Sparse Variational GP with Inducing Points
Sparse GPs approximate the full kernel with inducing points:
The resulting covariance is — a LowRankUpdate
in gaussx. We optimize the ELBO with respect to inducing point
locations and kernel hyperparameters via jax.grad.
Background¶
Sparse GP methods address the cost of exact GP inference by introducing inducing points . The key idea is to approximate the full GP posterior via a variational distribution that conditions on the inducing variables . The Nystrom approximation gives:
The variational free energy (VFE) framework (Titsias, 2009) optimizes a lower bound on the log-marginal likelihood:
The trace correction penalizes the approximation error, and the bound is tight when the inducing points explain all the data.
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import jax
import jax.numpy as jnp
import matplotlib.pyplot as plt
import gaussx
jax.config.update("jax_enable_x64", True)Generate data¶
key = jax.random.PRNGKey(123)
k1, k2 = jax.random.split(key)
n_data = 300
noise_std = 0.2
f_true = lambda x: jnp.sin(2 * x) + 0.3 * jnp.cos(5 * x)
x_data = jnp.sort(jax.random.uniform(k1, (n_data,), minval=-4.0, maxval=4.0))
y_data = f_true(x_data) + noise_std * jax.random.normal(k2, (n_data,))
x_plot = jnp.linspace(-4.5, 4.5, 500)Build Nystrom approximation¶
The Nystrom factor gives
, so the noisy covariance
is a rank- update of a diagonal -- exactly a LowRankUpdate.
This is why gaussx can apply the Woodbury identity for
solve and the matrix determinant lemma for
logdet.
n_inducing = 12
def rbf(x1, x2, ls, var):
return var * jnp.exp(-0.5 * (x1[:, None] - x2[None, :]) ** 2 / ls**2)
def build_sparse_gp(x_data, x_inducing, ls, var, noise):
"""Build the sparse GP covariance as a LowRankUpdate."""
K_nm = rbf(x_data, x_inducing, ls, var)
K_mm = rbf(x_inducing, x_inducing, ls, var)
# Nystrom: U = K_nm @ chol(K_mm)^{-T}
jitter = 1e-5 * jnp.eye(len(x_inducing))
L_mm = jnp.linalg.cholesky(K_mm + jitter)
U = jax.scipy.linalg.solve_triangular(L_mm, K_nm.T, lower=True).T
# Sigma = noise * I + U U^T
return gaussx.low_rank_plus_diag(noise * jnp.ones(len(x_data)), U)
# Initial setup
x_inducing = jnp.linspace(-3.5, 3.5, n_inducing)
ls, var, noise = 1.0, 1.0, noise_std**2
sigma = build_sparse_gp(x_data, x_inducing, ls, var, noise)
print(f"Data: {n_data}, Inducing: {n_inducing}")
print(f"Operator rank: {sigma.rank}")Data: 300, Inducing: 12
Operator rank: 12
ELBO (variational lower bound)¶
The first term uses gaussx solve + logdet. The trace correction penalizes information lost by the approximation.
def elbo(log_params, x_data, y_data, x_inducing):
ls = jnp.exp(log_params[0])
var = jnp.exp(log_params[1])
noise = jnp.exp(log_params[2])
sigma_op = build_sparse_gp(x_data, x_inducing, ls, var, noise)
# Log-likelihood term
alpha = gaussx.solve(sigma_op, y_data)
data_fit = -0.5 * jnp.dot(y_data, alpha)
complexity = -0.5 * gaussx.logdet(sigma_op)
const = -0.5 * len(y_data) * jnp.log(2 * jnp.pi)
# Trace correction: tr(K_nn - Q_nn) / (2 * noise)
# K_nn diagonal = var (for RBF with same point)
# Q_nn diagonal = diag(U U^T)
Q_diag = jnp.sum(sigma_op.U**2, axis=1)
trace_correction = -0.5 * jnp.sum(var - Q_diag) / noise
return data_fit + complexity + const + trace_correction
log_params = jnp.log(jnp.array([ls, var, noise]))
print(f"Initial ELBO: {elbo(log_params, x_data, y_data, x_inducing):.2f}")Initial ELBO: -165.14
Optimize hyperparameters¶
grad_fn = jax.jit(jax.grad(lambda p: -elbo(p, x_data, y_data, x_inducing)))
lr = 0.01
for _i in range(100):
g = grad_fn(log_params)
# Clip gradients for stability
g = jnp.clip(g, -10.0, 10.0)
log_params = log_params - lr * g
ls_opt, var_opt, noise_opt = jnp.exp(log_params)
print(f"Optimized: ls={ls_opt:.3f}, var={var_opt:.3f}, noise={noise_opt:.4f}")
print(f"Final ELBO: {elbo(log_params, x_data, y_data, x_inducing):.2f}")Optimized: ls=1.020, var=1.504, noise=0.0922
Final ELBO: -96.68
Predict¶
sigma_opt = build_sparse_gp(x_data, x_inducing, ls_opt, var_opt, noise_opt)
alpha_opt = gaussx.solve(sigma_opt, y_data)
# Prediction via inducing points
K_star_m = rbf(x_plot, x_inducing, ls_opt, var_opt)
K_mm = rbf(x_inducing, x_inducing, ls_opt, var_opt)
K_nm = rbf(x_data, x_inducing, ls_opt, var_opt)
weights = jnp.linalg.solve(
K_mm + 1e-5 * jnp.eye(n_inducing),
K_nm.T @ alpha_opt,
)
y_pred = K_star_m @ weightsVisualize¶
fig, ax = plt.subplots(figsize=(10, 4))
ax.plot(x_plot, f_true(x_plot), "k--", lw=1.5, label="True function", zorder=4)
ax.scatter(
x_data,
y_data,
s=30,
c="C0",
alpha=0.3,
edgecolors="k",
linewidths=0.5,
label="Data",
zorder=5,
)
ax.plot(x_plot, y_pred, "C1-", lw=2, label="Sparse GP mean", zorder=3)
ax.scatter(
x_inducing,
jnp.full_like(x_inducing, -1.8),
marker="^",
s=50,
c="C2",
zorder=5,
label=f"Inducing pts ({n_inducing})",
)
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.set_title(
f"Sparse Variational GP: {n_data} data, {n_inducing} inducing "
f"(ls={ls_opt:.2f}, var={var_opt:.2f})"
)
ax.legend(fontsize=9)
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.tight_layout()
plt.show()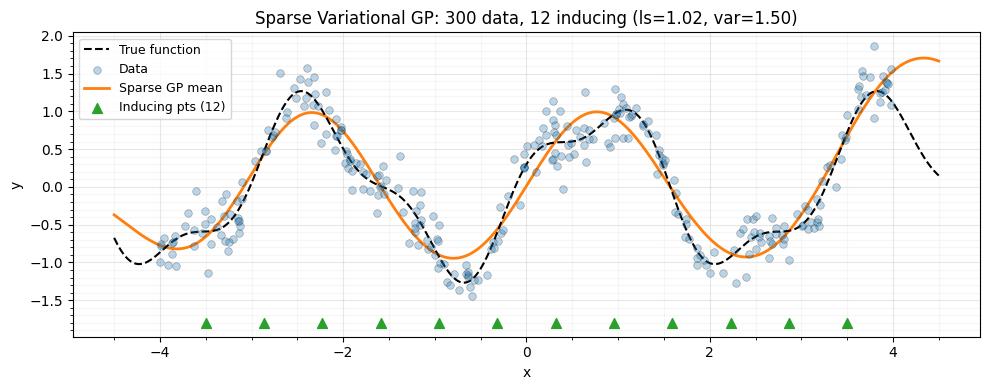
Summary¶
| Component | gaussx role |
|---|---|
low_rank_plus_diag | Build Nystrom covariance |
gaussx.solve | Woodbury solve for weights |
gaussx.logdet | Matrix determinant lemma for ELBO |
jax.grad | Differentiate through everything |
References¶
- Quinonero-Candela, J. & Rasmussen, C. E. (2005). A unifying view of sparse approximate Gaussian process regression. JMLR, 6, 1939--1959.
- Titsias, M. (2009). Variational learning of inducing variables in sparse Gaussian processes. Proc. AISTATS.
- Hensman, J., Fusi, N., & Lawrence, N. D. (2013). Gaussian processes for big data. Proc. UAI.