Sparse variational Markov GP
SparseMarkovGPPrior is the inducing-time-grid analogue of MarkovGPPrior. It wraps any SDEKernel and a sorted inducing grid Z of length M, exposing the standard SVGP building blocks derived from the SDE autocovariance .
Because the predictive-block contract matches the dense SparseGPPrior, the existing variational guides (FullRankGuide, MeanFieldGuide, WhitenedGuide) plug straight in. The sparse_markov_elbo function delivers a Gaussian-likelihood closed-form ELBO and accepts a gaussx integrator for non-conjugate likelihoods.
Cost: per ELBO evaluation. A truly linear-in- Kalman-aware ELBO (BayesNewton / MarkovFlow style) is a follow-up.
import jax.numpy as jnp
import jax.random as jr
import matplotlib.pyplot as plt
import numpy as np
from pyrox.gp import (
FullRankGuide,
GaussianLikelihood,
MarkovGPPrior,
MaternSDE,
SparseConditionedMarkovGP,
SparseMarkovGPPrior,
sparse_markov_elbo,
)
plt.rcParams["figure.dpi"] = 110
key = jr.PRNGKey(0)/home/azureuser/localfiles/pyrox/.venv/lib/python3.13/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
Synthetic regression data¶
Noisy mixture of two sinusoids on an irregular time grid.
N = 80
t_uniform = jnp.linspace(0.0, 8.0, N)
times = jnp.sort(t_uniform + 0.04 * jr.normal(jr.PRNGKey(1), (N,)))
f_true = 1.2 * jnp.sin(0.9 * times) + 0.4 * jnp.cos(2.7 * times)
y = f_true + 0.18 * jr.normal(key, (N,))
fig, ax = plt.subplots(figsize=(8, 3))
ax.scatter(times, y, c="C3", s=14, label="data")
ax.plot(times, f_true, "k--", lw=1, label="true f")
ax.set_xlabel("t")
ax.set_ylabel("y")
ax.legend(fontsize=8)
plt.tight_layout()
plt.show()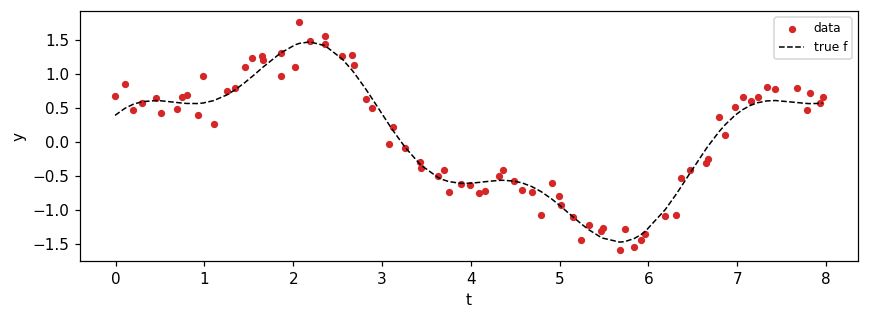
Sparse Markov GP setup¶
Pick M = 12 evenly-spaced inducing times and a Matern-3/2 SDE prior. The variational guide is a FullRankGuide over the inducing function values .
M = 12
Z = jnp.linspace(times[0], times[-1], M)
sde = MaternSDE(variance=1.0, lengthscale=0.6, order=1)
prior = SparseMarkovGPPrior(sde, Z, jitter=1e-6)
guide = FullRankGuide.init(M, scale=0.3)
lik = GaussianLikelihood(noise_var=0.05)
print(f"M (inducing) = {M}")
print(f"N (data) = {N}")
print(f"prior marginal variance = {float(prior._stationary_variance()):.3f}")
print(f"initial ELBO = {float(sparse_markov_elbo(prior, guide, lik, times, y)):.3f}")M (inducing) = 12
N (data) = 80
prior marginal variance = 1.000
initial ELBO = -771.188
Conjugate update — closed-form q(u) for Gaussian likelihood¶
For Gaussian observations the optimal is closed-form (Titsias 2009):
This sidesteps the need to optimise the ELBO and gives the optimal at the prescribed inducing locations directly. We use it here to demonstrate the sparse_markov_elbo / SparseConditionedMarkovGP API on a fitted guide.
import equinox as eqx
K_zz_op, K_xz, _K_xx_diag = prior.predictive_blocks(times)
K_zz = K_zz_op.as_matrix()
sigma2 = 0.05 # observation variance
mu_x = prior.mean(times)
# A = K_zz^{-1} K_xz^T (M, N)
A = jnp.linalg.solve(K_zz, K_xz.T)
# Posterior precision in the inducing-value coords:
# Lambda_q = K_zz^{-1} + sigma^{-2} A A^T
Lambda_q = jnp.linalg.inv(K_zz) + (1.0 / sigma2) * A @ A.T
S_q = jnp.linalg.inv(Lambda_q)
m_q = (1.0 / sigma2) * S_q @ A @ (y - mu_x)
# Build a guide carrying that closed-form posterior.
L_q = jnp.linalg.cholesky(0.5 * (S_q + S_q.T) + 1e-9 * jnp.eye(M))
guide = eqx.tree_at(
lambda g: (g.mean, g.scale_tril),
guide,
replace=(m_q, L_q),
)
print(
f"closed-form ELBO = {float(sparse_markov_elbo(prior, guide, lik, times, y)):.3f}"
)closed-form ELBO = -101.641
Posterior at the fitted guide¶
SparseConditionedMarkovGP.predict(t_star) produces the mean / variance at arbitrary test times via the standard SVGP predictive (cost ).
cond = SparseConditionedMarkovGP(prior=prior, guide=guide)
t_star = jnp.linspace(times[0] - 0.3, times[-1] + 1.0, 300)
m_star, v_star = cond.predict(t_star)
m_star, v_star = np.asarray(m_star), np.asarray(v_star)
sd_star = np.sqrt(np.maximum(v_star, 0))
fig, ax = plt.subplots(figsize=(8.5, 3.6))
ax.fill_between(
t_star,
m_star - 2 * sd_star,
m_star + 2 * sd_star,
color="C0",
alpha=0.25,
label="posterior ±2σ",
)
ax.plot(t_star, m_star, color="C0", lw=2, label="posterior mean")
ax.plot(times, f_true, "k--", lw=1, label="true f")
ax.scatter(times, y, c="C3", s=14, label="data", zorder=3)
ax.scatter(
Z,
jnp.full_like(Z, ax.get_ylim()[0] + 0.1),
c="C2",
marker="^",
s=40,
label="inducing Z",
)
ax.set_xlabel("t")
ax.set_ylabel("y / f")
ax.set_title("SparseMarkovGP posterior — closed-form q(u)")
ax.legend(loc="lower right", fontsize=8)
plt.tight_layout()
plt.show()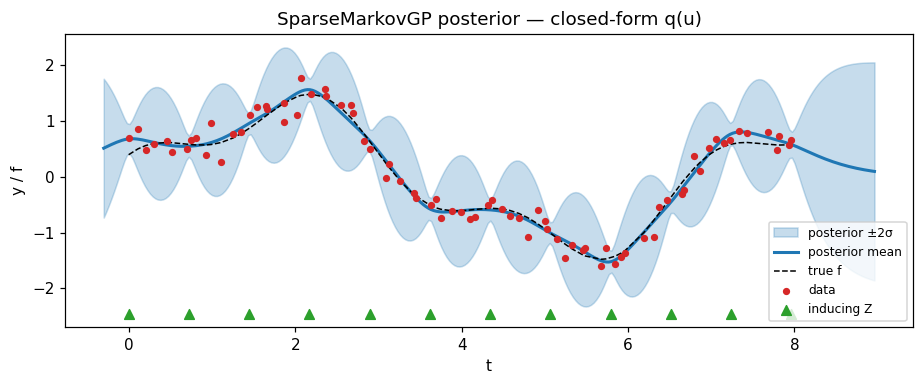
Comparison: sparse vs dense Markov GP¶
At the closed-form and well-spaced inducing points, the SVGP posterior should approximate the dense-Markov posterior. The remaining difference is the inducing-set bias from working with pseudo-points instead of all data points.
dense_prior = MarkovGPPrior(sde, times)
dense_cond = dense_prior.condition(y, jnp.asarray(0.05))
m_dense, v_dense = dense_cond.predict(t_star)
m_dense, v_dense = np.asarray(m_dense), np.asarray(v_dense)
sd_dense = np.sqrt(np.maximum(v_dense, 0))
fig, ax = plt.subplots(figsize=(8.5, 3.6))
ax.fill_between(
t_star,
m_dense - 2 * sd_dense,
m_dense + 2 * sd_dense,
color="C2",
alpha=0.15,
label="dense ±2σ",
)
ax.plot(t_star, m_dense, color="C2", lw=2, label="dense Markov mean")
ax.fill_between(
t_star,
m_star - 2 * sd_star,
m_star + 2 * sd_star,
color="C0",
alpha=0.15,
label="sparse ±2σ",
)
ax.plot(t_star, m_star, color="C0", lw=2, ls="--", label=f"sparse mean (M={M})")
ax.scatter(times, y, c="C3", s=10, alpha=0.5)
ax.set_xlabel("t")
ax.set_ylabel("f")
ax.set_title("Dense vs sparse Markov GP — same SDE prior, same noise")
ax.legend(loc="lower right", fontsize=8)
plt.tight_layout()
plt.show()
print(
f"max |sparse mean − dense mean| = {float(jnp.max(jnp.abs(m_star - m_dense))):.4f}"
)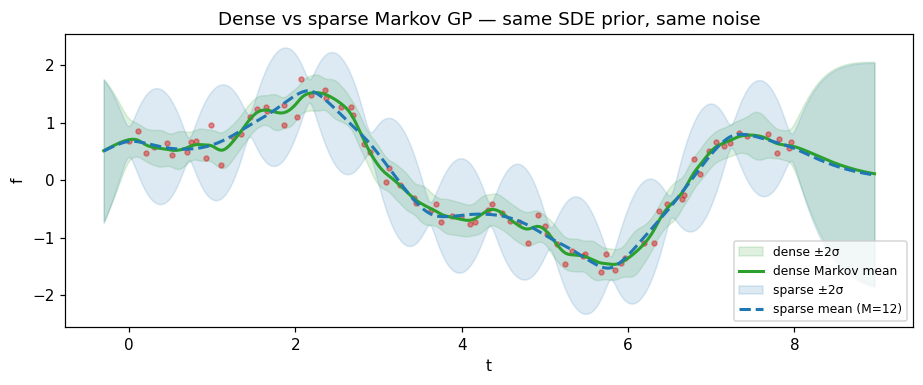
max |sparse mean − dense mean| = 0.2002
Sweep: how many inducing points do we need?¶
Increase M and re-fit the closed-form . Past a kernel-dependent threshold, the sparse posterior is indistinguishable from the dense one.
def fit_sparse(M_: int) -> SparseConditionedMarkovGP:
Z_ = jnp.linspace(times[0], times[-1], M_)
prior_ = SparseMarkovGPPrior(sde, Z_, jitter=1e-6)
K_zz_op_, K_xz_, _ = prior_.predictive_blocks(times)
K_zz_ = K_zz_op_.as_matrix()
A_ = jnp.linalg.solve(K_zz_, K_xz_.T)
Lambda_ = jnp.linalg.inv(K_zz_) + (1.0 / sigma2) * A_ @ A_.T
S_ = jnp.linalg.inv(Lambda_)
m_ = (1.0 / sigma2) * S_ @ A_ @ (y - prior_.mean(times))
L_ = jnp.linalg.cholesky(0.5 * (S_ + S_.T) + 1e-9 * jnp.eye(M_))
g_ = eqx.tree_at(
lambda g: (g.mean, g.scale_tril),
FullRankGuide.init(M_, scale=0.3),
replace=(m_, L_),
)
return SparseConditionedMarkovGP(prior=prior_, guide=g_)
fig, ax = plt.subplots(figsize=(8.5, 3.6))
ax.plot(t_star, m_dense, color="black", lw=2, label="dense (reference)")
for M_, color in zip([4, 8, 16, 32], ["C0", "C1", "C2", "C3"], strict=True):
cond_ = fit_sparse(M_)
m_, _ = cond_.predict(t_star)
ax.plot(t_star, np.asarray(m_), color=color, lw=1.5, ls="--", label=f"M={M_}")
ax.scatter(times, y, c="grey", s=8, alpha=0.4)
ax.set_xlabel("t")
ax.set_ylabel("posterior mean")
ax.set_title("Sparse-vs-dense posterior — sweep over inducing-point count")
ax.legend(loc="lower right", fontsize=8)
plt.tight_layout()
plt.show()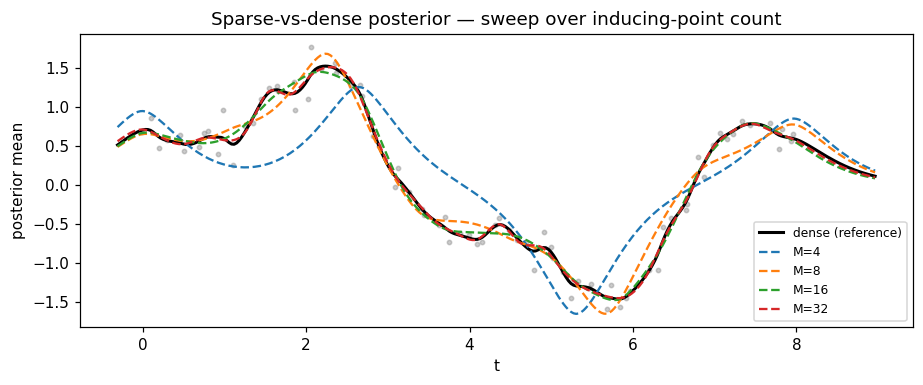
Summary¶
SparseMarkovGPPrior(sde_kernel, Z)exposes the SVGP predictive blocks for any state-space kernel.sparse_markov_elbo(...)plugs into the existing variational-guide infrastructure with a closed-form Gaussian-likelihood path and a numerical (gaussx-integrator) path for non-conjugate likelihoods.- For Gaussian observations the optimal is closed-form (used here for demonstration); for non-conjugate likelihoods, optimise
sparse_markov_elbovia standard SVI (seemarkov_gp_training.ipynbfor the optax-LBFGS pattern wrapping the SDE forward pass under JIT, which is the canonical way to train through the dynamic-stop expm inSDEKernel.discretise). - At large enough to span the posterior support of the kernel, the sparse posterior collapses to the dense one. Reducing trades fidelity for the ELBO cost.
- This surface is the SVGP-style foundation. A truly linear-in- Kalman-aware ELBO that exploits the Markov factorisation of is a follow-up.