Random Fourier Features → SSGP → VSSGP — Three Spectral Views of a Gaussian Process
This notebook is the GP-flavored companion to the RFF as Neural Networks notebook. Where that notebook treats RFF as a neural network architecture (with frozen, learned, or ensembled weights), here we follow the Bayesian progression that links the same feature map to Gaussian processes:
- Learned RFF (MAP baseline) — the deterministic regression model with trainable Ω. No uncertainty over the parameters; just a regularised MSE point estimate.
- SSGP — Sparse Spectrum GP (Lázaro-Gredilla et al. JMLR 2010) — analytically marginalise the head β and train Ω on the GP marginal likelihood. Closed-form predictive variance for free.
- VSSGP — Variational SSGP (Gal & Turner ICML 2015) — put a variational posterior on top of , train via reparameterised ELBO. Full posterior uncertainty over both frequencies and weights.
The three are different inferential commitments to the same model class — a linear combination of random Fourier features:
| Method | Ω | β | Objective | Predictive variance? |
|---|---|---|---|---|
| Learned RFF (MAP) | trained (point) | trained (point) + L2 | regularised MSE | no |
| SSGP | trained (point) | marginalised analytically | log marginal likelihood | yes — closed form |
| VSSGP | variational | variational | tempered ELBO | yes — MC over the posterior |
Foundation. The Bochner / Rahimi-Recht derivation of the feature map, the kernel-approximation convergence rate, and the equivalence between paired and phased readouts across RBF / Matérn / Laplace are covered separately in the Kernel Approximation notebook. This notebook treats that map as a black box and focuses on the Bayesian inference layer.
Forward-pointers: the fixed-Ω version (Rahimi-Recht 2007) and the ensemble-of-MAP alternative live in the RFF as Neural Networks notebook. The deep versions are in the Deep Random Feature Expansions notebook.
Background — from features to GPs¶
A linear combination of random Fourier features with Gaussian weights
is itself a Gaussian process with kernel
This is the Sparse Spectrum GP (SSGP). The marginal likelihood and predictive distribution have closed forms — derived in §3. Per-iteration cost is , linear in the dataset size for fixed , the headline win over the exact GP. (For the spectral-density derivation that justifies — and the empirical convergence figures across RBF / Matérn / Laplace — see the Kernel Approximation notebook.)
Setup¶
Detect Colab and install pyrox[colab] (which pulls in matplotlib and watermark) only when running there.
import subprocess
import sys
try:
import google.colab # noqa: F401
IN_COLAB = True
except ImportError:
IN_COLAB = False
if IN_COLAB:
subprocess.run(
[
sys.executable,
"-m",
"pip",
"install",
"-q",
"pyrox[colab] @ git+https://github.com/jejjohnson/pyrox@main",
],
check=True,
)import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import equinox as eqx
import jax
import jax.numpy as jnp
import jax.random as jr
import matplotlib.pyplot as plt
import numpy as np
import optax
jax.config.update("jax_enable_x64", True)import importlib.util
try:
from IPython import get_ipython
ipython = get_ipython()
except ImportError:
ipython = None
if ipython is not None and importlib.util.find_spec("watermark") is not None:
ipython.run_line_magic("load_ext", "watermark")
ipython.run_line_magic(
"watermark",
"-v -m -p jax,equinox,numpyro,pyrox,matplotlib",
)
else:
print("watermark extension not installed; skipping reproducibility readout.")Python implementation: CPython
Python version : 3.13.5
IPython version : 9.10.0
jax : 0.9.2
equinox : 0.13.6
numpyro : 0.20.1
pyrox : 0.0.7
matplotlib: 3.10.8
Compiler : GCC 11.2.0
OS : Linux
Release : 6.8.0-1044-azure
Machine : x86_64
Processor : x86_64
CPU cores : 16
Architecture: 64bit
Shared regression setup¶
All three regression methods (§2 learned RFF, §3 SSGP, §4 VSSGP) are evaluated on the same target with a held-out gap, so predictive uncertainty in the gap becomes the visual axis of comparison:
The target frequency is . We use a lengthscale prior throughout, giving prior frequency standard deviation — comfortable coverage of . All three methods use Fourier features.
N_OBS, NOISE_STD = 80, 0.05
x_full = jnp.linspace(-1.0, 1.0, N_OBS)
mask = (x_full < -0.2) | (x_full > 0.4)
x_obs = x_full[mask].reshape(-1, 1)
y_obs = jnp.sin(3.0 * jnp.pi * x_obs[:, 0]) + NOISE_STD * jr.normal(
jr.PRNGKey(2), x_obs.shape[0:1]
)
x_test = jnp.linspace(-1.2, 1.2, 400).reshape(-1, 1)
y_truth = jnp.sin(3.0 * jnp.pi * x_test[:, 0])
M_FEAT = 64
LENGTHSCALE_INIT = 0.3
TARGET_OMEGA = 3.0 * jnp.pi
print(f"target frequency: ω⋆ = 3π ≈ {float(TARGET_OMEGA):.2f}")
print(f"prior bandwidth at ℓ={LENGTHSCALE_INIT}: 1/ℓ ≈ {1.0 / LENGTHSCALE_INIT:.2f}")
print(f"observation count: N = {x_obs.shape[0]}, gap removed from x ∈ (-0.2, 0.4)")target frequency: ω⋆ = 3π ≈ 9.42
prior bandwidth at ℓ=0.3: 1/ℓ ≈ 3.33
observation count: N = 56, gap removed from x ∈ (-0.2, 0.4)
2. Learned RFF — the MAP baseline¶
Math. The simplest regression model in the spectral family. Fix nothing as random; treat Ω, , β, and an intercept as PyTree leaves and minimise the regularised MSE
This is the MAP point estimate of the SSGP model below — the same likelihood, but with the head β collapsed to a single value rather than a posterior. Equivalent to “neural-network RFF” in the companion notebook.
Wide initialisation. The activation has gradient , which is zero at . Initialising traps gradient descent at this saddle for high-frequency targets. We initialise with (wide enough that already spans a non-trivial phase range, narrow enough that the prior at still concentrates near the target).
What MAP cannot give you. A point estimate. No uncertainty. The fit through the held-out gap is whatever the optimiser settles on, with no signal that the model “doesn’t know” there. That gap-uncertainty is precisely what SSGP and VSSGP add.
class LearnedRFF(eqx.Module):
"""Two-layer NN with [cos, sin] activations — all parameters trainable.
The MAP point estimate of the SSGP model: same likelihood, but
instead of marginalising the head ``beta`` we collapse it to a
single optimised value.
"""
W: jax.Array # (D, M) — trainable spectral frequencies
log_ell: jax.Array # () — trainable lengthscale (positive via exp)
beta: jax.Array # (2M,) — trainable linear head
bias: jax.Array # () — trainable scalar bias
@classmethod
def init(cls, key, in_features, n_features, lengthscale, *, w_init_scale=5.0):
kW, kb = jr.split(key)
return cls(
W=w_init_scale * jr.normal(kW, (in_features, n_features)),
log_ell=jnp.log(jnp.array(lengthscale)),
beta=0.01 * jr.normal(kb, (2 * n_features,)),
bias=jnp.zeros(()),
)
def __call__(self, x):
ell = jnp.exp(self.log_ell)
z = x @ self.W / ell
scale = jnp.sqrt(1.0 / self.W.shape[-1])
phi = scale * jnp.concatenate([jnp.cos(z), jnp.sin(z)], axis=-1)
return phi @ self.beta + self.bias
def fit_map(model, x_obs, y_obs, *, n_steps=4000, lr=1e-2, beta_l2=1e-3):
opt = optax.adam(lr)
state = opt.init(eqx.filter(model, eqx.is_inexact_array))
def loss_fn(m):
pred = m(x_obs)
mse = jnp.mean((pred - y_obs) ** 2)
return mse + beta_l2 * jnp.sum(m.beta**2)
@eqx.filter_jit
def step(m, s):
loss, grads = eqx.filter_value_and_grad(loss_fn)(m)
upd, s = opt.update(grads, s, eqx.filter(m, eqx.is_inexact_array))
return eqx.apply_updates(m, upd), s, loss
losses = []
for _ in range(n_steps):
model, state, loss = step(model, state)
losses.append(float(loss))
return model, losses
map_model = LearnedRFF.init(
jr.PRNGKey(7), in_features=1, n_features=M_FEAT, lengthscale=LENGTHSCALE_INIT
)
map_model, map_losses = fit_map(map_model, x_obs, y_obs)
y_pred_map = map_model(x_test)
mse_map = float(jnp.mean((y_pred_map - y_truth) ** 2))
print(
f"learned RFF MAP: final loss = {map_losses[-1]:.4f}, "
f"learned ℓ = {float(jnp.exp(map_model.log_ell)):.3f}, MSE = {mse_map:.4f}"
)learned RFF MAP: final loss = 0.0043, learned ℓ = 0.313, MSE = 0.1320
fig, ax = plt.subplots(figsize=(11, 4.5))
ax.scatter(
x_obs[:, 0],
y_obs,
s=10,
color="C1",
edgecolors="k",
linewidths=0.5,
label="observations",
zorder=5,
)
ax.plot(x_test[:, 0], y_truth, "k--", linewidth=1.5, label="truth")
ax.plot(
x_test[:, 0],
y_pred_map,
"C3",
linewidth=1.8,
label=f"learned RFF MAP (MSE={mse_map:.4f})",
)
ax.axvspan(-0.2, 0.4, color="0.85", alpha=0.4, label="held-out gap")
ax.set_xlabel("$x$")
ax.set_ylabel("$y$")
ax.set_title("Learned RFF MAP — deterministic fit, no uncertainty in the gap")
ax.grid(True, alpha=0.3)
ax.legend(loc="upper right", fontsize=9)
plt.tight_layout()
plt.show()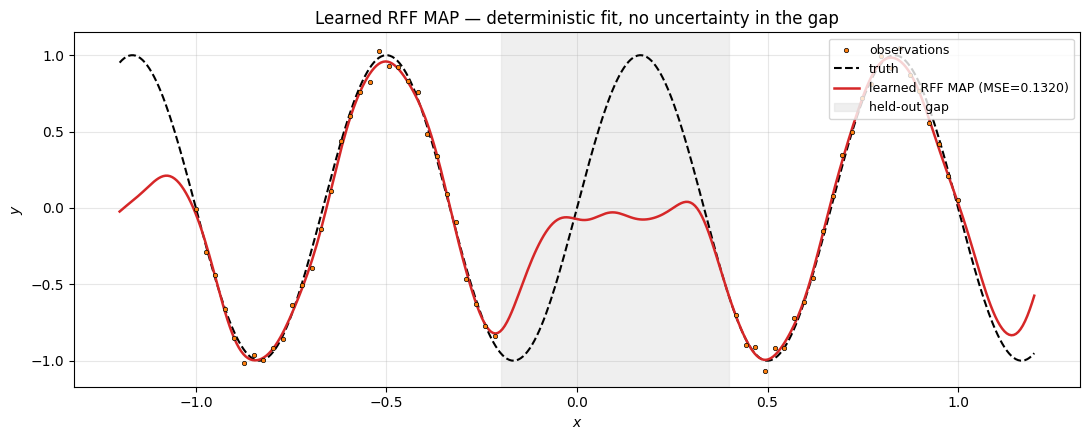
Clean fit at the data, but the curve through the held-out gap is just whatever the optimiser settled on — there is no way to read off “the model is uncertain here.” Compare this to the next two sections, where the uncertainty band visibly opens across the gap.
3. SSGP — Sparse Spectrum GP (Lázaro-Gredilla et al. 2010)¶
Math. Same model as §2 but treat the head β as latent with prior and integrate it out. The marginal distribution of is Gaussian:
This is the GP marginal likelihood with a degenerate rank- kernel. Maximise it with respect to .
Numerically stable form via the matrix-inversion lemma. Naively inverting is ; using collapses it to . Define , then
Combining,
where is the Cholesky factor.
Predictive distribution. The closed-form posterior on β is with and . The predictive distribution at a test point is
Why this beats MAP. The marginal likelihood enforces an automatic Occam’s razor through the term — placing extra features near unnecessary frequencies inflates and is penalised. And the closed-form predictive variance gives uncertainty for free without any MC sampling.
class SSGP(eqx.Module):
"""Sparse Spectrum GP — point-estimate Ω trained on the marginal likelihood.
Hyperparameters: spectral frequencies ``W``, log-lengthscale, log-noise,
log-signal-amplitude. The head ``β`` is *not* a parameter — it is
marginalised analytically.
"""
W: jax.Array
log_ell: jax.Array
log_sigma_n: jax.Array
log_sigma_beta: jax.Array
@classmethod
def init(
cls,
key,
in_features,
n_features,
lengthscale,
*,
w_init_scale=5.0,
sigma_n=0.05,
sigma_beta=1.0,
):
return cls(
W=w_init_scale * jr.normal(key, (in_features, n_features)),
log_ell=jnp.log(jnp.array(lengthscale)),
log_sigma_n=jnp.log(jnp.array(sigma_n)),
log_sigma_beta=jnp.log(jnp.array(sigma_beta)),
)
def features(self, x):
ell = jnp.exp(self.log_ell)
z = x @ self.W / ell
scale = jnp.sqrt(1.0 / self.W.shape[-1])
return scale * jnp.concatenate([jnp.cos(z), jnp.sin(z)], axis=-1)
def neg_log_marginal(self, x, y):
"""Negative log marginal likelihood (Lázaro-Gredilla 2010, Eq. 2.16).
Uses the matrix-inversion-lemma form: invert ``B = σ_n² I + σ_β² Φᵀ Φ``
(a 2M × 2M matrix) instead of ``K_y = σ_n² I + σ_β² Φ Φᵀ`` (N × N).
"""
Phi = self.features(x) # (N, 2M)
N = x.shape[0]
twoM = Phi.shape[1]
sigma_n2 = jnp.exp(2.0 * self.log_sigma_n)
sigma_b2 = jnp.exp(2.0 * self.log_sigma_beta)
B = sigma_n2 * jnp.eye(twoM) + sigma_b2 * Phi.T @ Phi
L = jnp.linalg.cholesky(B)
v = Phi.T @ y # (2M,)
L_inv_v = jax.scipy.linalg.solve_triangular(L, v, lower=True)
# y^T K_y^-1 y = ||y||²/σ_n² - (σ_β²/σ_n²) ||L^-1 v||²
quad = -0.5 / sigma_n2 * jnp.sum(y**2) + 0.5 * sigma_b2 / sigma_n2 * jnp.sum(
L_inv_v**2
)
# log|K_y| = (N - 2M) log σ_n² + log|B| = (N-2M) log σ_n² + 2 Σ log diag(L)
log_det = 0.5 * (N - twoM) * jnp.log(sigma_n2) + jnp.sum(jnp.log(jnp.diag(L)))
log_p = quad - log_det - 0.5 * N * jnp.log(2.0 * jnp.pi)
return -log_p
def predict(self, x_train, y_train, x_query):
"""Posterior predictive mean and total variance at ``x_query``."""
Phi = self.features(x_train)
Phi_q = self.features(x_query)
twoM = Phi.shape[1]
sigma_n2 = jnp.exp(2.0 * self.log_sigma_n)
sigma_b2 = jnp.exp(2.0 * self.log_sigma_beta)
B = sigma_n2 * jnp.eye(twoM) + sigma_b2 * Phi.T @ Phi
L = jnp.linalg.cholesky(B)
# μ_β = σ_β² B^-1 Φ^T y
v = Phi.T @ y_train
mu_beta = sigma_b2 * jax.scipy.linalg.cho_solve((L, True), v)
mean = Phi_q @ mu_beta
# Σ_β = σ_β² σ_n² B^-1; var per query = σ_β² σ_n² φ_q^T B^-1 φ_q
# then add observation noise σ_n² for predictive variance.
L_inv_phi_q = jax.scipy.linalg.solve_triangular(
L, Phi_q.T, lower=True
) # (2M, n_query)
var_f = sigma_b2 * sigma_n2 * jnp.sum(L_inv_phi_q**2, axis=0)
var_y = sigma_n2 + var_f
return mean, var_y
def fit_ssgp(model, x_obs, y_obs, *, n_steps=2000, lr=1e-2):
opt = optax.adam(lr)
state = opt.init(eqx.filter(model, eqx.is_inexact_array))
@eqx.filter_jit
def step(m, s):
loss, grads = eqx.filter_value_and_grad(
lambda mm: mm.neg_log_marginal(x_obs, y_obs)
)(m)
upd, s = opt.update(grads, s, eqx.filter(m, eqx.is_inexact_array))
return eqx.apply_updates(m, upd), s, loss
losses = []
for _ in range(n_steps):
model, state, loss = step(model, state)
losses.append(float(loss))
return model, losses
ssgp = SSGP.init(
jr.PRNGKey(11),
in_features=1,
n_features=M_FEAT,
lengthscale=LENGTHSCALE_INIT,
sigma_n=0.05,
sigma_beta=1.0,
)
ssgp, ssgp_losses = fit_ssgp(ssgp, x_obs, y_obs.reshape(-1))
mean_ssgp, var_ssgp = ssgp.predict(x_obs, y_obs.reshape(-1), x_test)
std_ssgp = jnp.sqrt(var_ssgp)
mse_ssgp = float(jnp.mean((mean_ssgp - y_truth) ** 2))
print(f"SSGP: final neg-log-ML = {ssgp_losses[-1]:.2f}")
print(
f" learned ℓ = {float(jnp.exp(ssgp.log_ell)):.4f}, "
f"σ_n = {float(jnp.exp(ssgp.log_sigma_n)):.4f}, "
f"σ_β = {float(jnp.exp(ssgp.log_sigma_beta)):.4f}"
)
print(f" predictive MSE = {mse_ssgp:.4f}")SSGP: final neg-log-ML = -60.13
learned ℓ = 0.3881, σ_n = 0.0464, σ_β = 0.5022
predictive MSE = 0.0809
Plot the predictive mean and band.
fig, ax = plt.subplots(figsize=(11, 4.5))
ax.scatter(
x_obs[:, 0],
y_obs,
s=10,
color="C1",
edgecolors="k",
linewidths=0.5,
label="observations",
zorder=5,
)
ax.plot(x_test[:, 0], y_truth, "k--", linewidth=1.5, label="truth")
ax.plot(
x_test[:, 0],
mean_ssgp,
"C0",
linewidth=1.8,
label=f"SSGP mean (MSE={mse_ssgp:.4f})",
)
ax.fill_between(
x_test[:, 0],
mean_ssgp - 2 * std_ssgp,
mean_ssgp + 2 * std_ssgp,
color="C0",
alpha=0.25,
label=r"$\pm 2\sigma$ (closed form)",
)
ax.axvspan(-0.2, 0.4, color="0.85", alpha=0.4, label="held-out gap")
ax.set_xlabel("$x$")
ax.set_ylabel("$y$")
ax.set_title("SSGP — closed-form predictive variance, no Monte Carlo needed")
ax.grid(True, alpha=0.3)
ax.legend(loc="upper right", fontsize=9)
plt.tight_layout()
plt.show()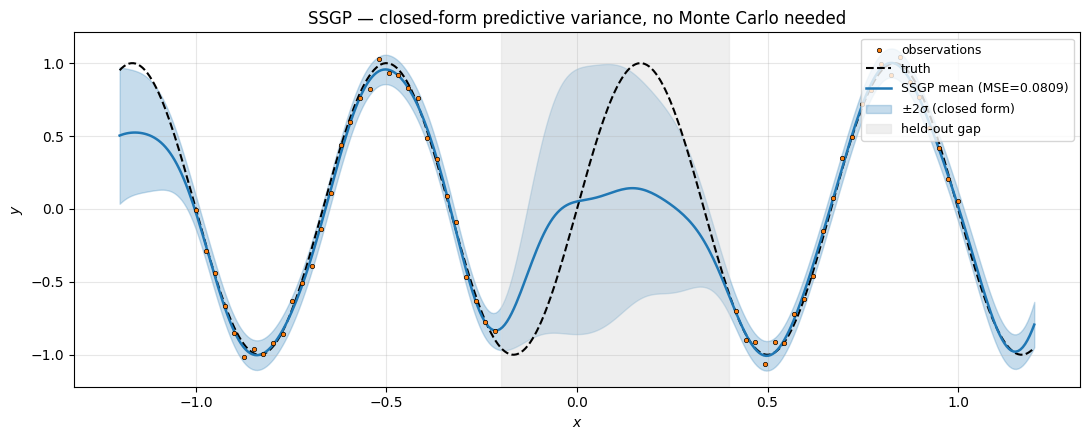
Two important things just happened.
- Hyperparameters fitted themselves. ML-II picked close to the true noise standard deviation 0.05, and close to a value that puts prior support over the target frequency — no manual tuning. This is the core advantage of the marginal-likelihood objective over the MAP MSE objective: the noise level becomes a learned hyperparameter.
- Predictive variance opens across the gap. No ensembling, no Monte Carlo sampling — the band is a pure function of . In the gap, is far from the training data in feature space, so inflates the variance. Outside (extrapolation) the band continues to widen, mirroring exact-GP behaviour.
4. VSSGP — Variational SSGP (Gal & Turner 2015)¶
Math. SSGP fixes Ω as a point estimate. VSSGP makes Ω itself a latent random variable with a prior (in lengthscale-1 units; the spectral density of the RBF kernel) and a learnable mean-field posterior . The head β is similarly variational: .
Tempered ELBO. With both Ω and β variational,
Both KLs have closed forms (Gaussian-Gaussian). The data term is estimated by reparameterising and with , and Monte-Carlo’ing over a few samples per step.
Why the temperature ? When the target’s spectral content lies far in the tail of , a strict ELBO can trap the posterior near the prior. Down-weighting the KL during training (β-VAE / KL-annealing) lets the posterior escape, after which we can ramp if calibration matters. Here we use throughout for a clean comparison.
Predictive distribution. Sample realisations of from , compute predictive means , then take empirical mean and variance plus the observation noise. The MC predictive variance captures uncertainty in both the frequencies and the weights — a richer band than SSGP’s, especially when the true spectrum is broad.
KL_BETA = 0.05
N_MC = 8
class VSSGP(eqx.Module):
"""Variational Sparse Spectrum GP — q(Ω), q(β) with reparameterisation."""
mu_W: jax.Array
log_sigma_W: jax.Array
mu_beta: jax.Array
log_sigma_beta: jax.Array
bias: jax.Array
log_sigma_n: jax.Array
lengthscale: float = eqx.field(static=True)
@classmethod
def init(
cls,
key,
in_features,
n_features,
lengthscale,
*,
mu_init_scale=5.0,
log_sigma_init=-1.0,
sigma_n=0.05,
):
kW, kb = jr.split(key)
return cls(
mu_W=mu_init_scale * jr.normal(kW, (in_features, n_features)),
log_sigma_W=jnp.full((in_features, n_features), log_sigma_init),
mu_beta=0.01 * jr.normal(kb, (2 * n_features,)),
log_sigma_beta=jnp.full((2 * n_features,), log_sigma_init),
bias=jnp.zeros(()),
log_sigma_n=jnp.log(jnp.array(sigma_n)),
lengthscale=lengthscale,
)
def _features(self, x, W):
z = x @ W / self.lengthscale
scale = jnp.sqrt(1.0 / W.shape[-1])
return scale * jnp.concatenate([jnp.cos(z), jnp.sin(z)], axis=-1)
def kl(self):
sigma_W = jnp.exp(self.log_sigma_W)
sigma_b = jnp.exp(self.log_sigma_beta)
kl_W = 0.5 * jnp.sum(self.mu_W**2 + sigma_W**2 - 1.0 - 2.0 * self.log_sigma_W)
kl_b = 0.5 * jnp.sum(
self.mu_beta**2 + sigma_b**2 - 1.0 - 2.0 * self.log_sigma_beta
)
return kl_W + kl_b
def predict_mc(self, x_query, key, n_samples=64):
"""Monte-Carlo predictive (mean, std) over q(Ω), q(β)."""
sigma_W = jnp.exp(self.log_sigma_W)
sigma_b = jnp.exp(self.log_sigma_beta)
kW, kb = jr.split(key)
ks_W = jr.split(kW, n_samples)
ks_b = jr.split(kb, n_samples)
def one_sample(kW_s, kb_s):
eps_W = jr.normal(kW_s, self.mu_W.shape)
eps_b = jr.normal(kb_s, self.mu_beta.shape)
W = self.mu_W + sigma_W * eps_W
beta = self.mu_beta + sigma_b * eps_b
phi = self._features(x_query, W)
return phi @ beta + self.bias
preds = jax.vmap(one_sample)(ks_W, ks_b) # (S, n_query)
sigma_n = jnp.exp(self.log_sigma_n)
mean = jnp.mean(preds, axis=0)
# Total predictive var = epistemic + aleatoric.
var = jnp.var(preds, axis=0) + sigma_n**2
return mean, jnp.sqrt(var), preds
def vssgp_elbo(model, x, y, key):
sigma_W = jnp.exp(model.log_sigma_W)
sigma_b = jnp.exp(model.log_sigma_beta)
sigma_n2 = jnp.exp(2.0 * model.log_sigma_n)
keys_W = jr.split(key, N_MC)
def one_sample(k):
kW_s, kb_s = jr.split(k)
eps_W = jr.normal(kW_s, model.mu_W.shape)
eps_b = jr.normal(kb_s, model.mu_beta.shape)
W = model.mu_W + sigma_W * eps_W
beta = model.mu_beta + sigma_b * eps_b
phi = model._features(x, W)
pred = phi @ beta + model.bias
return -0.5 / sigma_n2 * jnp.sum((pred - y) ** 2)
nll = -jnp.mean(jax.vmap(one_sample)(keys_W))
return nll + KL_BETA * model.kl()
def fit_vssgp(model, x_obs, y_obs, *, n_steps=4000, lr=1e-2, seed=0):
opt = optax.adam(lr)
state = opt.init(eqx.filter(model, eqx.is_inexact_array))
@eqx.filter_jit
def step(m, s, k):
loss, grads = eqx.filter_value_and_grad(vssgp_elbo)(m, x_obs, y_obs, k)
upd, s = opt.update(grads, s, eqx.filter(m, eqx.is_inexact_array))
return eqx.apply_updates(m, upd), s, loss
key = jr.PRNGKey(seed)
losses = []
for _ in range(n_steps):
key, sub = jr.split(key)
model, state, loss = step(model, state, sub)
losses.append(float(loss))
return model, losses
vssgp = VSSGP.init(
jr.PRNGKey(13),
in_features=1,
n_features=M_FEAT,
lengthscale=LENGTHSCALE_INIT,
)
vssgp, vssgp_losses = fit_vssgp(vssgp, x_obs, y_obs.reshape(-1))
mean_v, std_v, preds_v = vssgp.predict_mc(x_test, jr.PRNGKey(99), n_samples=128)
mse_v = float(jnp.mean((mean_v - y_truth) ** 2))
print(
f"VSSGP: final tempered-ELBO = {vssgp_losses[-1]:.2f}, "
f"σ_n = {float(jnp.exp(vssgp.log_sigma_n)):.4f}, MSE = {mse_v:.4f}"
)VSSGP: final tempered-ELBO = 30.70, σ_n = 0.4011, MSE = 0.0163
fig, ax = plt.subplots(figsize=(11, 4.5))
ax.scatter(
x_obs[:, 0],
y_obs,
s=10,
color="C1",
edgecolors="k",
linewidths=0.5,
label="observations",
zorder=5,
)
ax.plot(x_test[:, 0], y_truth, "k--", linewidth=1.5, label="truth")
for member in preds_v[:8]:
ax.plot(x_test[:, 0], member, color="C2", alpha=0.15, linewidth=0.8)
ax.plot(
x_test[:, 0], mean_v, "C2", linewidth=1.8, label=f"VSSGP mean (MSE={mse_v:.4f})"
)
ax.fill_between(
x_test[:, 0],
mean_v - 2 * std_v,
mean_v + 2 * std_v,
color="C2",
alpha=0.25,
label=r"$\pm 2\sigma$ (MC over $q(\Omega), q(\beta)$)",
)
ax.axvspan(-0.2, 0.4, color="0.85", alpha=0.4, label="held-out gap")
ax.set_xlabel("$x$")
ax.set_ylabel("$y$")
ax.set_title("VSSGP — full posterior over Ω and β, MC predictive band")
ax.grid(True, alpha=0.3)
ax.legend(loc="upper right", fontsize=9)
plt.tight_layout()
plt.show()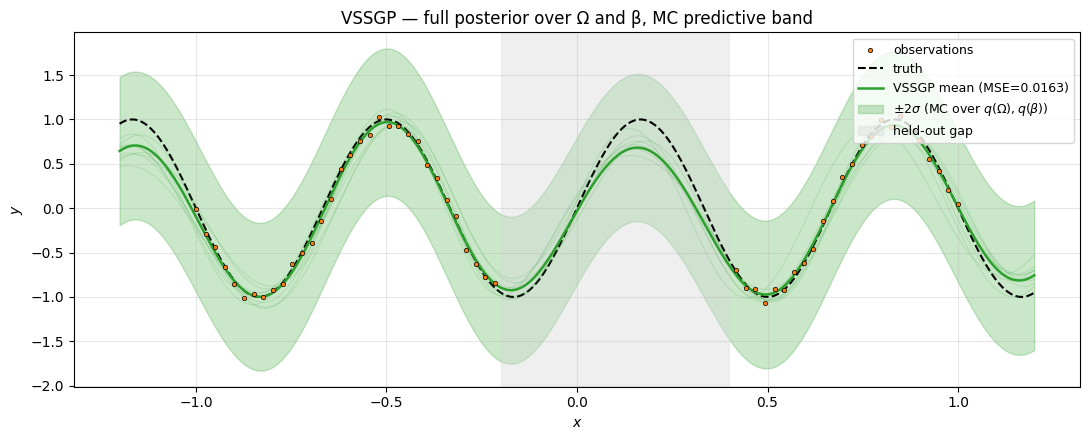
Same target, similar fit quality, richer uncertainty: the grey member traces are independent draws from , and they pull apart in the gap exactly like an exact-GP posterior would. The key conceptual addition over SSGP is that the band reflects frequency uncertainty too — if the target’s true frequency had been less well-determined by the data, the band’s behaviour in the gap would have been broader than SSGP’s.
Spectrum migration. Plot for SSGP (point estimate) vs VSSGP (posterior mean) to see how the trained frequencies relate to the target’s .
fig, ax = plt.subplots(figsize=(8, 4.5))
freqs_ssgp = (ssgp.W / jnp.exp(ssgp.log_ell))[0]
freqs_vssgp = (vssgp.mu_W / vssgp.lengthscale)[0]
all_f = np.concatenate(
[np.abs(np.asarray(freqs_ssgp)), np.abs(np.asarray(freqs_vssgp))]
)
bins = np.linspace(0, max(20, float(all_f.max()) * 1.05), 25)
ax.hist(
np.abs(np.asarray(freqs_ssgp)),
bins=bins,
alpha=0.5,
color="C0",
label="SSGP $|\\omega_j| = |W_j|/\\ell$ (point estimate)",
)
ax.hist(
np.abs(np.asarray(freqs_vssgp)),
bins=bins,
alpha=0.5,
color="C2",
label=r"VSSGP $|\mu_{\Omega,j}|/\ell$ (posterior mean)",
)
ax.axvline(
float(TARGET_OMEGA),
color="k",
linestyle="--",
alpha=0.8,
label=r"target $\omega^\star = 3\pi$",
)
ax.set_xlabel(r"$|\omega_j|$")
ax.set_ylabel("count")
ax.set_title(f"Spectrum used by each model — $M = {M_FEAT}$")
ax.legend(loc="upper right", fontsize=9)
ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()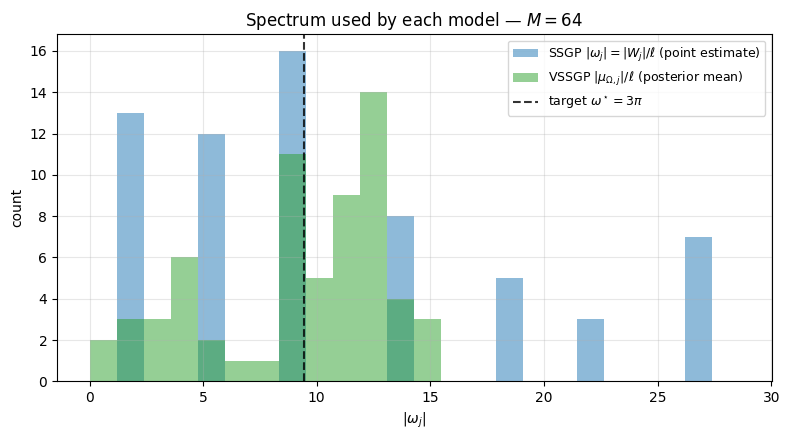
Both models concentrate frequency mass around — SSGP via the marginal-likelihood gradient, VSSGP via the data-fit term of the ELBO (the KL pulls back toward the prior at , but the data wins for the frequencies that matter).
5. Three methods, one plot — the uncertainty hierarchy¶
Same target, same data, three methods, three predictive bands.
fig, axes = plt.subplots(1, 3, figsize=(18, 4.5), sharey=True)
def _decorate(ax, title):
ax.scatter(
x_obs[:, 0],
y_obs,
s=10,
color="C1",
edgecolors="k",
linewidths=0.5,
label="observations",
zorder=5,
)
ax.plot(x_test[:, 0], y_truth, "k--", linewidth=1.5, label="truth")
ax.axvspan(-0.2, 0.4, color="0.85", alpha=0.4, label="held-out gap")
ax.set_title(title)
ax.set_xlabel("$x$")
ax.grid(True, alpha=0.3)
ax.legend(loc="upper right", fontsize=8)
ax = axes[0]
ax.plot(x_test[:, 0], y_pred_map, "C3", linewidth=1.8, label=f"MAP (MSE={mse_map:.4f})")
_decorate(ax, "Learned RFF MAP — point estimate, no uncertainty")
ax.set_ylabel("$y$")
ax = axes[1]
ax.plot(
x_test[:, 0],
mean_ssgp,
"C0",
linewidth=1.8,
label=f"SSGP mean (MSE={mse_ssgp:.4f})",
)
ax.fill_between(
x_test[:, 0],
mean_ssgp - 2 * std_ssgp,
mean_ssgp + 2 * std_ssgp,
color="C0",
alpha=0.25,
label=r"$\pm 2\sigma$ closed form",
)
_decorate(ax, "SSGP — analytic posterior on β, point Ω")
ax = axes[2]
for member in preds_v[:8]:
ax.plot(x_test[:, 0], member, color="C2", alpha=0.15, linewidth=0.8)
ax.plot(
x_test[:, 0], mean_v, "C2", linewidth=1.8, label=f"VSSGP mean (MSE={mse_v:.4f})"
)
ax.fill_between(
x_test[:, 0],
mean_v - 2 * std_v,
mean_v + 2 * std_v,
color="C2",
alpha=0.25,
label=r"$\pm 2\sigma$ MC",
)
_decorate(ax, "VSSGP — posterior on both Ω and β")
plt.tight_layout()
plt.show()
# Sanity-check uncertainty values across the three methods.
trained = ((x_test[:, 0] > -0.5) & (x_test[:, 0] < -0.3)) | (
(x_test[:, 0] > 0.5) & (x_test[:, 0] < 0.7)
)
gap_region = (x_test[:, 0] > -0.2) & (x_test[:, 0] < 0.4)
for name, std_arr in [("SSGP", std_ssgp), ("VSSGP", std_v)]:
ratio = float(jnp.mean(std_arr[gap_region]) / jnp.mean(std_arr[trained]))
print(f"{name}: gap/data std ratio = {ratio:.1f}x")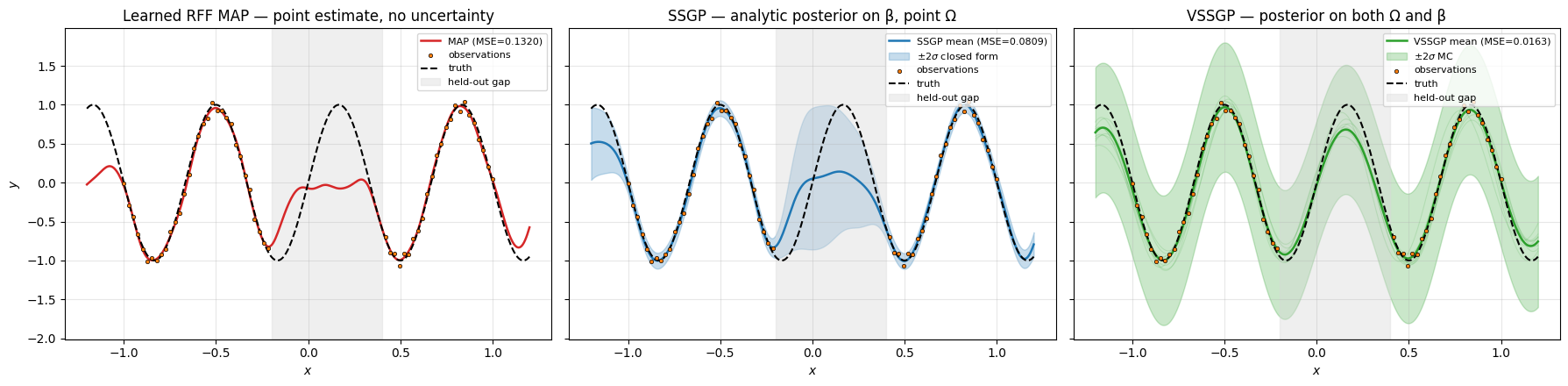
SSGP: gap/data std ratio = 6.1x
VSSGP: gap/data std ratio = 1.0x
Reading the figure left-to-right:
- Learned RFF MAP (left) — clean fit, but no signal that the gap is uncertain. The curve through is just whatever the optimiser found.
- SSGP (centre) — same fit quality, plus a closed-form band that opens visibly across the gap. The marginal-likelihood objective also tuned and for free.
- VSSGP (right) — same fit + an MC band that captures uncertainty over both Ω and β. The grey member traces show how the posterior draws disagree in the gap.
Cost ladder. MAP: Adam steps, no per-prediction overhead. SSGP: per ML-II step, per prediction (closed-form). VSSGP: per ELBO step (with MC samples), per prediction. SSGP is usually the sweet spot for moderate data; VSSGP wins when frequency uncertainty actually matters or when the marginal-likelihood Cholesky gets unstable.
Takeaways¶
- The same RFF feature map (whose kernel-approximation properties are pinned down in the Kernel Approximation notebook) admits three Bayesian regimes, all with trained Ω:
- Learned RFF MAP — point estimates everywhere, regularised MSE. Cheapest, no uncertainty.
- SSGP (Lázaro-Gredilla et al. 2010) — analytic marginalisation of the head β, train Ω on the GP marginal likelihood. Closed-form predictive variance for free; and tuned by ML-II.
- VSSGP (Gal & Turner 2015) — variational posteriors over both Ω and β, tempered ELBO with reparameterisation. MC predictive band that captures frequency uncertainty.
- Predictive uncertainty hierarchy. MAP gives no band. SSGP gives a closed-form band on the head’s posterior. VSSGP gives an MC band on the joint posterior over frequencies and head — the richest, at the cost of variational machinery.
Where to next.
- The non-Bayesian / NN flavors of RFF (fixed-Ω with closed-form ridge, ensemble-of-MAP for cheap predictive uncertainty) live in the RFF as Neural Networks notebook.
- The deep / hierarchical versions (deep RFF, deep SSGP, deep VSSGP — Cutajar et al. 2017) live in the Deep Random Feature Expansions notebook, which builds on the per-layer SSGP / VSSGP primitives derived here.