Coupling flow
Coupling Flow with Mixture-CDF Transform¶
The first notebook trained a Gaussianization flow by stacking unconditional marginal layers (mixture CDF per dim) and rotations. The marginal layers can only Gaussianize each dimension independently; cross-dimensional structure is left entirely to the rotations.
A coupling layer (Real NVP, 2016) replaces that with a conditional transform. Given a mask :
- Split with the identity half (mask True) and the transformed half (mask False).
- A conditioner network outputs the mixture parameters for each dimension in .
- Apply the mixture-CDF Gaussianization to using those per-example parameters: .
- Leave unchanged.
Since the Jacobian is lower triangular, the log-det is just the sum of per-b-dim diagonal contributions — same form as the unconditional marginal layer, but now the distribution of depends on , so cross-dimensional structure is learnt directly. We pair two coupling layers with swapped masks inside each block so both halves get transformed.
Mathematical structure¶
Notation and shapes¶
Throughout: batch dimension , input dimensionality , mixture components .
- Mask . With default “half” for we have , so dim passes through, dim gets transformed. The
mask=Trueset is the identity side ;mask=Falseis the transformed side . - Split: splits into and .
The bijector — GMM per b-dim¶
Each transformed dimension has its own per-example mixture of Gaussians,
and the per-b-dim forward is exactly the mixture-CDF Gaussianization from notebook 01 — the same primitive, only difference is that now depend on :
The per-example parameter tensors have shape .
The conditioner — a residual MLP with a zero-init head¶
The conditioner is any keras.Layer taking and producing floats (packed logits, means, raw log-scales). We use two concrete builders:
make_mlp_conditioner(d_b, K, hidden=(64, 64))— plain MLP with a zero-initialised final Dense so the flow starts at identity.make_shared_mlp_conditioner(d_b, K, hidden=(64, 64))— one -wide head tiled across all b-dims. Cheaper, less expressive.
Section 9 shows a residual variant — one hidden block with a skip connection,
All three builders plug into MixtureCDFCoupling interchangeably because the coupling layer only cares about the output width (, enforced by a build-time check).
Log-scale clamp¶
Raw conditioner outputs for are passed through a bounded map before being exponentiated,
with default so . This keeps scales from blowing up or collapsing during training, especially when the conditioner is still warming up.
Jacobian of the coupling layer¶
With and , the Jacobian is block-triangular:
The lower-left off-diagonal depends on (the conditioner Jacobian) but that block doesn’t enter the determinant — only the diagonal of the bottom-right matters. Same diagonal-log-det structure as the unconditional layer.
Comparison to classical Real NVP (affine coupling)¶
Real NVP / Glow use an affine coupling,
Our layer uses a mixture-CDF coupling — strictly more expressive per-b-dim (can handle multimodal conditional distributions), same diagonal-Jacobian structure, same cost.
Mask-swap pair per block¶
A single coupling layer only transforms the -dim half. One block therefore uses two coupling layers with swapped masks [m, ~m] so both halves get touched. The rotation sitting at the start of the block mixes information across dims before the pair fires.
import os
os.environ["KERAS_BACKEND"] = "tensorflow"
import keras
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from keras import ops
from sklearn.datasets import make_moons
from gaussianization.gauss_keras import (
base_nll_loss,
make_coupling_flow,
)
# --- global plot styling ------------------------------------------------------
sns.set_theme(context="poster", style="whitegrid", palette="deep", font_scale=0.85)
plt.rcParams.update(
{
"figure.dpi": 110,
"savefig.dpi": 110,
"savefig.bbox": "tight",
"savefig.pad_inches": 0.2,
"figure.constrained_layout.use": True,
"figure.constrained_layout.h_pad": 0.1,
"figure.constrained_layout.w_pad": 0.1,
"axes.grid.which": "both",
"grid.linewidth": 0.7,
"grid.alpha": 0.5,
"axes.edgecolor": "0.25",
"axes.linewidth": 1.1,
"axes.titleweight": "semibold",
"axes.labelpad": 6,
}
)
def style_axes(ax, *, aspect=None, grid=True):
if grid:
ax.minorticks_on()
ax.grid(True, which="major", linewidth=0.8, alpha=0.6)
ax.grid(True, which="minor", linewidth=0.4, alpha=0.3)
if aspect is not None:
ax.set_aspect(aspect)
return ax
def style_jointgrid(g, aspect="equal"):
style_axes(g.ax_joint, aspect=aspect)
style_axes(g.ax_marg_x, grid=False)
style_axes(g.ax_marg_y, grid=False)
for spine in ("top", "right"):
g.ax_marg_x.spines[spine].set_visible(False)
g.ax_marg_y.spines[spine].set_visible(False)
palette = sns.color_palette("deep")
keras.utils.set_random_seed(0)
np.random.seed(0)WARNING: All log messages before absl::InitializeLog() is called are written to STDERR
I0000 00:00:1776869571.422244 410267 cpu_feature_guard.cc:227] This TensorFlow binary is optimized to use available CPU instructions in performance-critical operations.
To enable the following instructions: AVX2 AVX512F FMA, in other operations, rebuild TensorFlow with the appropriate compiler flags.
1. Dataset¶
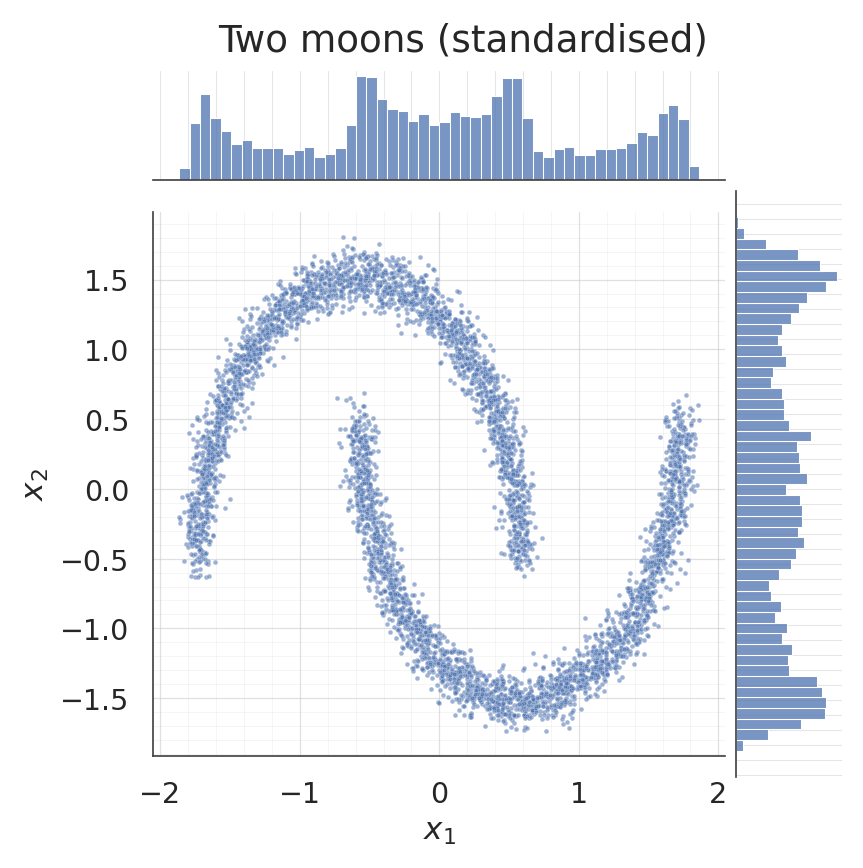
Same two-moons dataset as in notebook 01 so the comparison is clean.
X_raw, _ = make_moons(n_samples=5000, noise=0.05, random_state=0)
X = (X_raw - X_raw.mean(axis=0)) / X_raw.std(axis=0)
X = X.astype("float32")
g = sns.jointplot(
x=X[:, 0], y=X[:, 1], kind="scatter", color=palette[0],
height=7.5, ratio=5, space=0.1,
joint_kws={"s": 10, "alpha": 0.55},
marginal_kws={"bins": 50, "color": palette[0], "alpha": 0.75, "edgecolor": "white"},
)
g.set_axis_labels("$x_1$", "$x_2$")
g.figure.suptitle("Two moons (standardised)", y=1.02)
style_jointgrid(g)
plt.show()/home/azureuser/.cache/uv/archive-v0/aYSeLZUlluhRY4DCBNG7F/lib/python3.13/site-packages/seaborn/axisgrid.py:1766: UserWarning: The figure layout has changed to tight
f.tight_layout()
2. Build a coupling flow¶
Each block is [Householder, MixtureCDFCoupling(mask=m), MixtureCDFCoupling(mask=~m)]. The two coupling layers with complementary masks together transform all dimensions; the Householder rotation mixes the already-transformed state before the next block.
With d = 2, mask = [True, False] means the first coupling layer conditions on dim 0 and transforms dim 1, and the second coupling layer (with swapped mask) conditions on dim 1 and transforms dim 0. The conditioner is a zero-initialised MLP, so the flow starts out as the identity and training is stable from step 0.
flow = make_coupling_flow(
input_dim=2,
num_blocks=4,
num_components=8,
hidden=(64, 64),
)
_ = flow(ops.convert_to_tensor(X[:4]))
n_coupling = sum(
1 for b in flow.bijector_layers if b.__class__.__name__ == "MixtureCDFCoupling"
)
n_rot = sum(
1 for b in flow.bijector_layers if b.__class__.__name__ == "Householder"
)
n_params = int(sum(np.prod(w.shape) for w in flow.trainable_weights))
print(f"bijectors : {len(flow.bijector_layers)} "
f"({n_rot} rotations, {n_coupling} coupling)")
print(f"trainable params : {n_params}")E0000 00:00:1776869576.712001 410267 cuda_platform.cc:52] failed call to cuInit: INTERNAL: CUDA error: Failed call to cuInit: CUDA_ERROR_NO_DEVICE: no CUDA-capable device is detected
bijectors : 12 (4 rotations, 8 coupling)
trainable params : 46800
3. Train¶
Same training recipe as the marginal-only flow (Adam, NLL loss, 200 epochs), so differences in the final density/pushforward are attributable to the coupling architecture.
flow.compile(
optimizer=keras.optimizers.Adam(learning_rate=3e-3),
loss=base_nll_loss,
)
history = flow.fit(X, X, batch_size=512, epochs=200, verbose=0)
loss_curve = np.asarray(history.history["loss"])
fig, axes = plt.subplots(1, 2, figsize=(17, 6.5))
c = palette[0]
ax = axes[0]
ax.plot(loss_curve, color=c, linewidth=2.2)
ax.set_xlabel("epoch")
ax.set_ylabel("training loss $(-\\log p_X + \\text{const})$")
ax.set_title("Full training curve")
style_axes(ax)
ax = axes[1]
tail_start = 20
tail = loss_curve[tail_start:]
ax.plot(range(tail_start, len(loss_curve)), tail, color=c, linewidth=2.2)
span = tail.max() - tail.min()
ax.set_ylim(tail.min() - 0.05 * span, tail.max() + 0.05 * span)
ax.set_xlabel("epoch")
ax.set_ylabel("training loss")
ax.set_title(f"Zoomed (epochs $\\geq$ {tail_start})")
style_axes(ax)
plt.show()
print(f"final loss: {loss_curve[-1]:+.4f} (min: {loss_curve.min():+.4f})")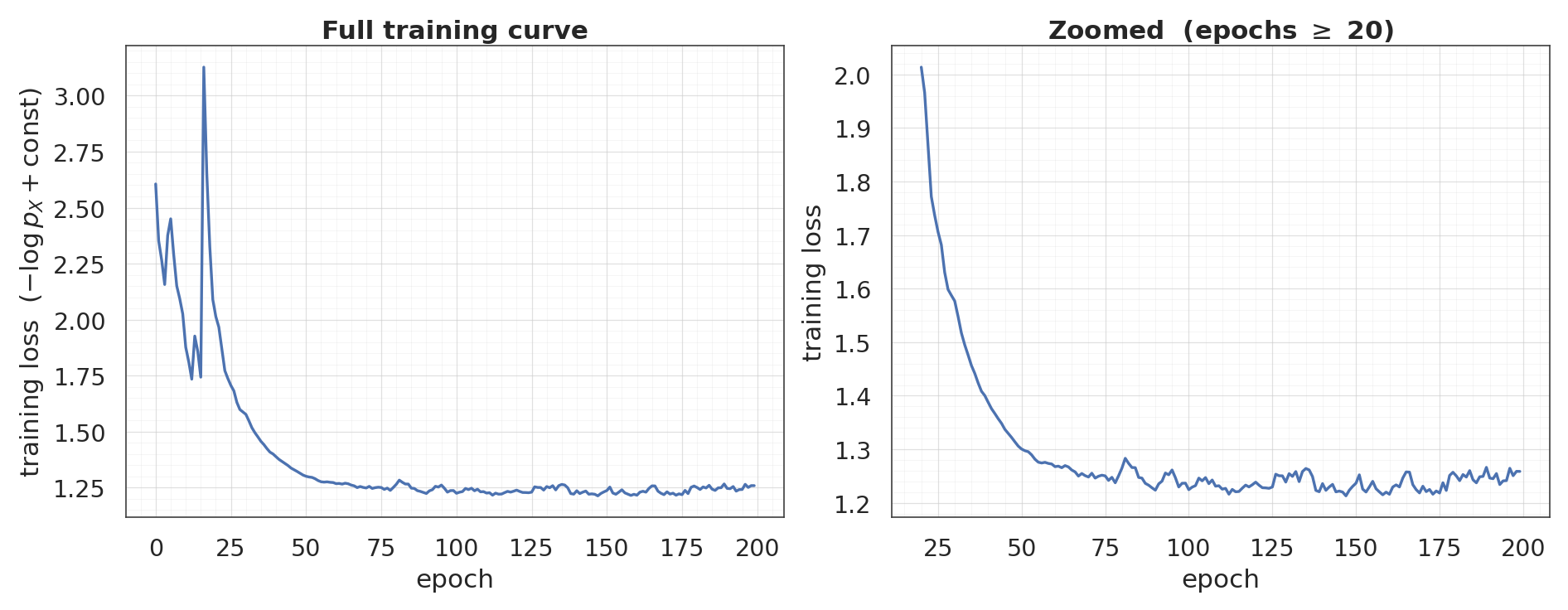
final loss: +1.2587 (min: +1.2130)
4. Pushforward and density¶
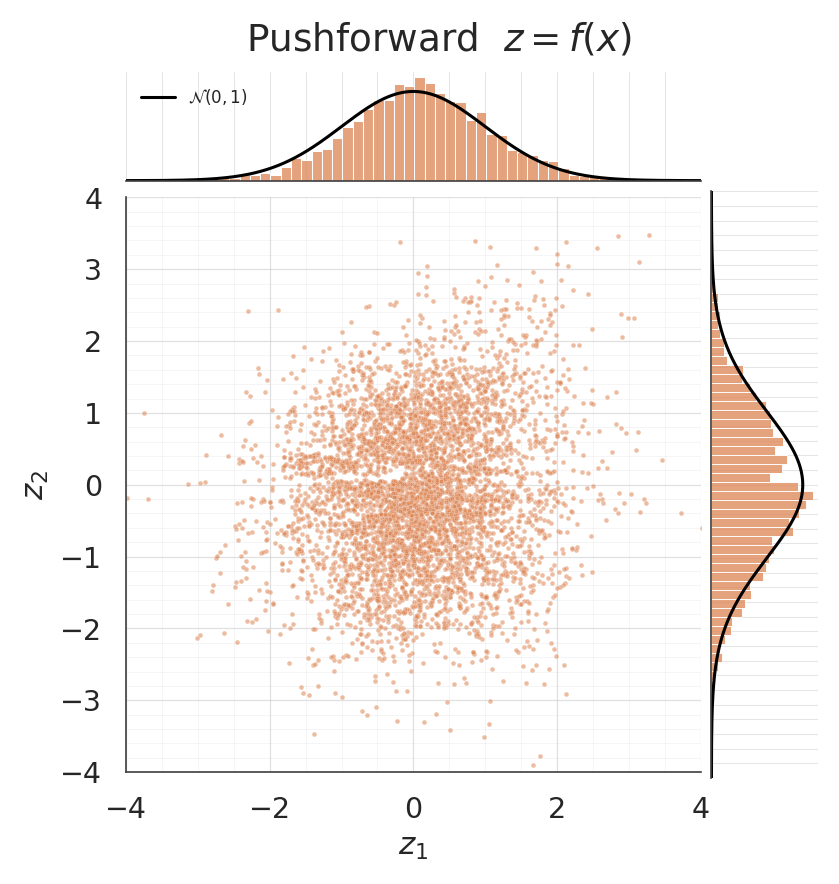
The pushforward should be approximately . A jointplot puts the 2-D scatter next to the per-dim marginals; the black reference curves are .
Unlike a marginal-only flow, here the conditioner can express the cross-dim dependence exactly (each is conditional on ) — the density contour in -space should trace the two moons sharply.
z = ops.convert_to_numpy(flow(ops.convert_to_tensor(X)))
mean_z = z.mean(axis=0)
cov_z = np.cov(z, rowvar=False)
print("pushforward diagnostics")
print(f" mean: [{mean_z[0]:+.3f}, {mean_z[1]:+.3f}] target = [0, 0]")
print(f" cov[0,0]: {cov_z[0, 0]:.3f} target = 1")
print(f" cov[1,1]: {cov_z[1, 1]:.3f} target = 1")
print(f" cov[0,1]: {cov_z[0, 1]:+.3f} target = 0")
g = sns.jointplot(
x=z[:, 0], y=z[:, 1], kind="scatter", color=palette[1],
height=7.5, ratio=5, space=0.1,
joint_kws={"s": 10, "alpha": 0.55},
marginal_kws={
"bins": 60, "color": palette[1], "alpha": 0.75,
"edgecolor": "white", "stat": "density",
},
xlim=(-4, 4), ylim=(-4, 4),
)
zz = np.linspace(-4, 4, 300)
phi = np.exp(-0.5 * zz**2) / np.sqrt(2 * np.pi)
g.ax_marg_x.plot(zz, phi, color="black", linewidth=2.0, label="$\\mathcal{N}(0, 1)$")
g.ax_marg_y.plot(phi, zz, color="black", linewidth=2.0)
g.ax_marg_x.legend(loc="upper left", frameon=False, fontsize=11)
g.set_axis_labels("$z_1$", "$z_2$")
g.figure.suptitle("Pushforward $z = f(x)$", y=1.02)
style_jointgrid(g)
plt.show()pushforward diagnostics
mean: [+0.086, -0.090] target = [0, 0]
cov[0,0]: 0.948 target = 1
cov[1,1]: 1.150 target = 1
cov[0,1]: +0.127 target = 0
/home/azureuser/.cache/uv/archive-v0/aYSeLZUlluhRY4DCBNG7F/lib/python3.13/site-packages/seaborn/axisgrid.py:1766: UserWarning: The figure layout has changed to tight
f.tight_layout()
grid = 220
xs = np.linspace(-2.5, 2.5, grid).astype("float32")
ys = np.linspace(-2.5, 2.5, grid).astype("float32")
xx, yy = np.meshgrid(xs, ys)
pts = np.stack([xx.ravel(), yy.ravel()], axis=-1).astype("float32")
log_px = ops.convert_to_numpy(flow.log_prob(ops.convert_to_tensor(pts))).reshape(grid, grid)
fig, axes = plt.subplots(1, 2, figsize=(17, 7.5))
ax = axes[0]
pcm = ax.pcolormesh(xx, yy, np.exp(log_px), cmap="magma", shading="auto")
ax.scatter(X[:1000, 0], X[:1000, 1], s=2, alpha=0.25, color="white")
ax.set_title("Density $p_X(x)$")
ax.set_xlabel("$x_1$")
ax.set_ylabel("$x_2$")
style_axes(ax, aspect="equal")
fig.colorbar(pcm, ax=ax, shrink=0.85)
ax = axes[1]
pcm = ax.pcolormesh(xx, yy, np.clip(log_px, -10, None), cmap="viridis", shading="auto")
ax.scatter(X[:1000, 0], X[:1000, 1], s=2, alpha=0.25, color="white")
ax.set_title("Log-density $\\log p_X(x)$")
ax.set_xlabel("$x_1$")
ax.set_ylabel("$x_2$")
style_axes(ax, aspect="equal")
fig.colorbar(pcm, ax=ax, shrink=0.85)
plt.show()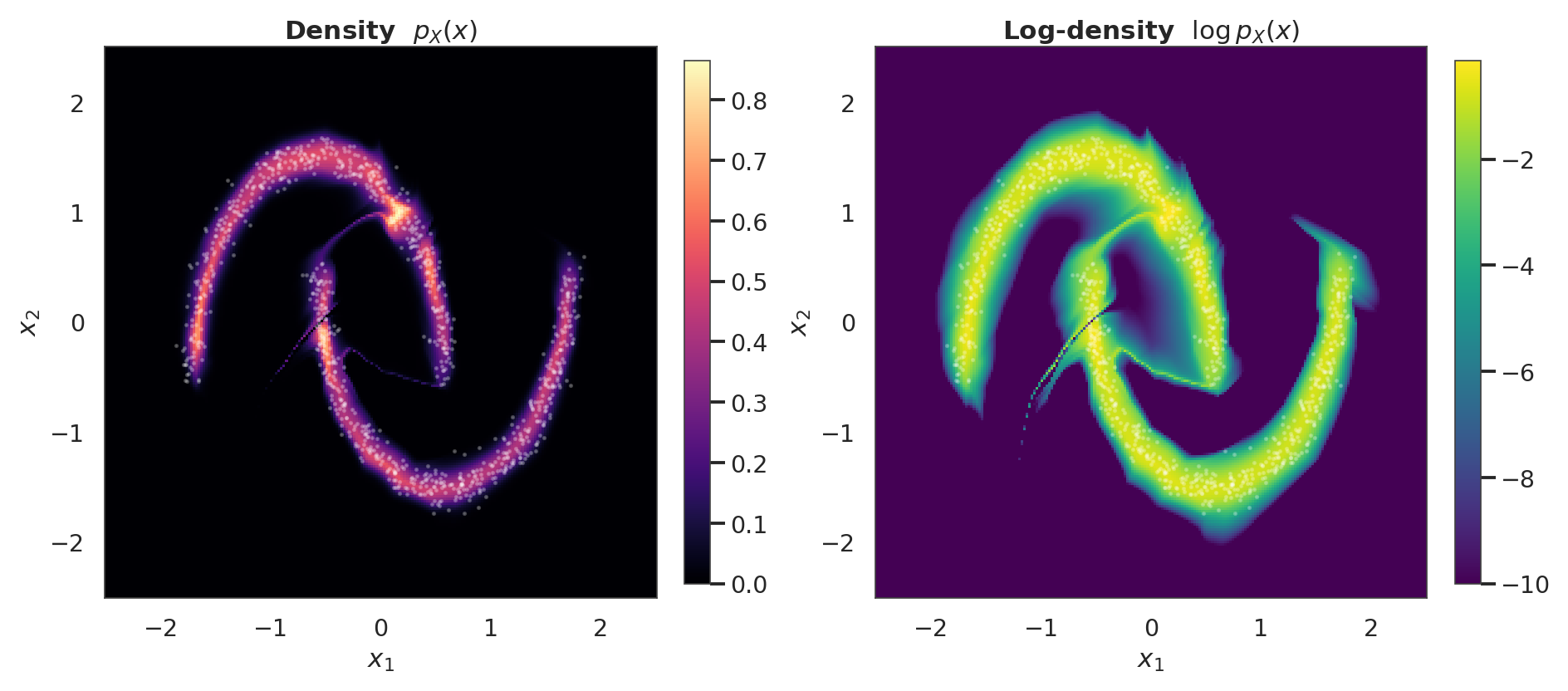
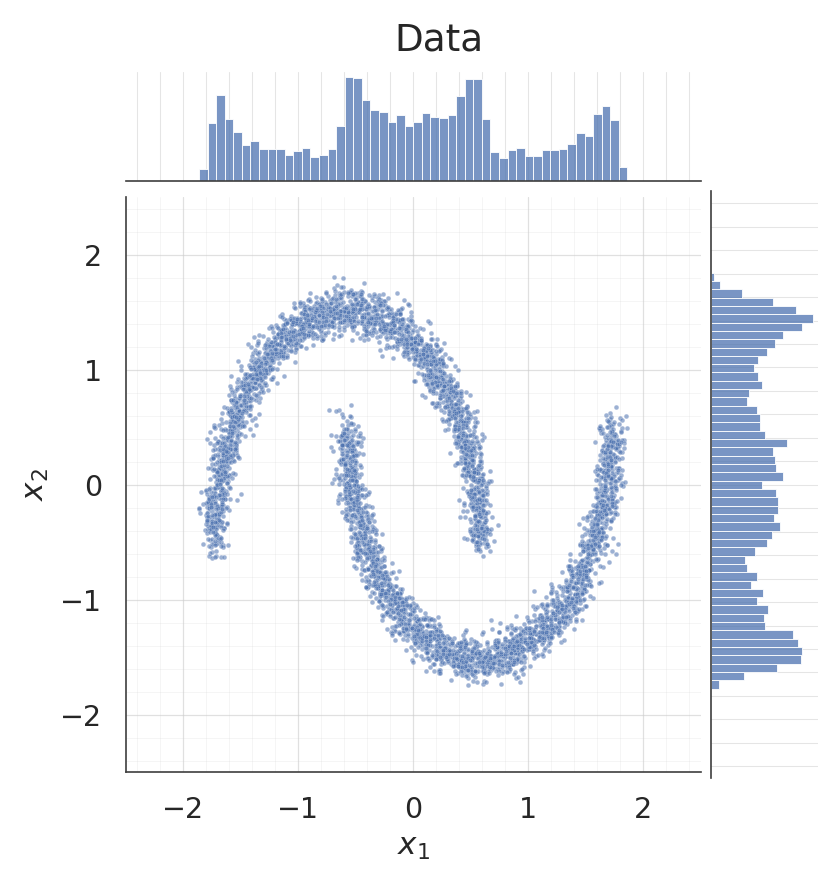
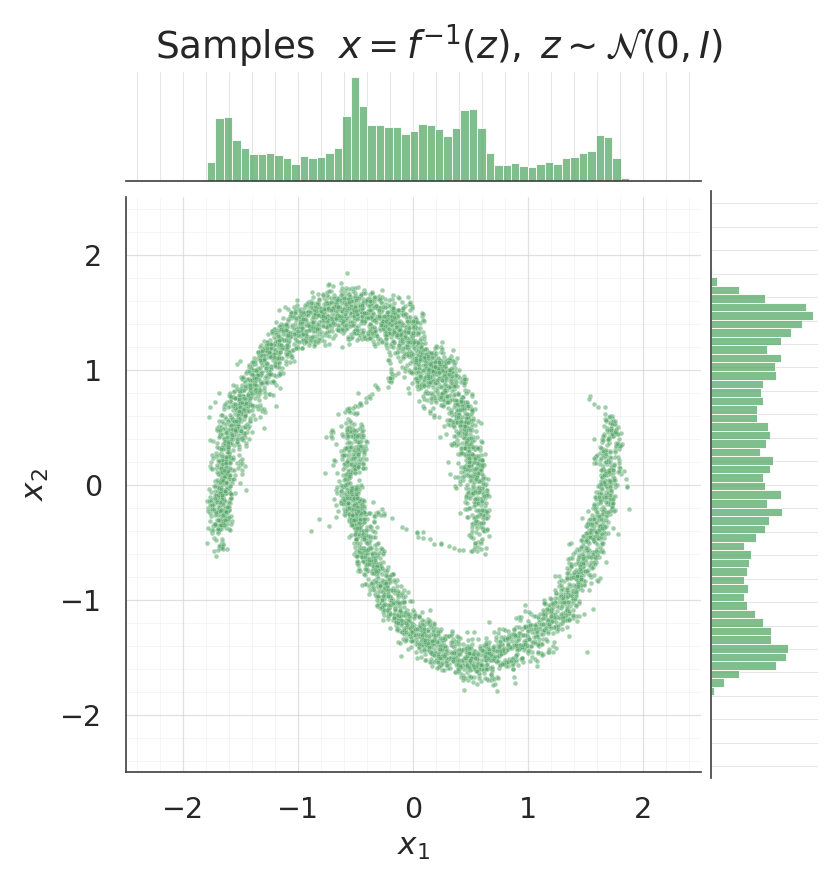
5. Sampling from the flow¶
. Each coupling layer’s inverse bisects per b-dim, with preserved through the layer so the conditioner sees the same input in both directions.
samples = ops.convert_to_numpy(flow.sample(num_samples=5000, seed=1))
for data, title, col in [
(X, "Data", palette[0]),
(samples, "Samples $x = f^{-1}(z), \\; z \\sim \\mathcal{N}(0, I)$", palette[2]),
]:
g = sns.jointplot(
x=data[:, 0], y=data[:, 1], kind="scatter", color=col,
height=7.5, ratio=5, space=0.1,
joint_kws={"s": 10, "alpha": 0.55},
marginal_kws={
"bins": 50, "color": col, "alpha": 0.75,
"edgecolor": "white",
},
xlim=(-2.5, 2.5), ylim=(-2.5, 2.5),
)
g.set_axis_labels("$x_1$", "$x_2$")
g.figure.suptitle(title, y=1.02)
style_jointgrid(g)
plt.show()/home/azureuser/.cache/uv/archive-v0/aYSeLZUlluhRY4DCBNG7F/lib/python3.13/site-packages/seaborn/axisgrid.py:1766: UserWarning: The figure layout has changed to tight
f.tight_layout()
/home/azureuser/.cache/uv/archive-v0/aYSeLZUlluhRY4DCBNG7F/lib/python3.13/site-packages/seaborn/axisgrid.py:1766: UserWarning: The figure layout has changed to tight
f.tight_layout()
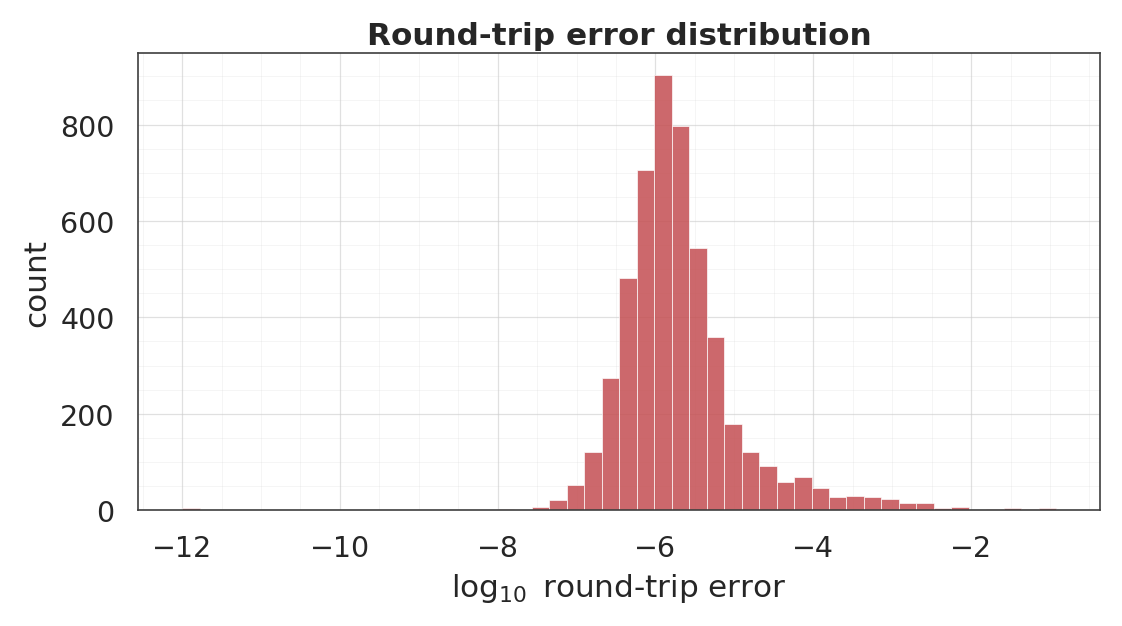
6. Round-trip inversion¶
Each coupling layer is a bijection by construction, so should recover to numerical precision.
x_t = ops.convert_to_tensor(X)
z_fwd = flow(x_t)
x_rt = ops.convert_to_numpy(flow.invert(z_fwd))
err = np.linalg.norm(x_rt - X, axis=-1)
print("round-trip L2 error ||f^{-1}(f(x)) - x||_2")
print(f" median : {np.median(err):.2e}")
print(f" 95th% : {np.percentile(err, 95):.2e}")
print(f" max : {err.max():.2e}")
fig, ax = plt.subplots(figsize=(10, 5.5))
ax.hist(
np.log10(err + 1e-12), bins=50, color=palette[3],
alpha=0.85, edgecolor="white", linewidth=0.5,
)
ax.set_xlabel("$\\log_{10}$ round-trip error")
ax.set_ylabel("count")
ax.set_title("Round-trip error distribution")
style_axes(ax)
plt.show()round-trip L2 error ||f^{-1}(f(x)) - x||_2
median : 1.55e-06
95th% : 7.06e-05
max : 1.20e-01
7. Layer-by-layer view¶
States . Each coupling layer transforms one half while keeping the other intact (you can see the passthrough half pinned, the other half morphing). Rotations remix the state between coupling pairs.
states = flow.forward_with_intermediates(ops.convert_to_tensor(X[:2000]))
states_np = [ops.convert_to_numpy(s) for s in states]
n_states = len(states_np)
ncol = 4
nrow = int(np.ceil(n_states / ncol))
fig, axes = plt.subplots(nrow, ncol, figsize=(4.5 * ncol, 4.5 * nrow))
axes = axes.ravel()
layer_names = ["input $x$"] + [
b.__class__.__name__ for b in flow.bijector_layers
]
for i, (state, ax) in enumerate(zip(states_np, axes)):
ax.scatter(state[:, 0], state[:, 1], s=5, alpha=0.6, color=palette[0])
lim = 3.4 if i > 0 else 2.8
ax.set_xlim(-lim, lim)
ax.set_ylim(-lim, lim)
if i == n_states - 1:
ax.set_title("Output $z$")
else:
ax.set_title(f"{i}: {layer_names[i]}")
style_axes(ax, aspect="equal")
for ax in axes[n_states:]:
ax.axis("off")
plt.show()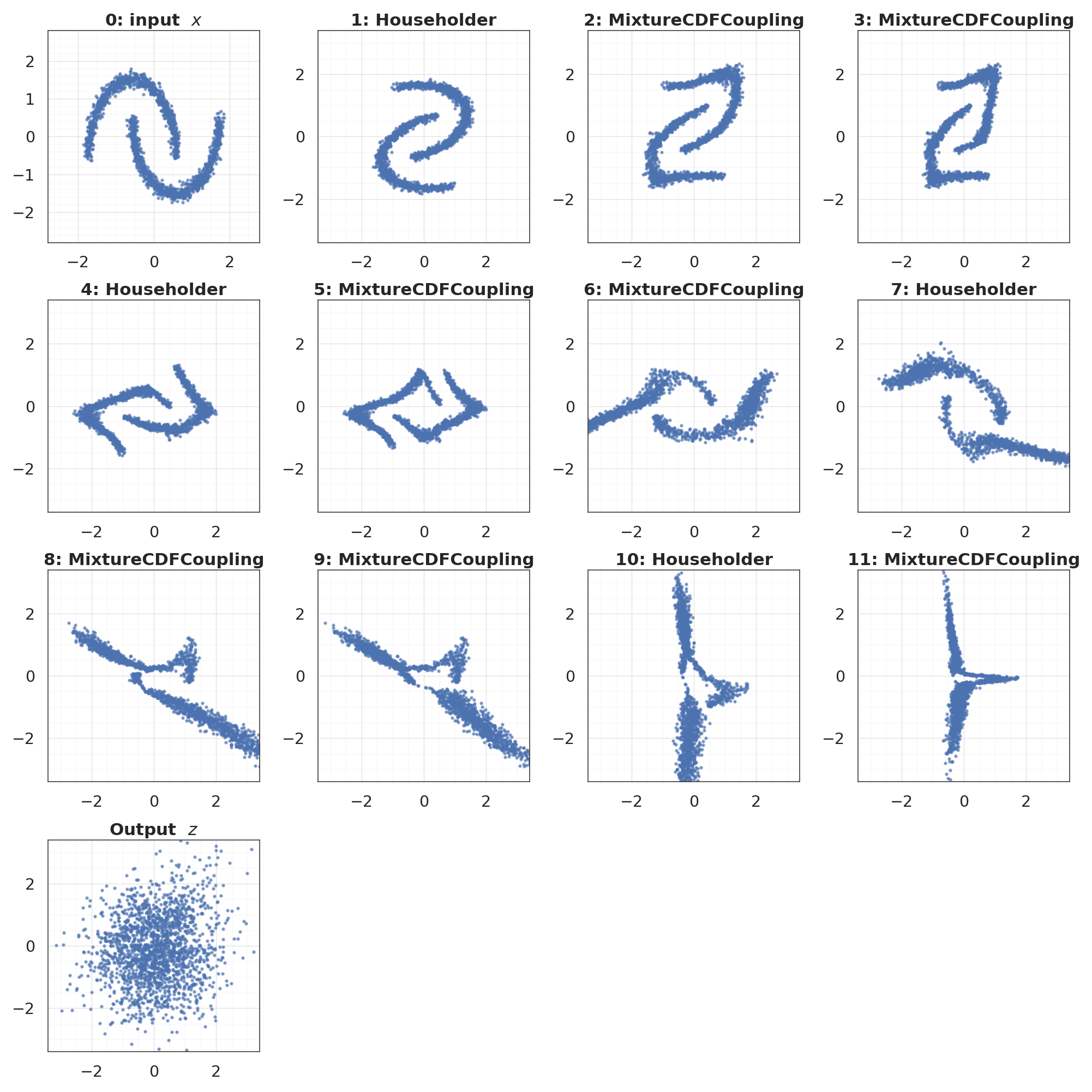
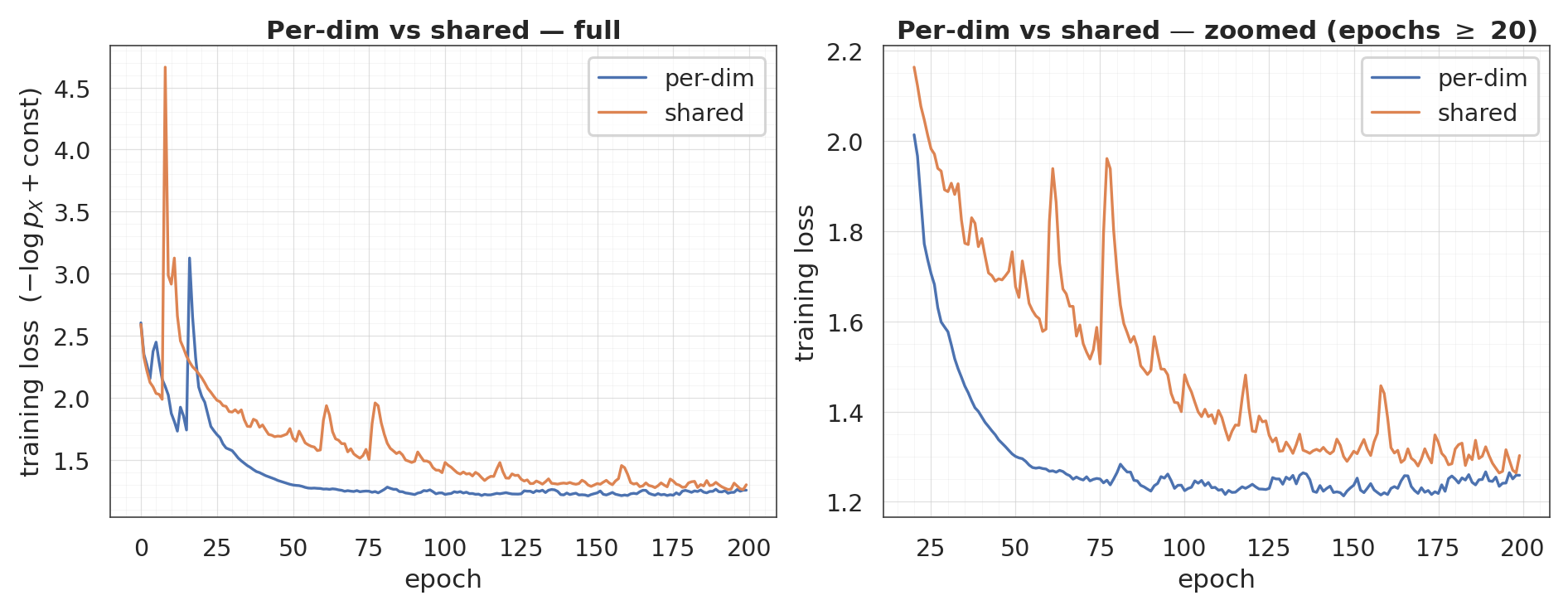
8. Shared-mixture conditioner¶
The per-dim conditioner emits floats — one mixture per transformed dimension. A shared-mixture conditioner emits only floats and reuses (tiles) those parameters across all dims. It is cheaper but strictly less expressive: every b-dim sees the same mixture given the same .
For two-moons (, ) the two are equivalent (only one b-dim per mask anyway). To see the difference we’d want ; here we just verify it trains and recovers the same density to similar fidelity.
flow_shared = make_coupling_flow(
input_dim=2,
num_blocks=4,
num_components=8,
hidden=(64, 64),
shared_mixture=True,
)
_ = flow_shared(ops.convert_to_tensor(X[:4]))
n_shared_params = int(sum(np.prod(w.shape) for w in flow_shared.trainable_weights))
print(f"shared-mixture flow trainable params : {n_shared_params} "
f"(per-dim reference : {n_params})")
flow_shared.compile(
optimizer=keras.optimizers.Adam(learning_rate=3e-3),
loss=base_nll_loss,
)
history_shared = flow_shared.fit(X, X, batch_size=512, epochs=200, verbose=0)
shared_curve = np.asarray(history_shared.history["loss"])
fig, axes = plt.subplots(1, 2, figsize=(17, 6.5))
ax = axes[0]
ax.plot(loss_curve, label="per-dim", color=palette[0], linewidth=2.2)
ax.plot(shared_curve, label="shared", color=palette[1], linewidth=2.2)
ax.set_xlabel("epoch")
ax.set_ylabel("training loss $(-\\log p_X + \\text{const})$")
ax.set_title("Per-dim vs shared — full")
ax.legend(frameon=True)
style_axes(ax)
ax = axes[1]
tail_start = 20
both_tail = np.concatenate([loss_curve[tail_start:], shared_curve[tail_start:]])
span = both_tail.max() - both_tail.min()
ax.plot(range(tail_start, len(loss_curve)), loss_curve[tail_start:],
label="per-dim", color=palette[0], linewidth=2.2)
ax.plot(range(tail_start, len(shared_curve)), shared_curve[tail_start:],
label="shared", color=palette[1], linewidth=2.2)
ax.set_ylim(both_tail.min() - 0.05 * span, both_tail.max() + 0.05 * span)
ax.set_xlabel("epoch")
ax.set_ylabel("training loss")
ax.set_title(f"Per-dim vs shared — zoomed (epochs $\\geq$ {tail_start})")
ax.legend(frameon=True)
style_axes(ax)
plt.show()shared-mixture flow trainable params : 46800 (per-dim reference : 46800)
log_px_shared = ops.convert_to_numpy(
flow_shared.log_prob(ops.convert_to_tensor(pts))
).reshape(grid, grid)
fig, ax = plt.subplots(figsize=(8.5, 7))
pcm = ax.pcolormesh(xx, yy, np.exp(log_px_shared), cmap="magma", shading="auto")
ax.scatter(X[:1000, 0], X[:1000, 1], s=2, alpha=0.25, color="white")
ax.set_title("Shared-mixture density $p_X(x)$")
ax.set_xlabel("$x_1$")
ax.set_ylabel("$x_2$")
style_axes(ax, aspect="equal")
fig.colorbar(pcm, ax=ax, shrink=0.85)
plt.show()
samples_shared = ops.convert_to_numpy(flow_shared.sample(num_samples=5000, seed=2))
g = sns.jointplot(
x=samples_shared[:, 0], y=samples_shared[:, 1], kind="scatter", color=palette[2],
height=7.5, ratio=5, space=0.1,
joint_kws={"s": 10, "alpha": 0.55},
marginal_kws={"bins": 50, "color": palette[2], "alpha": 0.75, "edgecolor": "white"},
xlim=(-2.5, 2.5), ylim=(-2.5, 2.5),
)
g.set_axis_labels("$x_1$", "$x_2$")
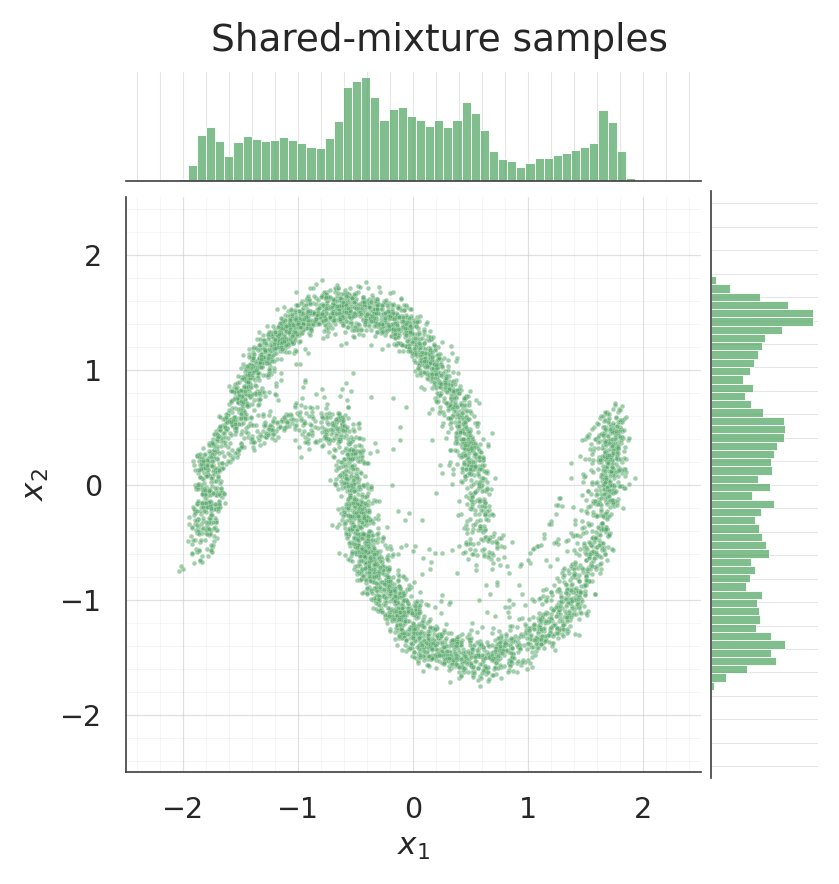
g.figure.suptitle("Shared-mixture samples", y=1.02)
style_jointgrid(g)
plt.show()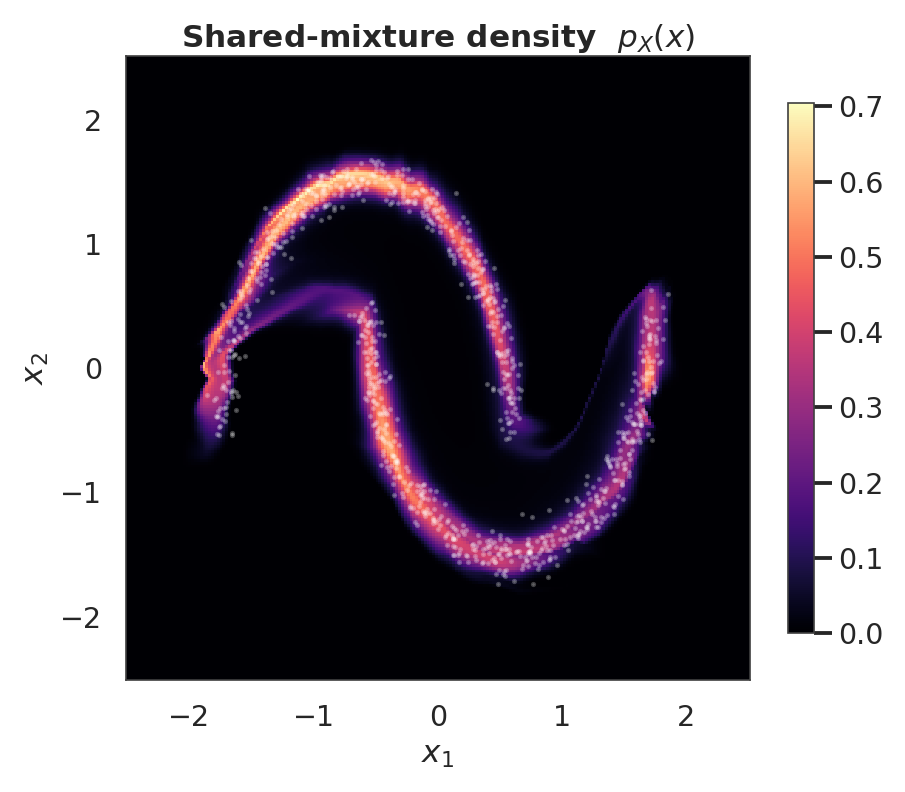
/home/azureuser/.cache/uv/archive-v0/aYSeLZUlluhRY4DCBNG7F/lib/python3.13/site-packages/seaborn/axisgrid.py:1766: UserWarning: The figure layout has changed to tight
f.tight_layout()
9. Custom conditioner — bring your own (residual network)¶
The coupling layer accepts any Keras layer/model whose output width is exactly . The build-time correctness check enforces this and raises a descriptive error otherwise. Below is the residual conditioner from the math section: one tanh hidden block with a skip connection, plus a zero-initialised output head so the layer starts at identity.
from gaussianization.gauss_keras import MixtureCDFCoupling, Householder
from gaussianization.gauss_keras.bijectors import GaussianizationFlow
D, D_B, K = 2, 1, 8
def make_residual_conditioner(d_b, num_components, hidden_width=32):
inp = keras.layers.Input(shape=(D - d_b,))
h = keras.layers.Dense(hidden_width, activation="tanh")(inp)
h = keras.layers.Dense(hidden_width, activation="tanh")(h) + h
out = keras.layers.Dense(
3 * d_b * num_components,
kernel_initializer="zeros",
bias_initializer="zeros",
)(h)
return keras.Model(inp, out)
mask = np.array([True, False])
custom_layers = []
for _ in range(4):
custom_layers.append(Householder(num_reflectors=2))
custom_layers.append(
MixtureCDFCoupling(
mask=mask,
conditioner=make_residual_conditioner(d_b=1, num_components=K),
num_components=K,
)
)
custom_layers.append(
MixtureCDFCoupling(
mask=~mask,
conditioner=make_residual_conditioner(d_b=1, num_components=K),
num_components=K,
)
)
flow_custom = GaussianizationFlow(custom_layers, input_dim=D)
_ = flow_custom(ops.convert_to_tensor(X[:4]))
flow_custom.compile(
optimizer=keras.optimizers.Adam(learning_rate=3e-3),
loss=base_nll_loss,
)
_ = flow_custom.fit(X, X, batch_size=512, epochs=100, verbose=0)
log_px_custom = ops.convert_to_numpy(
flow_custom.log_prob(ops.convert_to_tensor(pts))
).reshape(grid, grid)
fig, ax = plt.subplots(figsize=(8.5, 7))
pcm = ax.pcolormesh(xx, yy, np.exp(log_px_custom), cmap="magma", shading="auto")
ax.scatter(X[:1000, 0], X[:1000, 1], s=2, alpha=0.25, color="white")
ax.set_title("Custom-conditioner density $p_X(x)$")
ax.set_xlabel("$x_1$")
ax.set_ylabel("$x_2$")
style_axes(ax, aspect="equal")
fig.colorbar(pcm, ax=ax, shrink=0.85)
plt.show()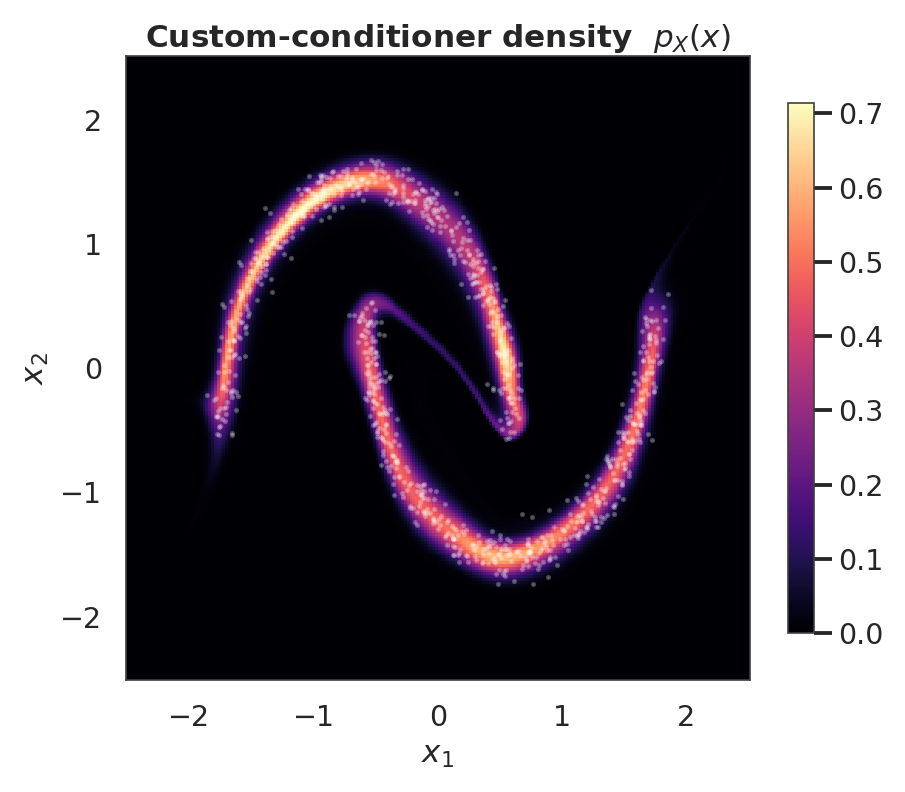
Recap¶
- Coupling = one half conditions, the other is transformed. Mask-swap in the same block so both halves get touched.
- The same
MixtureOfGaussiansprimitive drives both the unconditional marginal layer (notebook 01) and this layer — only difference is that the params now come from a network output instead of a learned per-dim lookup. - Zero-initialising the conditioner’s final Dense makes training stable from step 0 (the flow starts as the identity).
make_mlp_conditioner(per-dim) vsmake_shared_mlp_conditioner(tiled) trade off capacity against parameter count; custom architectures plug in cleanly as long as they output floats.