Kalman Filter and RTS Smoother
This notebook demonstrates the gaussx Kalman filter and Rauch-Tung-Striebel (RTS) smoother recipes on a simple linear dynamical system.
What you’ll learn:
- Setting up a linear-Gaussian state-space model
- Running
gaussx.kalman_filterfor online state estimation - Running
gaussx.rts_smootherfor offline smoothing - Comparing filtered vs smoothed estimates
- The filter is fully differentiable via JAX
Background¶
A linear-Gaussian state-space model (SSM) is defined by:
The Kalman filter computes the filtering distribution in a single forward pass. The RTS smoother then refines these estimates using future observations, giving the smoothing distribution .
The smoother always has lower (or equal) posterior variance than the filter, because it conditions on strictly more data.
Kalman filter equations¶
The filter alternates between a predict step and an update step:
Predict:
Update:
Note that each step involves a linear system solve (for ), which is where gaussx primitives are used.
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import jax
import jax.numpy as jnp
import matplotlib.pyplot as plt
import gaussx
jax.config.update("jax_enable_x64", True)Define the model¶
We use a 2D state with a damped oscillator transition (discretized spring-mass-damper). Only position is observed with noise.
- : discretized spring-mass-damper transition
- (observe position only)
- : process noise driving the oscillator
- : observation noise variance
dt = 0.1
T = 200
# Damped oscillator: omega=1.0 rad/s, damping gamma=0.15
omega, gamma = 1.0, 0.15
A = jnp.array([[1.0, dt], [-(omega**2) * dt, 1.0 - gamma * dt]])
# Observation matrix (observe position only)
H = jnp.array([[1.0, 0.0]])
# Process noise covariance
q_var = 0.3
Q = q_var * jnp.array([[dt**3 / 3, dt**2 / 2], [dt**2 / 2, dt]])
# Observation noise covariance
R = jnp.array([[0.5]])
print("A =\n", A)
print("H =", H)
print("Q =\n", Q)
print("R =", R)A =
[[ 1. 0.1 ]
[-0.1 0.985]]
H = [[1. 0.]]
Q =
[[0.0001 0.0015]
[0.0015 0.03 ]]
R = [[0.5]]
Generate data¶
We simulate the true trajectory from the model itself, so the Kalman filter’s assumptions are satisfied. This shows the filter performing as designed — tracking a randomly evolving state from noisy observations.
key = jax.random.PRNGKey(42)
# Simulate states from the model
def simulate_step(carry, key_t):
x = carry
k1, k2 = jax.random.split(key_t)
q_t = jax.random.multivariate_normal(k1, jnp.zeros(2), Q)
x_new = A @ x + q_t
r_t = jax.random.multivariate_normal(k2, jnp.zeros(1), R)
y_t = H @ x_new + r_t
return x_new, (x_new, y_t)
x0 = jnp.array([3.0, 0.0]) # displaced from equilibrium, at rest
keys = jax.random.split(key, T)
_, (true_states, observations) = jax.lax.scan(simulate_step, x0, keys)
times = jnp.arange(T) * dt
true_position = true_states[:, 0]
print("true_states shape:", true_states.shape)
print("observations shape:", observations.shape)true_states shape: (200, 2)
observations shape: (200, 1)
Run Kalman filter¶
gaussx.kalman_filter takes the model matrices and observations,
returning a FilterState with filtered means, covariances, and the
total log-likelihood.
# Initial state: zero mean, moderate uncertainty
init_mean = jnp.zeros(2)
init_cov = jnp.eye(2) * 4.0
filter_state = gaussx.kalman_filter(A, H, Q, R, observations, init_mean, init_cov)
print("Filtered means shape:", filter_state.filtered_means.shape)
print("Filtered covs shape:", filter_state.filtered_covs.shape)
print("Log-likelihood:", filter_state.log_likelihood)Filtered means shape: (200, 2)
Filtered covs shape: (200, 2, 2)
Log-likelihood: -223.3188576581507
Plot filtered results¶
The filtered estimate tracks the true position closely despite the noisy observations. The shaded band shows the credible interval.
filt_pos_mean = filter_state.filtered_means[:, 0]
filt_pos_std = jnp.sqrt(filter_state.filtered_covs[:, 0, 0])
filt_vel_mean = filter_state.filtered_means[:, 1]
filt_vel_std = jnp.sqrt(filter_state.filtered_covs[:, 1, 1])
fig, axes = plt.subplots(2, 1, figsize=(12, 6), sharex=True)
# Position
ax = axes[0]
ax.scatter(
times,
observations[:, 0],
s=30,
c="C0",
alpha=0.35,
edgecolors="k",
linewidths=0.5,
label="Observations",
zorder=5,
)
ax.plot(times, true_position, "k--", lw=1.5, label="True position", zorder=4)
ax.plot(times, filt_pos_mean, "C1-", lw=2, label="Filtered mean", zorder=3)
ax.fill_between(
times,
filt_pos_mean - 2 * filt_pos_std,
filt_pos_mean + 2 * filt_pos_std,
color="C1",
alpha=0.2,
label=r"$\pm 2\sigma$",
)
ax.set_ylabel("Position")
ax.set_title("Kalman Filter: Filtered Estimates")
ax.legend(loc="upper right", fontsize=9)
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
# Velocity (inferred, never directly observed)
ax = axes[1]
ax.plot(times, true_states[:, 1], "k--", lw=1.5, label="True velocity", zorder=4)
ax.plot(times, filt_vel_mean, "C3-", lw=2, label="Filtered mean", zorder=3)
ax.fill_between(
times,
filt_vel_mean - 2 * filt_vel_std,
filt_vel_mean + 2 * filt_vel_std,
color="C3",
alpha=0.2,
label=r"$\pm 2\sigma$",
)
ax.set_xlabel("Time")
ax.set_ylabel("Velocity (latent)")
ax.legend(loc="upper right", fontsize=9)
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.tight_layout()
plt.show()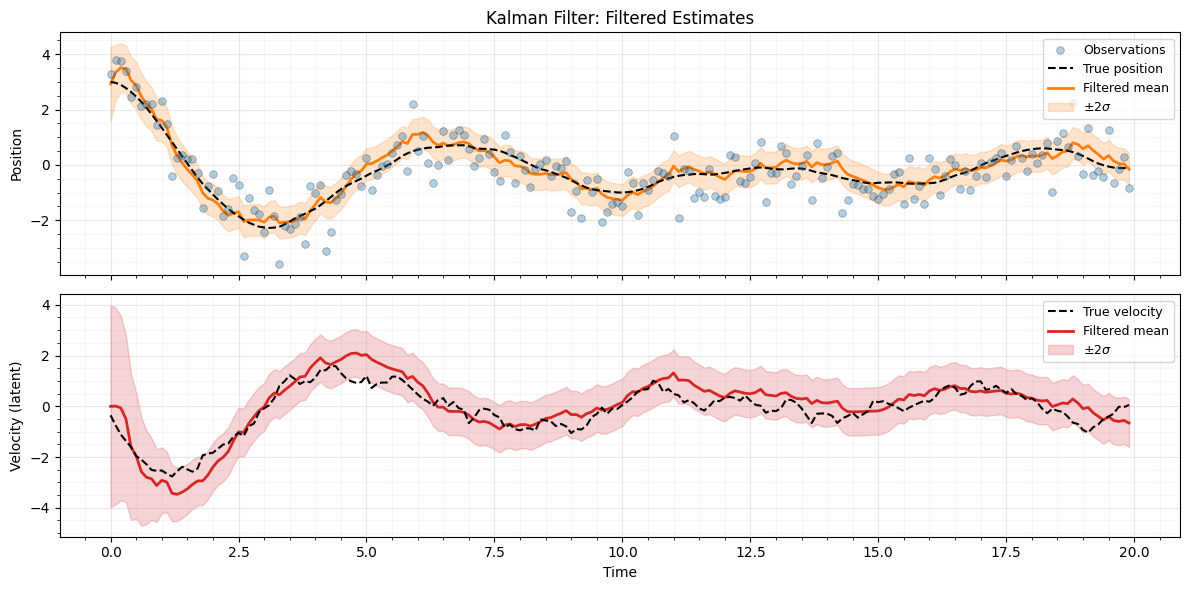
Run RTS smoother¶
The RTS smoother takes the filter output and refines the estimates using a backward pass. This produces tighter credible intervals, especially in the middle of the time series.
The smoother computes a backward recursion from down to :
The smoother refines filter estimates by incorporating future observations. This optimal smoother was derived by Rauch, Tung, & Striebel (1965).
smoothed_means, smoothed_covs = gaussx.rts_smoother(filter_state, A, Q)
print("Smoothed means shape:", smoothed_means.shape)
print("Smoothed covs shape:", smoothed_covs.shape)Smoothed means shape: (200, 2)
Smoothed covs shape: (200, 2, 2)
Plot filtered vs smoothed¶
The smoother uncertainty is everywhere less than or equal to the filter uncertainty, because it conditions on all observations .
smooth_pos_mean = smoothed_means[:, 0]
smooth_pos_std = jnp.sqrt(smoothed_covs[:, 0, 0])
smooth_vel_mean = smoothed_means[:, 1]
smooth_vel_std = jnp.sqrt(smoothed_covs[:, 1, 1])
fig, axes = plt.subplots(2, 1, figsize=(12, 7), sharex=True)
# Top: position — overlay filtered and smoothed
ax = axes[0]
ax.scatter(
times,
observations[:, 0],
s=30,
c="C0",
alpha=0.25,
edgecolors="k",
linewidths=0.5,
label="Observations",
zorder=5,
)
ax.plot(times, true_position, "k--", lw=1.5, label="True", zorder=4)
ax.fill_between(
times,
filt_pos_mean - 2 * filt_pos_std,
filt_pos_mean + 2 * filt_pos_std,
color="C1",
alpha=0.15,
label=r"Filtered $\pm 2\sigma$",
)
ax.plot(times, filt_pos_mean, "C1-", lw=1, alpha=0.6, label="Filtered", zorder=3)
ax.fill_between(
times,
smooth_pos_mean - 2 * smooth_pos_std,
smooth_pos_mean + 2 * smooth_pos_std,
color="C2",
alpha=0.25,
label=r"Smoothed $\pm 2\sigma$",
)
ax.plot(times, smooth_pos_mean, "C2-", lw=2, label="Smoothed", zorder=3)
ax.set_ylabel("Position")
ax.set_title("Filter vs Smoother: Position")
ax.legend(loc="upper right", fontsize=9, ncol=2)
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
# Bottom: velocity — overlay filtered and smoothed
ax = axes[1]
ax.plot(times, true_states[:, 1], "k--", lw=1.5, label="True", zorder=4)
ax.fill_between(
times,
filt_vel_mean - 2 * filt_vel_std,
filt_vel_mean + 2 * filt_vel_std,
color="C1",
alpha=0.15,
label=r"Filtered $\pm 2\sigma$",
)
ax.plot(times, filt_vel_mean, "C1-", lw=1, alpha=0.6, label="Filtered", zorder=3)
ax.fill_between(
times,
smooth_vel_mean - 2 * smooth_vel_std,
smooth_vel_mean + 2 * smooth_vel_std,
color="C3",
alpha=0.25,
label=r"Smoothed $\pm 2\sigma$",
)
ax.plot(times, smooth_vel_mean, "C3-", lw=2, label="Smoothed", zorder=3)
ax.set_xlabel("Time")
ax.set_ylabel("Velocity (latent)")
ax.set_title("Filter vs Smoother: Velocity")
ax.legend(loc="upper right", fontsize=9, ncol=2)
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.tight_layout()
plt.show()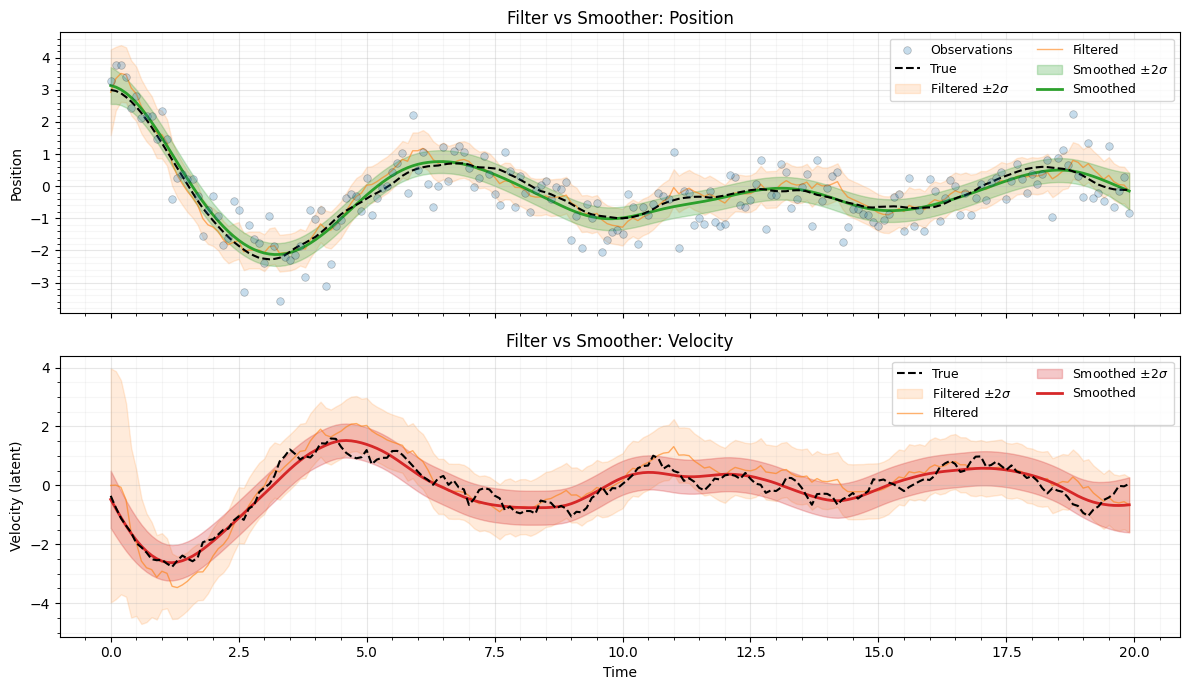
Differentiability¶
The Kalman filter is implemented via jax.lax.scan, so it is fully
differentiable. We can compute gradients of the log-likelihood with
respect to model parameters — useful for learning SSM parameters
via gradient-based optimization.
def neg_log_likelihood(log_obs_noise_var):
"""Negative log-likelihood as a function of log observation noise variance."""
R_param = jnp.exp(log_obs_noise_var) * jnp.eye(1)
fs = gaussx.kalman_filter(A, H, Q, R_param, observations, init_mean, init_cov)
return -fs.log_likelihood
# Evaluate at the true value
log_R_true = jnp.log(R[0, 0])
nll = neg_log_likelihood(log_R_true)
grad_nll = jax.grad(neg_log_likelihood)(log_R_true)
print(f"log(R) = {log_R_true:.4f}")
print(f"Negative log-likelihood = {nll:.4f}")
print(f"Gradient d(-LL)/d(log R) = {grad_nll:.4f}")log(R) = -0.6931
Negative log-likelihood = 223.3189
Gradient d(-LL)/d(log R) = 12.2965
Summary¶
gaussx.kalman_filterimplements the standard Kalman filter forward pass, returning filtered means, covariances, and the total log-likelihood.gaussx.rts_smootherrefines filtered estimates using a backward pass, producing tighter credible intervals.- Both are implemented with
jax.lax.scanand are fully compatible with JAX transforms:jit,vmap, andgradall work out of the box. - This makes gaussx suitable for learning SSM parameters via gradient-based optimization of the log-likelihood.
References¶
- Kalman, R. E. (1960). A new approach to linear filtering and prediction problems. J. Basic Engineering, 82(1), 35--45.
- Rauch, H. E., Tung, F., & Striebel, C. T. (1965). Maximum likelihood estimates of linear dynamic systems. AIAA Journal, 3(8), 1445--1450.
- Sarkka, S. (2013). Bayesian Filtering and Smoothing. Cambridge University Press.
- Anderson, B. D. O. & Moore, J. B. (1979). Optimal Filtering. Prentice-Hall.