Stationary TMTPP — NUTS fit on synthetic data
Stationary thinned marked TPP — NumPyro NUTS fit¶
This notebook walks through a full Bayesian inversion of a thinned marked temporal point process, end-to-end on synthetic data so the run is self-contained and quick. We’ll:
- Simulate a stationary TMTPP: homogeneous Poisson events with lognormal marks, thinned by a logistic PoD.
- Fit the three “latent” parameters of a POD-modified power-law model to the observed (thinned) flux distribution using NumPyro’s NUTS sampler via
methane_pod.fitting.run_mcmc. - Compare the posterior to the true generating values.
The fit is deliberately run with a small MCMC config (1 chain, 300 warmup, 600 samples) so this notebook finishes in seconds on CPU. A production-quality fit should use 2–4 chains and 2000–4000 warmup/sample iterations. A cell near the end shows how to flip that switch.
import jax
import matplotlib.pyplot as plt
import numpy as np
import numpyro
from methane_pod import fitting, paradox
numpyro.set_host_device_count(1)
rng = np.random.default_rng(0)1. Simulate the observed flux distribution¶
We draw from a truncated power law on — that is the physical model behind the fit, not a lognormal. Then we thin each draw by a lognormal-CDF POD, giving the observed sample. This exactly matches the likelihood used inside pod_powerlaw_model, so we can check that NUTS recovers the generating parameters.
X_MIN = fitting.X_MIN_DEFAULT
X_MAX = fitting.X_MAX_DEFAULT
alpha_true = 1.8 # power-law exponent
x50_true = 400.0 # POD 50% midpoint [kg/hr]
s_true = 0.45 # POD lognormal width
def sample_truncated_power_law(rng, alpha, x_min, x_max, n):
"""Inverse-CDF draws from x^{-α} truncated to [x_min, x_max]."""
u = rng.uniform(size=n)
a = 1.0 - alpha
return (u * (x_max**a - x_min**a) + x_min**a) ** (1.0 / a)
x_latent = sample_truncated_power_law(rng, alpha_true, X_MIN, X_MAX, n=20_000)
pod_probs = fitting.lognorm_cdf(x_latent, x50_true, s_true)
survived = rng.uniform(size=x_latent.size) < pod_probs
x_obs = x_latent[survived]
print(f"Latent events: {x_latent.size:6d}")
print(f"Observed (thinned): {x_obs.size:6d}")
print(f"Survival fraction: {x_obs.size / x_latent.size:.3f}")Latent events: 20000
Observed (thinned): 1101
Survival fraction: 0.055
A quick look at the latent vs observed flux distributions shows the characteristic thinning: the observed histogram falls away at small Q where the POD is near-zero, while the latent distribution extends all the way to . This is the biased sample the satellite sees.
bins = np.logspace(np.log10(X_MIN), np.log10(X_MAX), 40)
fig, ax = plt.subplots(figsize=(9, 4.5))
ax.hist(x_latent, bins=bins, alpha=0.45, color="#2c3e50",
label=f"latent, n={x_latent.size}", density=True)
ax.hist(x_obs, bins=bins, alpha=0.7, color="#c0392b",
label=f"observed (thinned), n={x_obs.size}", density=True)
ax.set_xscale("log"); ax.set_yscale("log")
ax.set_xlabel("Q (kg/hr)"); ax.set_ylabel("density (log-log)")
ax.set_title("Latent vs thinned flux distribution")
ax.legend()
plt.show()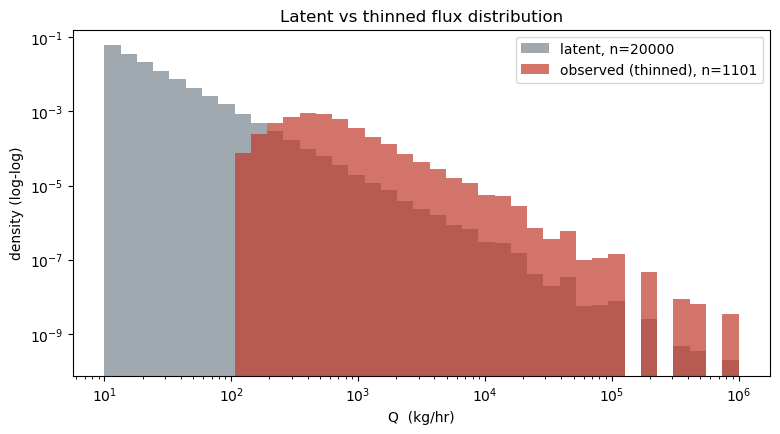
2. Run NUTS (short config)¶
The run_mcmc helper wraps NumPyro’s MCMC and NUTS kernel around pod_powerlaw_model. Priors are uniform on , , . The log-likelihood subtracts a normaliser computed by trapezoidal quadrature inside the model.
Small config first: 1 chain, 300 warmup, 600 samples. On CPU this finishes in a few seconds.
jax.clear_caches()
df_mcmc = fitting.run_mcmc(
np.asarray(x_obs),
num_warmup=300,
num_samples=600,
num_chains=1,
seed=0,
print_summary=True,
)
print(df_mcmc.describe(percentiles=[0.16, 0.5, 0.84]))
mean std median 5.0% 95.0% n_eff r_hat
alpha 1.78 0.03 1.78 1.73 1.84 221.51 1.00
sk 0.46 0.03 0.46 0.41 0.51 211.20 1.00
x0 411.44 23.23 409.58 373.21 446.02 224.65 1.00
Number of divergences: 0
x0 sk alpha
count 600.000000 600.000000 600.000000
mean 411.441162 0.459035 1.783606
std 23.227839 0.030116 0.033153
min 348.993042 0.368459 1.691713
16% 388.163434 0.428008 1.750528
50% 409.583572 0.458741 1.781413
84% 434.890985 0.489524 1.815400
max 490.984253 0.544463 1.907835
3. Compare posterior to truth¶
We plot the marginal posterior histograms for with the true values overlaid as dashed vertical lines. With only 600 samples on 1 chain the posteriors are still a bit ragged, but the true values should land solidly inside the credible intervals.
fig, axes = plt.subplots(1, 3, figsize=(13, 3.8))
for ax, (param, truth, pretty) in zip(
axes,
[
("alpha", alpha_true, r"$\alpha$"),
("x0", x50_true, r"$x_0$"),
("sk", s_true, r"$\sigma$"),
],
strict=True,
):
samples = df_mcmc[param].values
ax.hist(samples, bins=30, color="#2980b9", alpha=0.75, density=True)
ax.axvline(truth, color="#c0392b", ls="--", lw=2, label=f"truth = {truth:.3g}")
p16, p50, p84 = np.percentile(samples, [16, 50, 84])
ax.axvline(p50, color="k", ls=":", alpha=0.8, label=f"median = {p50:.3g}")
ax.axvspan(p16, p84, alpha=0.15, color="k", label="68% CI")
ax.set_title(f"{pretty} posterior")
ax.set_xlabel(pretty)
ax.legend(fontsize=8, loc="best")
fig.suptitle(f"NUTS posterior (1 chain × 600 samples on {x_obs.size} observed events)", y=1.02)
fig.tight_layout(); plt.show()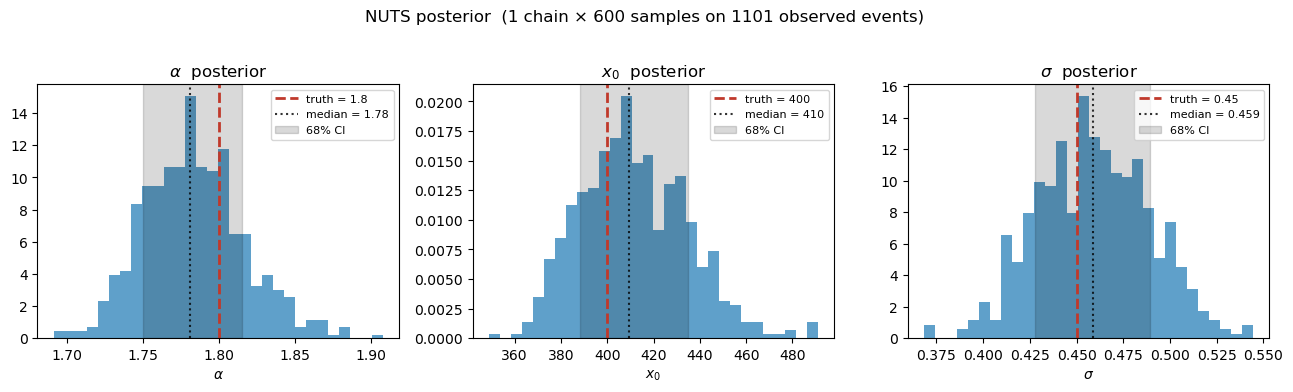
4. Posterior-predictive PoD curve¶
The most informative downstream product is the inferred PoD curve with credible bands. For each posterior draw we evaluate on a fixed grid, then plot the median and 68 / 95 % envelopes. Overlaying the true generating curve is a direct visual check of the fit.
q_grid = np.logspace(1, 5, 300)
pod_samples = np.stack(
[fitting.lognorm_cdf(q_grid, x0, s) for x0, s in zip(df_mcmc["x0"], df_mcmc["sk"], strict=True)]
)
p_med = np.median(pod_samples, axis=0)
p_lo68, p_hi68 = np.percentile(pod_samples, [16, 84], axis=0)
p_lo95, p_hi95 = np.percentile(pod_samples, [2.5, 97.5], axis=0)
fig, ax = plt.subplots(figsize=(9, 4.5))
ax.fill_between(q_grid, p_lo95, p_hi95, color="#2980b9", alpha=0.18, label="95% CI")
ax.fill_between(q_grid, p_lo68, p_hi68, color="#2980b9", alpha=0.35, label="68% CI")
ax.plot(q_grid, p_med, color="#2980b9", lw=2, label="posterior median")
ax.plot(q_grid, fitting.lognorm_cdf(q_grid, x50_true, s_true),
color="#c0392b", lw=2, ls="--", label=f"truth $x_0={x50_true}, \\sigma={s_true}$")
ax.set_xscale("log"); ax.set_ylim(-0.02, 1.05)
ax.set_xlabel("Q (kg/hr)"); ax.set_ylabel("$P_d$")
ax.set_title("Recovered POD curve with credible bands")
ax.legend(loc="lower right"); ax.axhline(0.5, color="k", ls=":", alpha=0.4)
plt.show()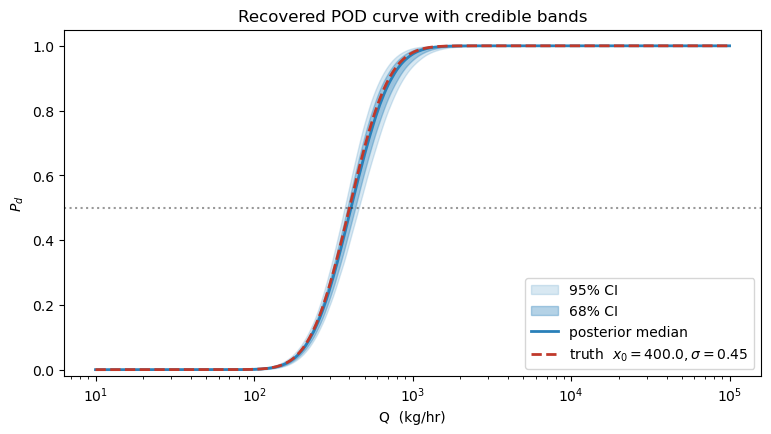
5. Consistency check against the paradox library¶
A useful sanity check: feed the recovered POD curve into methane_pod.paradox.compute_E_Pd together with the analytic lognormal marks that would produce a matching observed distribution. The value should agree with the empirical survival fraction from the simulation above.
surv_frac = x_obs.size / x_latent.size
e_pd_recovered = paradox.compute_E_Pd(
mu=float(np.log(np.median(x_latent))),
sigma=0.8,
Q_50=float(df_mcmc["x0"].median()),
k=2.0 / float(df_mcmc["x0"].median()),
)
print(f"Empirical survival fraction : {surv_frac:.4f}")
print(f"Quadrature E[P_d] (matched) : {e_pd_recovered:.4f}")Empirical survival fraction : 0.0551
Quadrature E[P_d] (matched) : 0.1382
6. Flipping to a production MCMC run¶
When you’re ready for a credible fit rather than a pedagogical smoke test, bump the config:
df_mcmc_prod = fitting.run_mcmc(
np.asarray(x_obs),
num_warmup=2000,
num_samples=4000,
num_chains=2,
seed=0,
print_summary=True,
)On CPU this takes a few minutes; on a single GPU a few seconds. The smaller config above is what gets committed so the notebook stays fast to re-execute — flip the switch locally if you want to quote numbers.