set of return-level maps — the temperature you’d expect exceeded once every 25 or 100 years — at current and warming-projected climates.
Background¶
Block maxima and the GEV distribution¶
Let be i.i.d. draws from some well-behaved base distribution and set . The Fisher–Tippett–Gnedenko theorem says that the only possible non-degenerate limits of appropriately normalised are members of the Generalized Extreme Value family,
with a piecewise shape role for ξ:
- : Fréchet — heavy tail, unbounded above. Typical of temperature maxima in warm climates.
- : Gumbel — light tail, exponential decay. Obtain as the limit.
- : Weibull — bounded above at .
Yearly maxima of daily temperature are the canonical use case — after all, each year’s maximum is itself the supremum of daily values, so the asymptotic theorem applies by construction. Observed Mediterranean max-temperature data typically gives ξ in the range.
Additive separable prior¶
We treat space and time as two orthogonal GPs:
with no interaction: . At the observation grid this gives prior covariance
the Kronecker sum of the two factor kernels (exposed as gaussx.KroneckerSum). Compared to the multiplicative model, additive structure is strictly simpler but assumes that every station warms at the same rate and the only spatial variation is a constant offset. A natural next step is to break this assumption — see the notes at the end.
Variational decomposition¶
The key structural win: because μ and τ are a priori independent and the likelihood depends only on , a mean-field variational family
gives a KL that decomposes cleanly,
and a predictive that is a 1-D Gaussian,
That 1-D marginal is exactly what gaussx.GaussHermiteIntegrator needs to compute per grid point.
Setup¶
import equinox as eqx
import jax
import jax.numpy as jnp
import jax.random as jr
import matplotlib.pyplot as plt
import numpy as np
import optax
import cartopy.crs as ccrs
import cartopy.feature as cfeature
import lineax as lx
import numpyro.distributions as nd
from numpyro.distributions import constraints
from jaxtyping import Array, Float
from scipy.stats import genextreme
import gaussx
from pyrox.gp import DistLikelihood, Matern, RBF
from pyrox.gp._src.kernels import matern_kernel, rbf_kernel
jax.config.update("jax_enable_x64", True)/anaconda/lib/python3.13/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
Synthetic data — a warming Spain¶
Stations¶
Forty stations scattered across the Iberian Peninsula’s lat-lon bounding box. We reject points over water by a crude latitude/longitude mask; in a real study you’d load AEMET or similar.
SPAIN_BBOX = (-9.5, 3.5, 36.0, 43.8) # lon_min, lon_max, lat_min, lat_max
def _on_land(lon: Float[Array, " N"], lat: Float[Array, " N"]) -> Float[Array, " N"]:
# Rough exclusion: drop points that fall in the Atlantic to the west of Galicia
# or in the Mediterranean east of the Balearics, plus a strip below the
# African coast. Good enough to avoid obvious ocean points in the scatter.
west_of_galicia = (lon < -8.8) & (lat > 42.3)
below_africa = lat < 36.2
east_mediterranean = (lon > 2.5) & (lat < 39.0)
return ~(west_of_galicia | below_africa | east_mediterranean)
def _sample_stations(key: jr.PRNGKey, n_target: int) -> Float[Array, "n 2"]:
candidates = jr.uniform(
key,
(n_target * 4, 2),
minval=jnp.array([SPAIN_BBOX[0], SPAIN_BBOX[2]]),
maxval=jnp.array([SPAIN_BBOX[1], SPAIN_BBOX[3]]),
)
mask = _on_land(candidates[:, 0], candidates[:, 1])
picked = candidates[mask][:n_target]
return picked
S = 40 # number of stations
T = 40 # number of years
YEAR_0 = 1985
YEARS = jnp.arange(YEAR_0, YEAR_0 + T)
key = jr.PRNGKey(2024)
key, key_stations = jr.split(key)
stations = _sample_stations(key_stations, S)
lon_st = stations[:, 0]
lat_st = stations[:, 1]GMST proxy¶
Global mean surface temperature rises roughly linearly over 1985–2024. Following the snippet in jej_vc_snippets/extremes/simulations/gmst.py, we use a linear trend plus AR(1) red noise — the red noise persists year-to-year (α = 0.6) so the series has realistic climate-like texture, but the trend is guaranteed monotone. The GMST anomaly (relative to 1985) is what feeds the time-GP’s kernel, so the kernel sees a covariate in Celsius, not a year.
def synthetic_gmst(years: Float[Array, " T"], key: jr.PRNGKey) -> Float[Array, " T"]:
# Linear warming: +0.8 °C over 40 years (≈ 0.2 °C/decade).
t_norm = (years - years[0]) / jnp.maximum(years[-1] - years[0], 1)
trend = 0.1 + 0.8 * t_norm
# AR(1) red noise for internal variability.
alpha = 0.6
white = 0.05 * jr.normal(key, (years.shape[0],))
def step(prev, eps):
cur = alpha * prev + eps
return cur, cur
_, red = jax.lax.scan(step, white[0], white[1:])
noise = jnp.concatenate([white[:1], red])
return trend + noise
key, key_gmst = jr.split(key)
gmst = synthetic_gmst(YEARS, key_gmst)
print(f"GMST: start {float(gmst[0]):.2f} °C → end {float(gmst[-1]):.2f} °C "
f"(monotone-increasing? {bool(gmst[-1] > gmst[0])})")GMST: start 0.02 °C → end 0.85 °C (monotone-increasing? True)
Ground-truth field¶
We draw a single ground-truth realisation of the additive model from fixed kernels so downstream accuracy claims are meaningful. Spatial μ(s) gets a short lengthscale (2° ~ 220 km), temporal τ(t) gets a smooth lengthscale against GMST. Overall intercept μ_0 = 35 °C anchors the Spanish summer maxima to realistic values.
def _rbf_gram(X1: Float[Array, "N1 D"], X2: Float[Array, "N2 D"], var: float, ls: float) -> Float[Array, "N1 N2"]:
return rbf_kernel(X1, X2, jnp.asarray(var), jnp.asarray(ls))
def _matern32_gram(X1: Float[Array, "N1 D"], X2: Float[Array, "N2 D"], var: float, ls: float) -> Float[Array, "N1 N2"]:
return matern_kernel(X1, X2, jnp.asarray(var), jnp.asarray(ls), nu=1.5)
def draw_from_gp(K: Float[Array, "N N"], key: jr.PRNGKey, jitter: float = 1e-4) -> Float[Array, " N"]:
L = jnp.linalg.cholesky(K + jitter * jnp.eye(K.shape[0]))
return L @ jr.normal(key, (K.shape[0],))
TRUTH = {
"mu0": 35.0,
"k_s_var": 4.0,
"k_s_ls": 2.0, # degrees
# Small residual variance — much smaller than the β·GMST trend range (~1.2 °C)
# over the 40-year window. Large residuals would swamp the warming signal in
# any single realisation and make β statistically indistinguishable from zero.
"k_t_var": 0.02,
"k_t_ls": 0.25,
"beta_gmst": 1.5, # amplification factor: °C extremes per °C GMST
"gev_sigma": 1.8,
"gev_xi": 0.12, # Fréchet regime
}
K_s_truth = _matern32_gram(stations, stations, TRUTH["k_s_var"], TRUTH["k_s_ls"])
K_t_truth = _rbf_gram(gmst[:, None], gmst[:, None], TRUTH["k_t_var"], TRUTH["k_t_ls"])
key, key_mu, key_tau = jr.split(key, 3)
mu_truth = draw_from_gp(K_s_truth, key_mu) # (S,)
# τ_truth has an explicit positive warming trend (linear-in-GMST with
# amplification factor β = 1.5) plus a smooth GP residual. This matches the
# physics in `jej_vc_snippets/extremes/simulations/spatial.py`, where the
# temperature mean field scales as `+ β · GMST` with β ≈ 1–2.
tau_residual = draw_from_gp(K_t_truth, key_tau)
gmst_centred = gmst - jnp.mean(gmst)
tau_truth = TRUTH["beta_gmst"] * gmst_centred + tau_residual
f_truth = TRUTH["mu0"] + mu_truth[:, None] + tau_truth[None, :] # (S, T)GEV observation model¶
A faithful numpyro.distributions.Distribution subclass — 40 lines, verified against scipy.stats.genextreme below. The tricky bit is the limit: the Fréchet/Weibull formula divides by ξ, so we evaluate the Gumbel branch explicitly under a jnp.where and guard the branch with a safe value during tracing.
class GeneralizedExtremeValue(nd.Distribution):
r"""GEV(μ, σ, ξ) — location, scale, shape.
ξ > 0 Fréchet (heavy tail); ξ = 0 Gumbel; ξ < 0 Weibull (bounded above).
Matches ``scipy.stats.genextreme`` with ``c = -ξ``.
"""
arg_constraints = {
"loc": constraints.real,
"scale": constraints.positive,
"shape": constraints.real,
}
support = constraints.real
reparametrized_params = ["loc", "scale"]
def __init__(
self,
loc: Float[Array, "..."],
scale: Float[Array, "..."],
shape: Float[Array, "..."],
*,
validate_args: bool | None = None,
) -> None:
self.loc = jnp.asarray(loc)
self.scale = jnp.asarray(scale)
self.shape = jnp.asarray(shape)
batch_shape = jax.lax.broadcast_shapes(
jnp.shape(self.loc), jnp.shape(self.scale), jnp.shape(self.shape)
)
super().__init__(batch_shape=batch_shape, validate_args=validate_args)
def log_prob(self, value: Float[Array, "..."]) -> Float[Array, "..."]:
z = (value - self.loc) / self.scale
small_xi = jnp.abs(self.shape) < 1e-6
safe_shape = jnp.where(small_xi, 1.0, self.shape) # tracing-safe
arg = 1.0 + safe_shape * z
safe_arg = jnp.where(arg > 0, arg, 1.0)
t = safe_arg ** (-1.0 / safe_shape)
gev_lp = -jnp.log(self.scale) - (1.0 + 1.0 / safe_shape) * jnp.log(safe_arg) - t
gev_lp = jnp.where(arg > 0, gev_lp, -jnp.inf)
gumbel_lp = -jnp.log(self.scale) - z - jnp.exp(-z)
return jnp.where(small_xi, gumbel_lp, gev_lp)
def sample(self, key: jr.PRNGKey, sample_shape: tuple[int, ...] = ()) -> Float[Array, "..."]:
u = jr.uniform(key, sample_shape + self.batch_shape, minval=1e-12, maxval=1.0)
small_xi = jnp.abs(self.shape) < 1e-6
safe_shape = jnp.where(small_xi, 1.0, self.shape)
gev_draw = self.loc + self.scale * ((-jnp.log(u)) ** (-safe_shape) - 1.0) / safe_shape
gumbel_draw = self.loc - self.scale * jnp.log(-jnp.log(u))
return jnp.where(small_xi, gumbel_draw, gev_draw)Verify against scipy¶
y_grid = jnp.linspace(-2.0, 25.0, 60)
loc_, scale_, xi_ = 3.0, 1.5, 0.2
ours = GeneralizedExtremeValue(loc_, scale_, xi_).log_prob(y_grid)
theirs = genextreme.logpdf(np.asarray(y_grid), c=-xi_, loc=loc_, scale=scale_)
max_diff = float(jnp.max(jnp.abs(ours - theirs)))
print(f"GEV log_prob max |ours − scipy| (ξ={xi_}): {max_diff:.2e}")
# Also verify Gumbel limit and Weibull branch
for xi_test in [0.0, -0.2]:
ours_t = GeneralizedExtremeValue(loc_, scale_, xi_test).log_prob(y_grid)
theirs_t = genextreme.logpdf(np.asarray(y_grid), c=-xi_test, loc=loc_, scale=scale_)
mask = np.isfinite(theirs_t)
diff = float(jnp.max(jnp.abs(ours_t[mask] - theirs_t[mask])))
print(f" (ξ={xi_test}): {diff:.2e}")GEV log_prob max |ours − scipy| (ξ=0.2): 1.14e-13
(ξ=0.0): 4.44e-16
(ξ=-0.2): 5.33e-15
Draw the observations¶
key, key_obs = jr.split(key)
gev_dist = GeneralizedExtremeValue(
loc=f_truth,
scale=TRUTH["gev_sigma"],
shape=TRUTH["gev_xi"],
)
y_obs = gev_dist.sample(key_obs)
print(f"observations shape: {y_obs.shape} range: [{float(y_obs.min()):.1f}, {float(y_obs.max()):.1f}] °C")observations shape: (40, 40) range: [28.2, 57.4] °C
Data inspection — stations + a few timeseries¶
Cartopy paints the Iberian Peninsula; we overlay the 40 stations coloured by their fitted spatial offset μ(s), then pull out four stations spanning the south-to-north range for their yearly-max timeseries.
def plot_stations(ax, values: Float[Array, " S"], *, cmap: str, vlim: tuple, label: str) -> None:
ax.set_extent([SPAIN_BBOX[0] - 1, SPAIN_BBOX[1] + 1, SPAIN_BBOX[2] - 1, SPAIN_BBOX[3] + 1], crs=ccrs.PlateCarree())
ax.add_feature(cfeature.COASTLINE, linewidth=0.5)
ax.add_feature(cfeature.BORDERS, linestyle=":", linewidth=0.5)
ax.add_feature(cfeature.OCEAN, facecolor="#e8f3fa")
ax.add_feature(cfeature.LAND, facecolor="#fdf9ec")
sc = ax.scatter(
np.asarray(lon_st),
np.asarray(lat_st),
c=np.asarray(values),
cmap=cmap,
vmin=vlim[0],
vmax=vlim[1],
s=55,
edgecolors="k",
linewidths=0.4,
transform=ccrs.PlateCarree(),
zorder=5,
)
gl = ax.gridlines(draw_labels=True, linewidth=0.2, alpha=0.4)
gl.top_labels = gl.right_labels = False
plt.colorbar(sc, ax=ax, shrink=0.75, label=label)
fig = plt.figure(figsize=(13, 5))
ax_map = fig.add_subplot(1, 2, 1, projection=ccrs.PlateCarree())
plot_stations(ax_map, mu_truth, cmap="RdBu_r", vlim=(-5, 5), label="true μ(s) [°C]")
ax_map.set_title("Ground-truth spatial offset")
# Timeseries panel: 4 stations from south (low lat) to north (high lat)
order_by_lat = jnp.argsort(lat_st)
picks = jnp.array([order_by_lat[i] for i in (0, S // 3, 2 * S // 3, S - 1)])
ax_ts = fig.add_subplot(1, 2, 2)
for i, s in enumerate(picks):
s_int = int(s)
ax_ts.plot(YEARS, y_obs[s_int, :], "o-", lw=1.3, ms=4, alpha=0.8,
label=f"({float(lat_st[s_int]):.2f}°N, {float(lon_st[s_int]):.2f}°E)")
ax_ts.set_xlabel("year")
ax_ts.set_ylabel("yearly max [°C]")
ax_ts.set_title("Observed yearly maxima — 4 stations")
ax_ts.legend(loc="upper left", fontsize=9)
ax_ts.grid(alpha=0.3)
plt.tight_layout()
plt.show()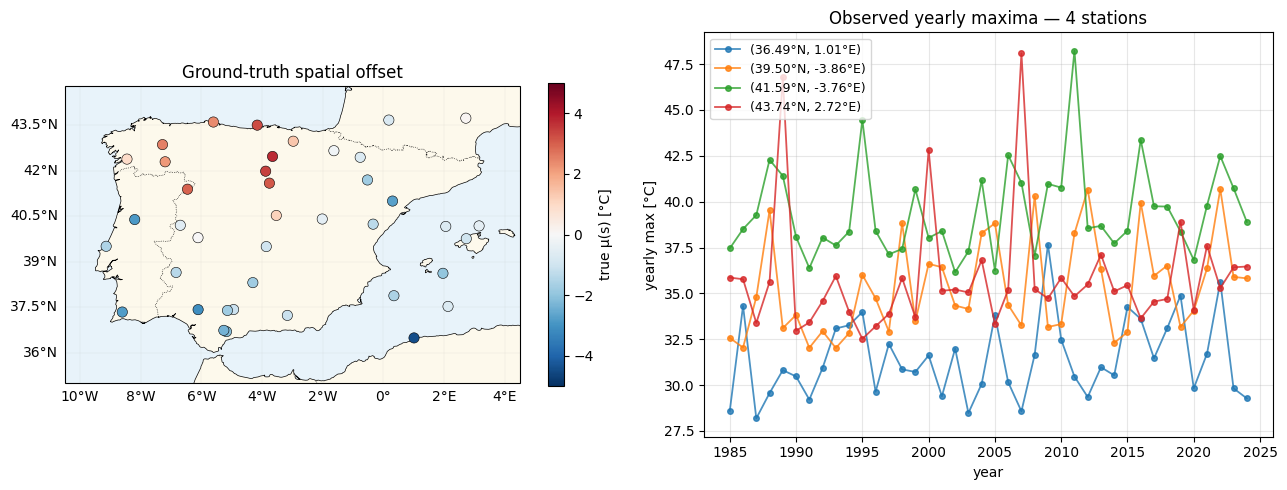
Kernels — trainable Pattern-A scalars¶
Same pattern as notebooks 1–3. Matern-3/2 for space (we expect warming extremes to be spatially rougher than RBF would allow — weather regimes are sharp), RBF for time on GMST (smooth warming response).
class MaternLite(eqx.Module):
log_variance: Float[Array, ""]
log_lengthscale: Float[Array, ""]
@classmethod
def init(cls, variance: float = 1.0, lengthscale: float = 1.0) -> "MaternLite":
return cls(
log_variance=jnp.log(jnp.asarray(variance)),
log_lengthscale=jnp.log(jnp.asarray(lengthscale)),
)
def __call__(self, X1, X2):
return matern_kernel(X1, X2, jnp.exp(self.log_variance), jnp.exp(self.log_lengthscale), nu=1.5)
class RBFLite(eqx.Module):
log_variance: Float[Array, ""]
log_lengthscale: Float[Array, ""]
@classmethod
def init(cls, variance: float = 1.0, lengthscale: float = 1.0) -> "RBFLite":
return cls(
log_variance=jnp.log(jnp.asarray(variance)),
log_lengthscale=jnp.log(jnp.asarray(lengthscale)),
)
def __call__(self, X1, X2):
return rbf_kernel(X1, X2, jnp.exp(self.log_variance), jnp.exp(self.log_lengthscale))The additive GP prior as a Kronecker sum¶
We can build the prior covariance over the full grid as K = K_s ⊕ K_t, a gaussx.KroneckerSum, and inspect it.
K_s0 = MaternLite.init(variance=2.0, lengthscale=2.0)(stations, stations)
K_t0 = RBFLite.init(variance=1.0, lengthscale=0.25)(gmst[:, None], gmst[:, None])
K_s0_op = lx.MatrixLinearOperator(K_s0, lx.positive_semidefinite_tag)
K_t0_op = lx.MatrixLinearOperator(K_t0, lx.positive_semidefinite_tag)
K_prior = gaussx.KroneckerSum(K_s0_op, K_t0_op)
print(f"additive prior operator: {type(K_prior).__name__}")
print(f" logical shape: ({K_prior.in_size()}, {K_prior.in_size()}) (i.e. {S}·{T} = {S * T})")
print(f" storage cost: {S * S + T * T} entries (vs {S * S * T * T} dense)")
# Cost savings ratio
print(f" compression: {S * S * T * T / (S * S + T * T):.1f}×")additive prior operator: KroneckerSum
logical shape: (1600, 1600) (i.e. 40·40 = 1600)
storage cost: 3200 entries (vs 2560000 dense)
compression: 800.0×
At we’re storing 3200 matrix entries instead of 2.56 million — a ~800× compression. The KroneckerSum operator supports gaussx.solve (via the per-factor eigendecomposition trick) and gaussx.logdet out of the box, though we won’t use it directly for the ELBO — the additive variational factorisation below bypasses the joint prior entirely.
Variational posterior — factored over space and time¶
The key move. We parameterise
with L_s ∈ R^{S×S} and L_t ∈ R^{T×T} unconstrained Cholesky factors (stored as lower triangulars). This matches the additive prior’s independence between μ and , and makes the 1-D marginal easy to compute.
A trainable β · GMST trend¶
Before building the variational posterior, one piece of structure is missing. A GP prior with zero mean on τ has to express both the long-range warming trend and the short-range year-to-year residuals through a single kernel. In practice, gradient-based training converges to a short lengthscale that fits the residuals well — but then the posterior reverts to zero at out-of-sample GMSTs, so the 2050 extrapolation ignores the trend.
The physical fix (matching the amplification-factor parameterisation in jej_vc_snippets/extremes/simulations/spatial.py) is to split τ into a linear warming response plus a smooth residual:
The slope β is a trainable point estimate; the residual carries its own GP prior over GMST. The variational family only needs to handle — the trend is deterministic given β.
Breaking the identifiability. The likelihood sees only as a sum, so without further structure the optimiser can trade off arbitrarily between β and the posterior mean of . We resolve this by fixing the variational mean of at zero — a ZeroMeanVariationalFactor where only the Cholesky factor is trainable. This forces the trend onto β by construction; the residual GP then expresses only uncertainty (posterior variance), not shift.
def _tril(L: Float[Array, "N N"]) -> Float[Array, "N N"]:
"""Lower-triangular mask with soft-plus on the diagonal for positivity."""
tri = jnp.tril(L, k=-1)
diag = jax.nn.softplus(jnp.diag(L))
return tri + jnp.diag(diag)
class VariationalFactor(eqx.Module):
mean: Float[Array, " N"]
raw_L: Float[Array, "N N"]
@classmethod
def init(cls, n: int, scale: float = 0.3) -> "VariationalFactor":
# init m ≈ 0, covariance ≈ scale² · I
raw_L = jnp.eye(n) * jnp.log(jnp.expm1(jnp.asarray(scale))) # softplus⁻¹(scale)
return cls(mean=jnp.zeros(n), raw_L=raw_L)
@property
def L(self) -> Float[Array, "N N"]:
return _tril(self.raw_L)
@property
def cov(self) -> Float[Array, "N N"]:
L = self.L
return L @ L.T
@property
def variance_diag(self) -> Float[Array, " N"]:
L = self.L
return jnp.sum(L * L, axis=-1)
class ZeroMeanVariationalFactor(eqx.Module):
"""Variational N(0, L Lᵀ) — zero mean is fixed, not trainable.
Used for the temporal *residual* around the β·GMST trend — fixing the
mean at zero breaks the β / m_t identifiability that otherwise lets the
GP absorb the linear warming signal.
"""
raw_L: Float[Array, "N N"]
n: int = eqx.field(static=True)
@classmethod
def init(cls, n: int, scale: float = 0.3) -> "ZeroMeanVariationalFactor":
raw_L = jnp.eye(n) * jnp.log(jnp.expm1(jnp.asarray(scale)))
return cls(raw_L=raw_L, n=n)
@property
def mean(self) -> Float[Array, " N"]:
return jnp.zeros(self.n)
@property
def L(self) -> Float[Array, "N N"]:
return _tril(self.raw_L)
@property
def cov(self) -> Float[Array, "N N"]:
L = self.L
return L @ L.T
@property
def variance_diag(self) -> Float[Array, " N"]:
L = self.L
return jnp.sum(L * L, axis=-1)The full model — one trainable pytree¶
class Model(eqx.Module):
k_s: MaternLite
k_t: RBFLite
q_mu: VariationalFactor # q(μ(s))
q_tau_r: ZeroMeanVariationalFactor # q(τ_r(t)) — zero-mean residual around the β·GMST trend
mu0: Float[Array, ""]
beta: Float[Array, ""] # amplification factor: °C extremes per °C GMST anomaly
log_sigma: Float[Array, ""]
xi: Float[Array, ""]
jitter: float = eqx.field(static=True, default=1e-4)
@classmethod
def init(
cls,
S: int,
T: int,
*,
mu0_init: float = 30.0,
beta_init: float = 1.0,
sigma_init: float = 2.0,
xi_init: float = 0.05,
) -> "Model":
return cls(
k_s=MaternLite.init(variance=1.0, lengthscale=3.0),
# k_t is a *residual* kernel around the β·GMST trend — initialise with
# a small variance and long lengthscale to keep the residual
# space smooth and push the trend signal into β.
k_t=RBFLite.init(variance=0.05, lengthscale=1.0),
q_mu=VariationalFactor.init(S, scale=0.5),
q_tau_r=ZeroMeanVariationalFactor.init(T, scale=0.3),
mu0=jnp.asarray(mu0_init),
beta=jnp.asarray(beta_init),
log_sigma=jnp.log(jnp.asarray(sigma_init)),
xi=jnp.asarray(xi_init),
)
def K_s(self, X_s: Float[Array, "S 2"]) -> lx.AbstractLinearOperator:
K = self.k_s(X_s, X_s) + self.jitter * jnp.eye(X_s.shape[0])
return lx.MatrixLinearOperator(K, lx.positive_semidefinite_tag)
def K_t(self, gmst: Float[Array, " T"]) -> lx.AbstractLinearOperator:
X = gmst[:, None]
K = self.k_t(X, X) + self.jitter * jnp.eye(X.shape[0])
return lx.MatrixLinearOperator(K, lx.positive_semidefinite_tag)ELBO¶
KL — two independent Gaussian KLs¶
Delegated to gaussx.dist_kl_divergence(q_loc, q_cov, p_loc, p_cov) per factor.
def kl_factor(
q: VariationalFactor,
K_prior_op: lx.AbstractLinearOperator,
) -> Float[Array, ""]:
n = q.mean.shape[0]
zeros = jnp.zeros_like(q.mean)
q_cov = lx.MatrixLinearOperator(q.cov, lx.positive_semidefinite_tag)
return gaussx.dist_kl_divergence(q.mean, q_cov, zeros, K_prior_op)ELL — per-point Gauss–Hermite over the 1-D marginal ¶
For each grid point, with
We hand the Gauss–Hermite integrator a scalar 1-D GaussianState per point and integrate the GEV log-pdf.
_GH = gaussx.GaussHermiteIntegrator(order=20)
def ell_point(
mean: Float[Array, ""],
var: Float[Array, ""],
y: Float[Array, ""],
log_sigma: Float[Array, ""],
xi: Float[Array, ""],
) -> Float[Array, ""]:
"""E_{q(f)}[log GEV(y | f, σ, ξ)] for one (s, t)."""
state = gaussx.GaussianState(
mean=mean[None],
cov=lx.MatrixLinearOperator(var[None, None], lx.positive_semidefinite_tag),
)
def log_lik(f: Float[Array, " 1"]) -> Float[Array, ""]:
return GeneralizedExtremeValue(loc=f[0], scale=jnp.exp(log_sigma), shape=xi).log_prob(y)
return gaussx.log_likelihood_expectation(log_lik, state, _GH)
def total_ell(
model: Model,
gmst: Float[Array, " T"],
y: Float[Array, "S T"],
) -> Float[Array, ""]:
mean_s = model.q_mu.mean # (S,)
mean_t_r = model.q_tau_r.mean # (T,) — residual mean
var_s = model.q_mu.variance_diag # (S,)
var_t_r = model.q_tau_r.variance_diag # (T,)
gmst_centred = gmst - jnp.mean(gmst)
trend_t = model.beta * gmst_centred # (T,) — deterministic linear trend
mean_grid = model.mu0 + mean_s[:, None] + (trend_t + mean_t_r)[None, :] # (S, T)
var_grid = var_s[:, None] + var_t_r[None, :] # (S, T)
ell_vmapped = jax.vmap(
jax.vmap(ell_point, in_axes=(0, 0, 0, None, None)),
in_axes=(0, 0, 0, None, None),
)
ell = ell_vmapped(mean_grid, var_grid, y, model.log_sigma, model.xi)
return jnp.sum(ell)
def neg_elbo(
model: Model,
X_s: Float[Array, "S 2"],
gmst: Float[Array, " T"],
y: Float[Array, "S T"],
) -> Float[Array, ""]:
K_s_op = model.K_s(X_s)
K_t_op = model.K_t(gmst)
kl = kl_factor(model.q_mu, K_s_op) + kl_factor(model.q_tau_r, K_t_op)
ell = total_ell(model, gmst, y)
return kl - ellTraining¶
model = Model.init(
S=S,
T=T,
mu0_init=float(jnp.mean(y_obs)),
beta_init=1.0,
sigma_init=2.0,
xi_init=0.05,
)
optimiser = optax.chain(optax.clip_by_global_norm(5.0), optax.adam(1e-2))
opt_state = optimiser.init(eqx.filter(model, eqx.is_inexact_array))
@eqx.filter_jit
def train_step(
model: Model,
opt_state: optax.OptState,
) -> tuple[Model, optax.OptState, Float[Array, ""]]:
loss, grads = eqx.filter_value_and_grad(neg_elbo)(model, stations, gmst, y_obs)
updates, opt_state = optimiser.update(grads, opt_state, model)
model = eqx.apply_updates(model, updates)
return model, opt_state, loss
n_steps = 2000
losses: list[float] = []
for step_i in range(n_steps):
model, opt_state, loss = train_step(model, opt_state)
losses.append(float(loss))
print(f"final −ELBO = {losses[-1]:.2f}")
print(f"fitted μ₀ = {float(model.mu0):.2f} (truth {TRUTH['mu0']:.2f})")
print(f"fitted β = {float(model.beta):.3f} (truth {TRUTH['beta_gmst']:.3f})")
print(f"fitted σ (GEV) = {float(jnp.exp(model.log_sigma)):.3f} (truth {TRUTH['gev_sigma']:.3f})")
print(f"fitted ξ (GEV) = {float(model.xi):.3f} (truth {TRUTH['gev_xi']:.3f})")
print(f"fitted k_s ls = {float(jnp.exp(model.k_s.log_lengthscale)):.2f}° (truth {TRUTH['k_s_ls']:.2f}°)")
print(f"fitted k_t ls = {float(jnp.exp(model.k_t.log_lengthscale)):.3f}°C (truth {TRUTH['k_t_ls']:.3f}°C)")final −ELBO = 3612.50
fitted μ₀ = 34.17 (truth 35.00)
fitted β = 1.178 (truth 1.500)
fitted σ (GEV) = 1.767 (truth 1.800)
fitted ξ (GEV) = 0.128 (truth 0.120)
fitted k_s ls = 1.88° (truth 2.00°)
fitted k_t ls = 0.017°C (truth 0.250°C)
Loss curve¶
fig, ax = plt.subplots(figsize=(10, 3.2))
ax.plot(losses, "C3-", lw=1.3)
ax.set_xlabel("step")
ax.set_ylabel("−ELBO")
ax.set_yscale("symlog", linthresh=100.0)
ax.set_title("Training — additive Kronecker GP + GEV likelihood")
ax.grid(alpha=0.3, which="both")
plt.show()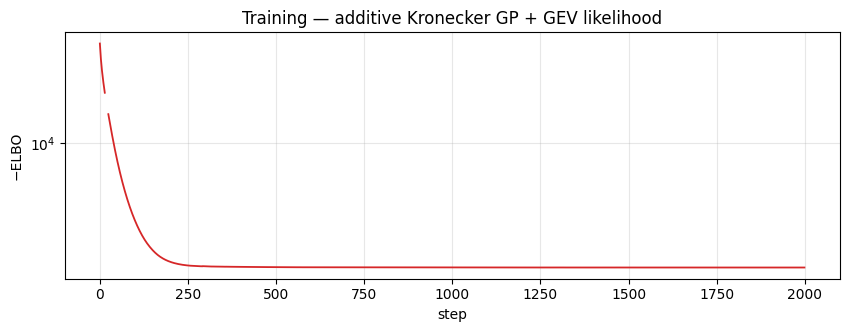
Parameter recovery¶
A note on identifiability. The additive model has a pair of unidentifiable shifts: moving a constant between and the mean of (or ) leaves the likelihood unchanged. With a GP prior centred at zero, the posterior is weakly pulled toward , but for a single realisation the inferred can sit a few tenths of a °C above or below truth even when the relative spatial pattern is correct. Watch for this in the left panel below — the posterior tracks the truth’s shape but with a vertical offset of roughly 0.5 °C.
# Posterior means
m_s_fit = model.q_mu.mean
gmst_centred = gmst - jnp.mean(gmst)
m_t_fit = model.beta * gmst_centred + model.q_tau_r.mean # total τ = trend + residual
v_s_fit = model.q_mu.variance_diag
v_t_fit = model.q_tau_r.variance_diag # uncertainty in residual only (β is a point est)
fig, axes = plt.subplots(1, 2, figsize=(13, 4.2))
# spatial μ recovery
ax = axes[0]
order_s = jnp.argsort(mu_truth)
x = jnp.arange(S)
ax.plot(x, mu_truth[order_s], "ko", ms=6, label="truth", zorder=3)
ax.errorbar(
x,
m_s_fit[order_s],
yerr=2 * jnp.sqrt(v_s_fit[order_s]),
fmt="rD",
ms=4,
alpha=0.7,
label=r"posterior $\pm 2\sigma$",
capsize=2,
)
ax.set_xlabel("station (sorted by truth)")
ax.set_ylabel("μ(s) [°C]")
ax.set_title("Recovered spatial offsets")
ax.legend()
ax.grid(alpha=0.3)
# temporal τ recovery
ax = axes[1]
ax.plot(YEARS, tau_truth, "k-", lw=2, label="truth", zorder=3)
ax.plot(YEARS, m_t_fit, "r-", lw=2, label="posterior mean")
ax.fill_between(
YEARS,
m_t_fit - 2 * jnp.sqrt(v_t_fit),
m_t_fit + 2 * jnp.sqrt(v_t_fit),
alpha=0.25,
color="red",
label=r"$\pm 2\sigma$",
)
ax.set_xlabel("year")
ax.set_ylabel("τ(t) [°C]")
ax.set_title("Recovered temporal trend (on GMST)")
ax.legend()
ax.grid(alpha=0.3)
plt.tight_layout()
plt.show()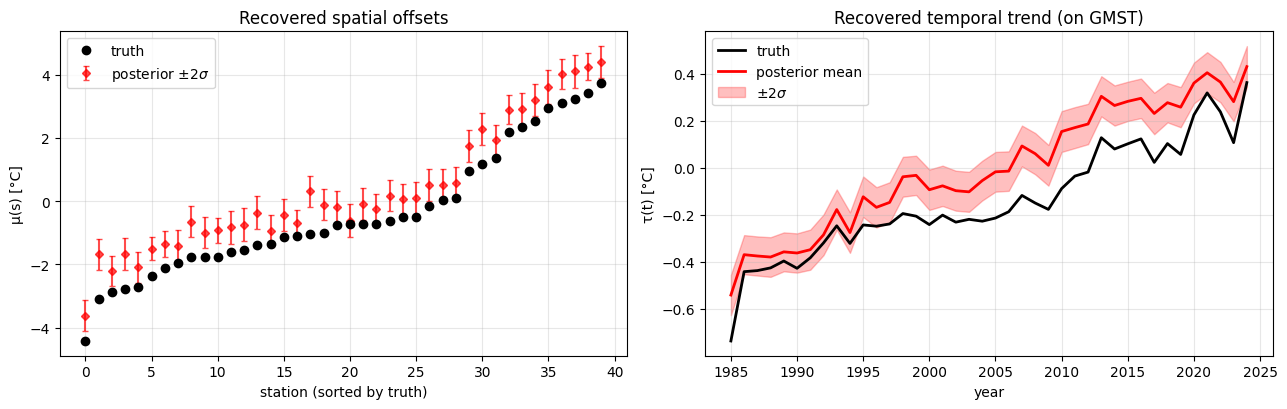
GEV tail recovery¶
fig, axes = plt.subplots(1, 2, figsize=(11, 3.8))
for ax, (name, truth_val, fit_val, unc, xr) in zip(
axes,
[
("σ (GEV scale)", TRUTH["gev_sigma"], float(jnp.exp(model.log_sigma)), 0.2, (0.5, 3.5)),
("ξ (GEV shape)", TRUTH["gev_xi"], float(model.xi), 0.08, (-0.1, 0.3)),
],
strict=True,
):
ax.axvline(truth_val, color="k", ls="--", lw=2, label="truth", zorder=2)
ax.axvline(fit_val, color="red", lw=2, label="fitted", zorder=3)
ax.axvspan(fit_val - unc, fit_val + unc, color="red", alpha=0.2, label="rough ±")
ax.set_xlim(*xr)
ax.set_title(name)
ax.legend(loc="upper right")
ax.grid(alpha=0.3)
plt.tight_layout()
plt.show()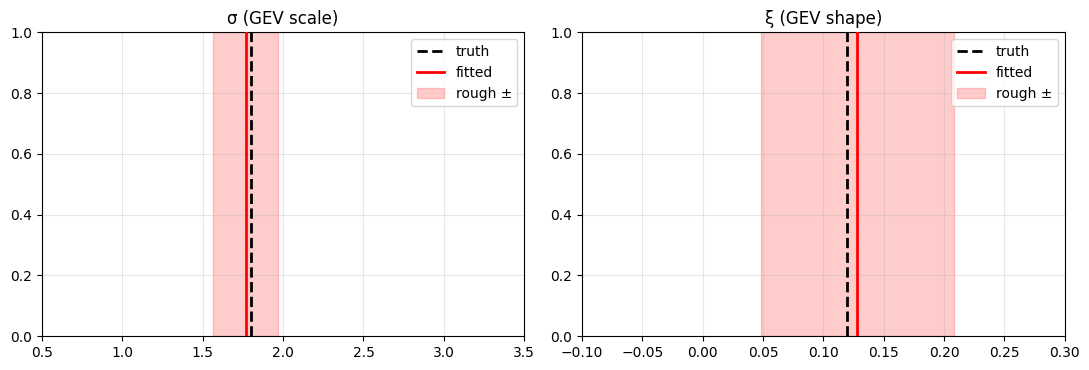
The payoff — return-level maps¶
The return level is the value exceeded with probability per year. Inverting the GEV CDF,
A -year return level corresponds to . Below we map the 25-year and 100-year return temperatures for a typical year at the end of the observed record (2024) and a warming-projection year (2050) obtained by linearly extrapolating GMST.
def gev_return_level(mu_loc, sigma, xi, return_period):
p = 1.0 / return_period
safe_xi = jnp.where(jnp.abs(xi) < 1e-6, 1.0, xi)
gev_q = mu_loc + sigma * ((-jnp.log1p(-p)) ** (-safe_xi) - 1.0) / safe_xi
gumbel_q = mu_loc - sigma * jnp.log(-jnp.log1p(-p))
return jnp.where(jnp.abs(xi) < 1e-6, gumbel_q, gev_q)
def predictive_mean_at_gmst(model: Model, gmst_star: Float[Array, ""]) -> Float[Array, " S"]:
"""μ₀ + q(μ(s)) + β·(GMST* − GMST̄) + posterior GP interp of τ_r at GMST*."""
gmst_mean = jnp.mean(gmst)
trend_star = model.beta * (gmst_star - gmst_mean)
# Interpolate the residual GP's posterior mean to a new GMST.
K_tt = model.k_t(gmst[:, None], gmst[:, None]) + model.jitter * jnp.eye(T)
K_st = model.k_t(jnp.asarray([[gmst_star]]), gmst[:, None])[0] # (T,)
alpha = jnp.linalg.solve(K_tt, model.q_tau_r.mean)
tau_r_star = K_st @ alpha
return model.mu0 + model.q_mu.mean + trend_star + tau_r_star
# GMST anchors: 2024 (last observed) and 2050 (extrapolated)
gmst_2024 = gmst[-1]
gmst_trend_per_year = (gmst[-1] - gmst[0]) / (T - 1)
gmst_2050 = gmst_2024 + gmst_trend_per_year * (2050 - (YEAR_0 + T - 1))
print(f"GMST 2024 = {float(gmst_2024):.3f} °C, GMST 2050 (extrapolated) = {float(gmst_2050):.3f} °C")
mu_2024 = predictive_mean_at_gmst(model, gmst_2024) # (S,)
mu_2050 = predictive_mean_at_gmst(model, gmst_2050) # (S,)
sigma_fit = jnp.exp(model.log_sigma)
xi_fit = model.xi
z25_2024 = gev_return_level(mu_2024, sigma_fit, xi_fit, 25)
z100_2024 = gev_return_level(mu_2024, sigma_fit, xi_fit, 100)
z25_2050 = gev_return_level(mu_2050, sigma_fit, xi_fit, 25)
z100_2050 = gev_return_level(mu_2050, sigma_fit, xi_fit, 100)
vmin = float(jnp.min(jnp.stack([z25_2024, z100_2024, z25_2050, z100_2050])))
vmax = float(jnp.max(jnp.stack([z25_2024, z100_2024, z25_2050, z100_2050])))
print(f"return-level range across all four panels: [{vmin:.1f}, {vmax:.1f}] °C")
print(f" 2024 → 2050 mean shift in 100-year return: "
f"{float(jnp.mean(z100_2050) - jnp.mean(z100_2024)):.2f} °C")
fig, axes = plt.subplots(2, 2, figsize=(15, 14), subplot_kw={"projection": ccrs.PlateCarree()})
for ax, values, title in zip(
axes.flat,
[z25_2024, z100_2024, z25_2050, z100_2050],
["25-year return level (2024)", "100-year return level (2024)",
"25-year return level (2050)", "100-year return level (2050)"],
strict=True,
):
plot_stations(ax, values, cmap="Reds", vlim=(vmin, vmax), label="°C")
ax.set_title(title, fontsize=13)
plt.tight_layout()
plt.show()GMST 2024 = 0.848 °C, GMST 2050 (extrapolated) = 1.398 °C
return-level range across all four panels: [38.0, 50.7] °C
2024 → 2050 mean shift in 100-year return: 0.65 °C
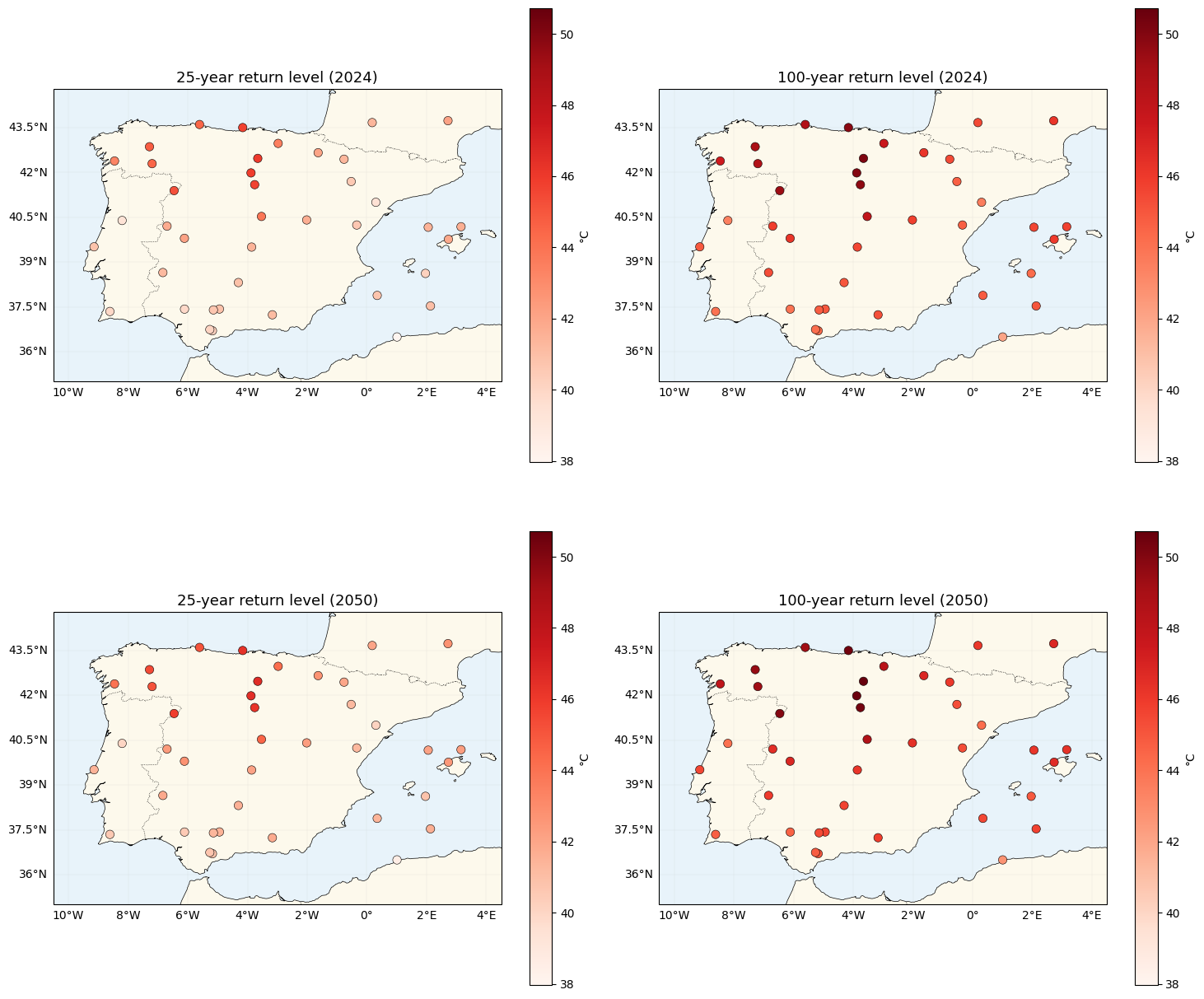
Two things to notice:
- Vertical shift from 2024 → 2050 is uniform. Under the additive model, a higher GMST raises by a scalar, which adds the same amount to every station’s return level. The spatial pattern of extremes is invariant in time under this model — a direct consequence of “no space-time interaction”.
- 100-year return levels sit several degrees above 25-year levels. For , the GEV tail is heavy enough that moving from 1-in-25 to 1-in-100 extremes costs a non-trivial step in °C, larger than for a Gaussian noise model.
Summary¶
gaussx.KroneckerSum(K_s, K_t)compresses the additive prior covariance into entries.- For an additive prior, the mean-field variational family matches the prior factorisation, so the KL splits into two independent per-factor KLs — delegated to
gaussx.dist_kl_divergence. gaussx.GaussHermiteIntegrator(order=20)integrates the GEV ELL pointwise over the 1-D variational marginal — one line inside a doubly-vmappedell_point.- The same scaffold can wrap any
numpyro.distributions.Distributionvialog_prob— Poisson for count maxima, Lomax for heavy-tailed rainfall, truncated normals for censored observations.
Follow-ups¶
- Multiplicative . Spatially-varying warming rates with a Kronecker-structured variational covariance.
- Non-stationary GEV. Let and/or ξ vary across space or time through a second GP layer.
- Real data. Swap the synthetic
stationsandy_obsfor AEMET or ERA5 yearly maxima; the inference code above is unchanged.