IG warm-start init
Iterative-Gaussianization warm start¶
Notebooks 01 and 02 trained Gaussianization / coupling flows from random initialisation. That works, but spends the first hundreds of epochs pulling the flow from a near-identity map towards the data.
Iterative Gaussianization (RBIG — Laparra & Malo 2011) is a non-gradient, greedy procedure that warm-starts the flow by walking block-by-block through the architecture and fitting each block’s parameters directly:
- Rotation block — fit PCA on the current state ; push the eigenvectors into the rotation layer (for Householder this uses a QR decomposition to extract reflector vectors).
- Marginal block — fit a per-dim mixture-of-Gaussians on the rotated data with scikit-learn’s
GaussianMixture, and assign directly into theMixtureCDFGaussianizationlayer weights. - Coupling block — fit a per-b-dim GMM on the current ; zero the conditioner’s final Dense kernel and set its bias to the flattened GMM params. At initialisation the conditioner output is constant-in-, so the coupling layer acts like a diagonal marginal with the fitted mixture. Gradient training then lets the conditioner learn to modulate on .
Between blocks the current is propagated forward through the newly-parameterised bijector, so the next block sees data that has already been partially Gaussianized. A classical result (Laparra & Malo 2011, Chen & Gopinath 2000) is that this drives towards at a geometric rate — even with no gradient descent at all, the flow is already a decent density estimator after a handful of blocks.
The point of this notebook: a Keras flow initialised this way starts at a much lower NLL than random init and reaches its training optimum in a fraction of the epochs.
import os
os.environ["KERAS_BACKEND"] = "tensorflow"
import keras
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from keras import ops
from sklearn.datasets import make_moons
from gaussianization.gauss_keras import (
base_nll_loss,
initialize_flow_from_ig,
make_coupling_flow,
make_gaussianization_flow,
)
# --- global plot styling ------------------------------------------------------
sns.set_theme(context="poster", style="whitegrid", palette="deep", font_scale=0.85)
plt.rcParams.update(
{
"figure.dpi": 110,
"savefig.dpi": 110,
"savefig.bbox": "tight",
"savefig.pad_inches": 0.2,
"figure.constrained_layout.use": True,
"figure.constrained_layout.h_pad": 0.1,
"figure.constrained_layout.w_pad": 0.1,
"axes.grid.which": "both",
"grid.linewidth": 0.7,
"grid.alpha": 0.5,
"axes.edgecolor": "0.25",
"axes.linewidth": 1.1,
"axes.titleweight": "semibold",
"axes.labelpad": 6,
}
)
def style_axes(ax, *, aspect=None, grid=True):
if grid:
ax.minorticks_on()
ax.grid(True, which="major", linewidth=0.8, alpha=0.6)
ax.grid(True, which="minor", linewidth=0.4, alpha=0.3)
if aspect is not None:
ax.set_aspect(aspect)
return ax
def style_jointgrid(g, aspect="equal"):
style_axes(g.ax_joint, aspect=aspect)
style_axes(g.ax_marg_x, grid=False)
style_axes(g.ax_marg_y, grid=False)
for spine in ("top", "right"):
g.ax_marg_x.spines[spine].set_visible(False)
g.ax_marg_y.spines[spine].set_visible(False)
palette = sns.color_palette("deep")
COLOR_RANDOM = palette[3]
COLOR_IG = palette[2]
keras.utils.set_random_seed(0)
np.random.seed(0)WARNING: All log messages before absl::InitializeLog() is called are written to STDERR
I0000 00:00:1776868596.826946 386554 cpu_feature_guard.cc:227] This TensorFlow binary is optimized to use available CPU instructions in performance-critical operations.
To enable the following instructions: AVX2 AVX512F FMA, in other operations, rebuild TensorFlow with the appropriate compiler flags.
1. Dataset¶
Two moons, standardised. Same data as notebooks 01/02 so the comparison is apples-to-apples.
X_raw, _ = make_moons(n_samples=5000, noise=0.05, random_state=0)
X = (X_raw - X_raw.mean(axis=0)) / X_raw.std(axis=0)
X = X.astype("float32")2. Diagonal flow — random vs IG init¶
Build two identical diagonal flows, warm-start one of them from IG, and train both with the same optimiser and schedule.
def build_diagonal_flow():
return make_gaussianization_flow(
input_dim=2,
num_blocks=8,
num_reflectors=2,
num_components=8,
pca_init_data=X,
mixture_init_data=X,
)
keras.utils.set_random_seed(1)
flow_random = build_diagonal_flow()
_ = flow_random(ops.convert_to_tensor(X[:4]))
keras.utils.set_random_seed(1)
flow_ig = build_diagonal_flow()
_ = flow_ig(ops.convert_to_tensor(X[:4]))
initialize_flow_from_ig(flow_ig, X)E0000 00:00:1776868601.498397 386554 cuda_platform.cc:52] failed call to cuInit: INTERNAL: CUDA error: Failed call to cuInit: CUDA_ERROR_NO_DEVICE: no CUDA-capable device is detected
array([[ 0.21370773, 0.68232536],
[-1.675768 , -0.09274741],
[ 0.5233947 , 1.6713623 ],
...,
[ 1.3039094 , -0.8307837 ],
[ 0.17580122, -0.15649901],
[ 0.17348166, -0.39425728]], shape=(5000, 2), dtype=float32)Before any training: what did IG init already buy us?¶
RBIG on its own produces a density estimate. Below we compare the mean log-likelihood (per sample) of the IG-initialised flow against the random-init flow, both before a single gradient step.
lp_random0 = float(np.mean(ops.convert_to_numpy(
flow_random.log_prob(ops.convert_to_tensor(X))
)))
lp_ig0 = float(np.mean(ops.convert_to_numpy(
flow_ig.log_prob(ops.convert_to_tensor(X))
)))
print(f"mean log-likelihood before training:")
print(f" random init : {lp_random0:+.3f} nats/sample")
print(f" IG init : {lp_ig0:+.3f} nats/sample (improvement: {lp_ig0 - lp_random0:+.3f})")mean log-likelihood before training:
random init : -881.520 nats/sample
IG init : -1.652 nats/sample (improvement: +879.868)
Pushforward at IG-init time¶
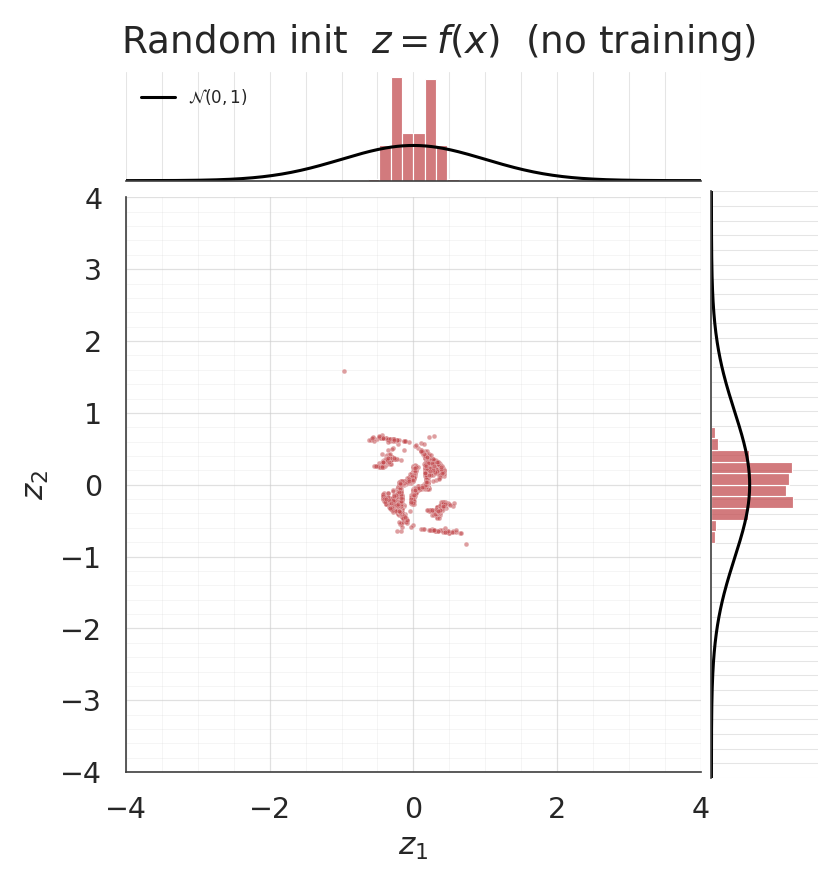
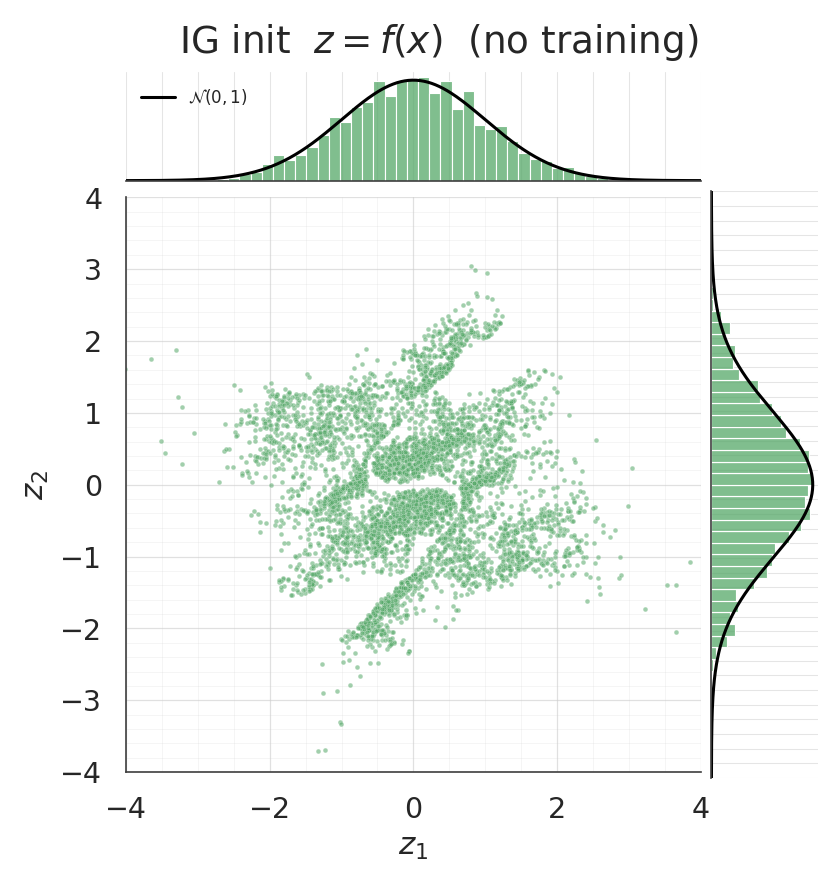
With no gradient training whatsoever, the pushforward of the IG-initialised flow should already be approximately . Jointplots make this immediately visible — the IG scatter should look isotropic and the marginals should line up with the black curves, while the random-init pushforward won’t.
z_ig0 = ops.convert_to_numpy(flow_ig(ops.convert_to_tensor(X)))
z_rnd0 = ops.convert_to_numpy(flow_random(ops.convert_to_tensor(X)))
zz = np.linspace(-4, 4, 300)
phi = np.exp(-0.5 * zz**2) / np.sqrt(2 * np.pi)
for z_sample, title, col in [
(z_rnd0, "Random init $z = f(x)$ (no training)", COLOR_RANDOM),
(z_ig0, "IG init $z = f(x)$ (no training)", COLOR_IG),
]:
g = sns.jointplot(
x=z_sample[:, 0], y=z_sample[:, 1], kind="scatter", color=col,
height=7.5, ratio=5, space=0.1,
joint_kws={"s": 10, "alpha": 0.55},
marginal_kws={
"bins": 60, "color": col, "alpha": 0.75,
"edgecolor": "white", "stat": "density",
},
xlim=(-4, 4), ylim=(-4, 4),
)
g.ax_marg_x.plot(zz, phi, color="black", linewidth=2.0, label="$\\mathcal{N}(0, 1)$")
g.ax_marg_y.plot(phi, zz, color="black", linewidth=2.0)
g.ax_marg_x.legend(loc="upper left", frameon=False, fontsize=11)
g.set_axis_labels("$z_1$", "$z_2$")
g.figure.suptitle(title, y=1.02)
style_jointgrid(g)
plt.show()
mean_ig0 = z_ig0.mean(axis=0)
cov_ig0 = np.cov(z_ig0, rowvar=False)
print("IG-init pushforward (pre-training)")
print(f" mean: [{mean_ig0[0]:+.3f}, {mean_ig0[1]:+.3f}] target = [0, 0]")
print(f" cov[0,0]:{cov_ig0[0, 0]:.3f} cov[1,1]:{cov_ig0[1, 1]:.3f} target = 1, 1")
print(f" cov[0,1]:{cov_ig0[0, 1]:+.3f} target = 0")/home/azureuser/.cache/uv/archive-v0/aYSeLZUlluhRY4DCBNG7F/lib/python3.13/site-packages/seaborn/axisgrid.py:1766: UserWarning: The figure layout has changed to tight
f.tight_layout()
/home/azureuser/.cache/uv/archive-v0/aYSeLZUlluhRY4DCBNG7F/lib/python3.13/site-packages/seaborn/axisgrid.py:1766: UserWarning: The figure layout has changed to tight
f.tight_layout()
IG-init pushforward (pre-training)
mean: [+0.000, -0.000] target = [0, 0]
cov[0,0]:1.003 cov[1,1]:1.006 target = 1, 1
cov[0,1]:+0.004 target = 0
Training comparison¶
Both flows, same optimiser, same 150 epochs. The IG curve starts at (and finishes near) the random curve’s final value — the zoomed panel on the right makes the small end-of-training gap legible.
flow_random.compile(optimizer=keras.optimizers.Adam(5e-3), loss=base_nll_loss)
flow_ig.compile(optimizer=keras.optimizers.Adam(5e-3), loss=base_nll_loss)
hist_random = flow_random.fit(X, X, batch_size=512, epochs=150, verbose=0)
hist_ig = flow_ig.fit(X, X, batch_size=512, epochs=150, verbose=0)
rnd_curve = np.asarray(hist_random.history["loss"])
ig_curve = np.asarray(hist_ig.history["loss"])
fig, axes = plt.subplots(1, 2, figsize=(17, 6.5))
ax = axes[0]
ax.plot(rnd_curve, label="random init", color=COLOR_RANDOM, linewidth=2.2)
ax.plot(ig_curve, label="IG init", color=COLOR_IG, linewidth=2.2)
ax.set_xlabel("epoch")
ax.set_ylabel("training loss $(-\\log p_X + \\text{const})$")
ax.set_title("Diagonal flow — full")
ax.legend(frameon=True)
style_axes(ax)
ax = axes[1]
tail_start = 15
tail_rnd = rnd_curve[tail_start:]
tail_ig = ig_curve[tail_start:]
both = np.concatenate([tail_rnd, tail_ig])
span = both.max() - both.min()
ax.plot(range(tail_start, len(rnd_curve)), tail_rnd,
label="random init", color=COLOR_RANDOM, linewidth=2.2)
ax.plot(range(tail_start, len(ig_curve)), tail_ig,
label="IG init", color=COLOR_IG, linewidth=2.2)
ax.set_ylim(both.min() - 0.05 * span, both.max() + 0.05 * span)
ax.set_xlabel("epoch")
ax.set_ylabel("training loss")
ax.set_title(f"Zoomed (epochs $\\geq$ {tail_start})")
ax.legend(frameon=True)
style_axes(ax)
plt.show()
print(f"final loss: random init = {rnd_curve[-1]:+.3f} "
f"IG init = {ig_curve[-1]:+.3f}")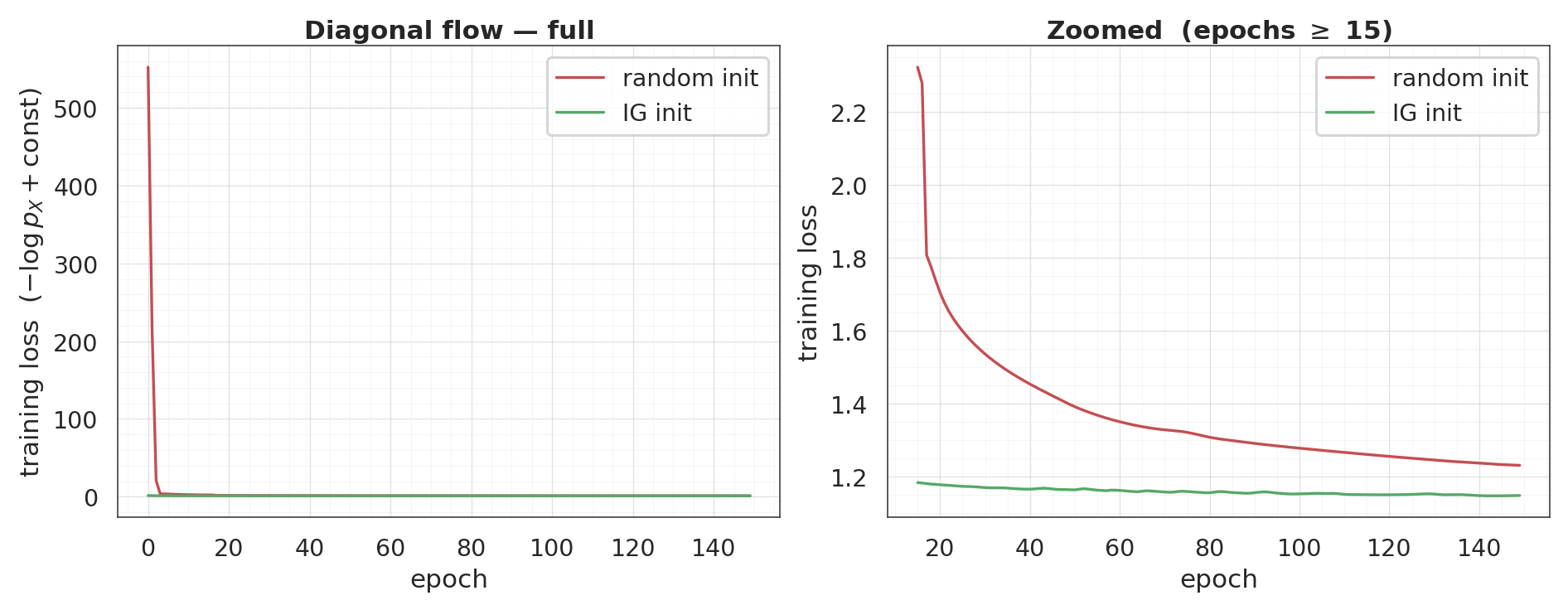
final loss: random init = +1.232 IG init = +1.149
Density and samples after training¶
grid = 220
xs = np.linspace(-2.5, 2.5, grid).astype("float32")
ys = np.linspace(-2.5, 2.5, grid).astype("float32")
xx, yy = np.meshgrid(xs, ys)
pts = np.stack([xx.ravel(), yy.ravel()], axis=-1).astype("float32")
log_px_rnd = ops.convert_to_numpy(
flow_random.log_prob(ops.convert_to_tensor(pts))
).reshape(grid, grid)
log_px_ig = ops.convert_to_numpy(
flow_ig.log_prob(ops.convert_to_tensor(pts))
).reshape(grid, grid)
fig, axes = plt.subplots(1, 2, figsize=(17, 7.5))
for ax, lp, title in zip(
axes, [log_px_rnd, log_px_ig], ["Random init — density", "IG init — density"]
):
pcm = ax.pcolormesh(xx, yy, np.exp(lp), cmap="magma", shading="auto")
ax.scatter(X[:800, 0], X[:800, 1], s=2, alpha=0.25, color="white")
ax.set_title(title)
ax.set_xlabel("$x_1$")
ax.set_ylabel("$x_2$")
style_axes(ax, aspect="equal")
fig.colorbar(pcm, ax=ax, shrink=0.85)
plt.show()
samples_rnd = ops.convert_to_numpy(flow_random.sample(num_samples=3000, seed=2))
samples_ig = ops.convert_to_numpy(flow_ig.sample(num_samples=3000, seed=2))
for s, title, col in [
(samples_rnd, "Random init — samples", COLOR_RANDOM),
(samples_ig, "IG init — samples", COLOR_IG),
]:
g = sns.jointplot(
x=s[:, 0], y=s[:, 1], kind="scatter", color=col,
height=7.5, ratio=5, space=0.1,
joint_kws={"s": 10, "alpha": 0.55},
marginal_kws={"bins": 50, "color": col, "alpha": 0.75, "edgecolor": "white"},
xlim=(-2.5, 2.5), ylim=(-2.5, 2.5),
)
g.set_axis_labels("$x_1$", "$x_2$")
g.figure.suptitle(title, y=1.02)
style_jointgrid(g)
plt.show()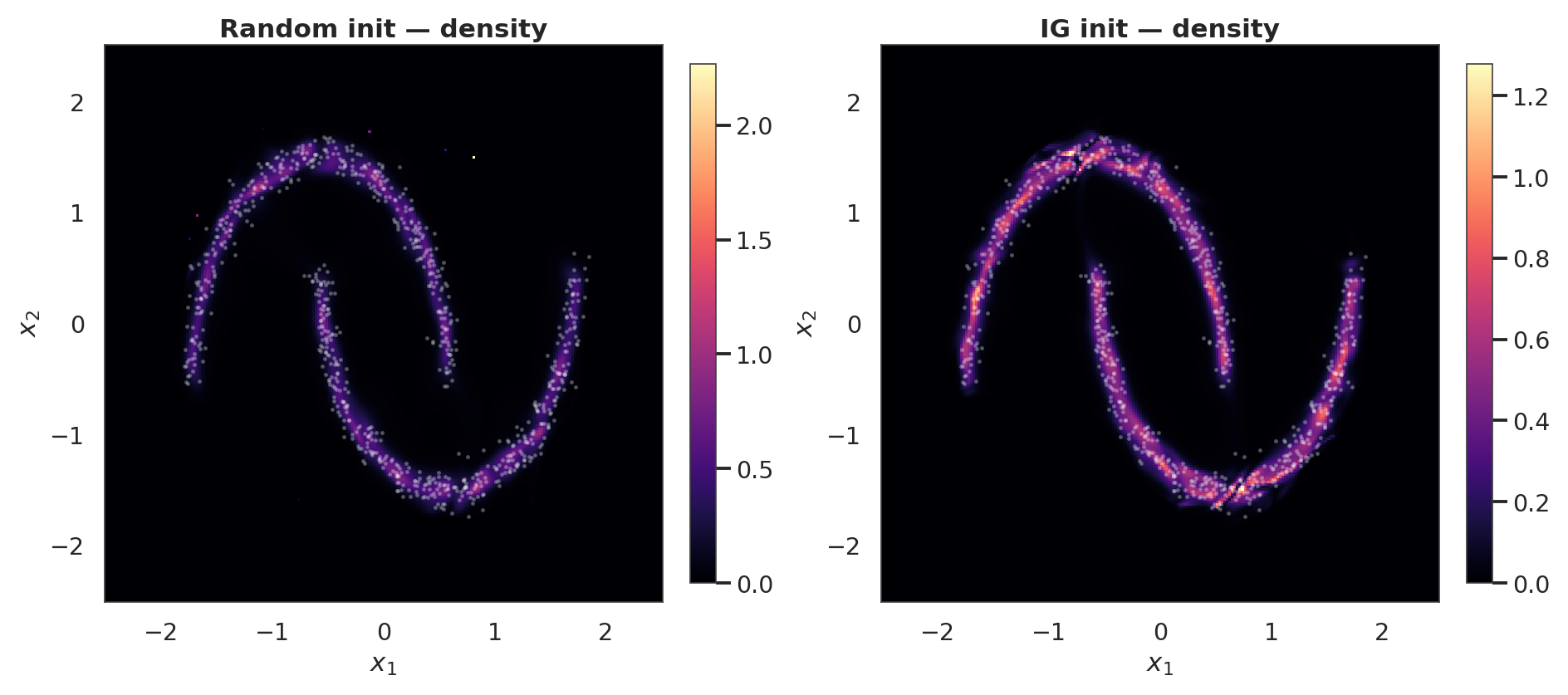
/home/azureuser/.cache/uv/archive-v0/aYSeLZUlluhRY4DCBNG7F/lib/python3.13/site-packages/seaborn/axisgrid.py:1766: UserWarning: The figure layout has changed to tight
f.tight_layout()
/home/azureuser/.cache/uv/archive-v0/aYSeLZUlluhRY4DCBNG7F/lib/python3.13/site-packages/seaborn/axisgrid.py:1766: UserWarning: The figure layout has changed to tight
f.tight_layout()
3. Coupling flow — random vs IG init¶
Same experiment with a coupling flow. IG init for coupling layers is weaker in isolation (the conditioner is not exercised at init — it emits a constant bias), but it still lets training skip the “the conditioner is effectively random, so the coupling transform is random” phase by handing it a sensible starting point where the conditioner behaves as a well-fit diagonal marginal.
def build_coupling_flow():
return make_coupling_flow(
input_dim=2,
num_blocks=4,
num_components=8,
hidden=(64, 64),
)
keras.utils.set_random_seed(3)
cpl_random = build_coupling_flow()
_ = cpl_random(ops.convert_to_tensor(X[:4]))
keras.utils.set_random_seed(3)
cpl_ig = build_coupling_flow()
_ = cpl_ig(ops.convert_to_tensor(X[:4]))
initialize_flow_from_ig(cpl_ig, X)array([[ 0.33592415, -0.63961166],
[-1.7532271 , -0.11788122],
[ 0.52066326, -1.6266781 ],
...,
[ 1.1722953 , 0.8738398 ],
[ 0.15655102, 0.20274866],
[ 0.06255252, 0.3414428 ]], shape=(5000, 2), dtype=float32)cpl_random.compile(optimizer=keras.optimizers.Adam(3e-3), loss=base_nll_loss)
cpl_ig.compile(optimizer=keras.optimizers.Adam(3e-3), loss=base_nll_loss)
hist_c_random = cpl_random.fit(X, X, batch_size=512, epochs=100, verbose=0)
hist_c_ig = cpl_ig.fit(X, X, batch_size=512, epochs=100, verbose=0)
crnd_curve = np.asarray(hist_c_random.history["loss"])
cig_curve = np.asarray(hist_c_ig.history["loss"])
fig, axes = plt.subplots(1, 2, figsize=(17, 6.5))
ax = axes[0]
ax.plot(crnd_curve, label="random init", color=COLOR_RANDOM, linewidth=2.2)
ax.plot(cig_curve, label="IG init", color=COLOR_IG, linewidth=2.2)
ax.set_xlabel("epoch")
ax.set_ylabel("training loss $(-\\log p_X + \\text{const})$")
ax.set_title("Coupling flow — full")
ax.legend(frameon=True)
style_axes(ax)
ax = axes[1]
tail_start = 10
tail_rnd = crnd_curve[tail_start:]
tail_ig = cig_curve[tail_start:]
both = np.concatenate([tail_rnd, tail_ig])
span = both.max() - both.min()
ax.plot(range(tail_start, len(crnd_curve)), tail_rnd,
label="random init", color=COLOR_RANDOM, linewidth=2.2)
ax.plot(range(tail_start, len(cig_curve)), tail_ig,
label="IG init", color=COLOR_IG, linewidth=2.2)
ax.set_ylim(both.min() - 0.05 * span, both.max() + 0.05 * span)
ax.set_xlabel("epoch")
ax.set_ylabel("training loss")
ax.set_title(f"Zoomed (epochs $\\geq$ {tail_start})")
ax.legend(frameon=True)
style_axes(ax)
plt.show()
print(f"final loss: random init = {crnd_curve[-1]:+.3f} "
f"IG init = {cig_curve[-1]:+.3f}")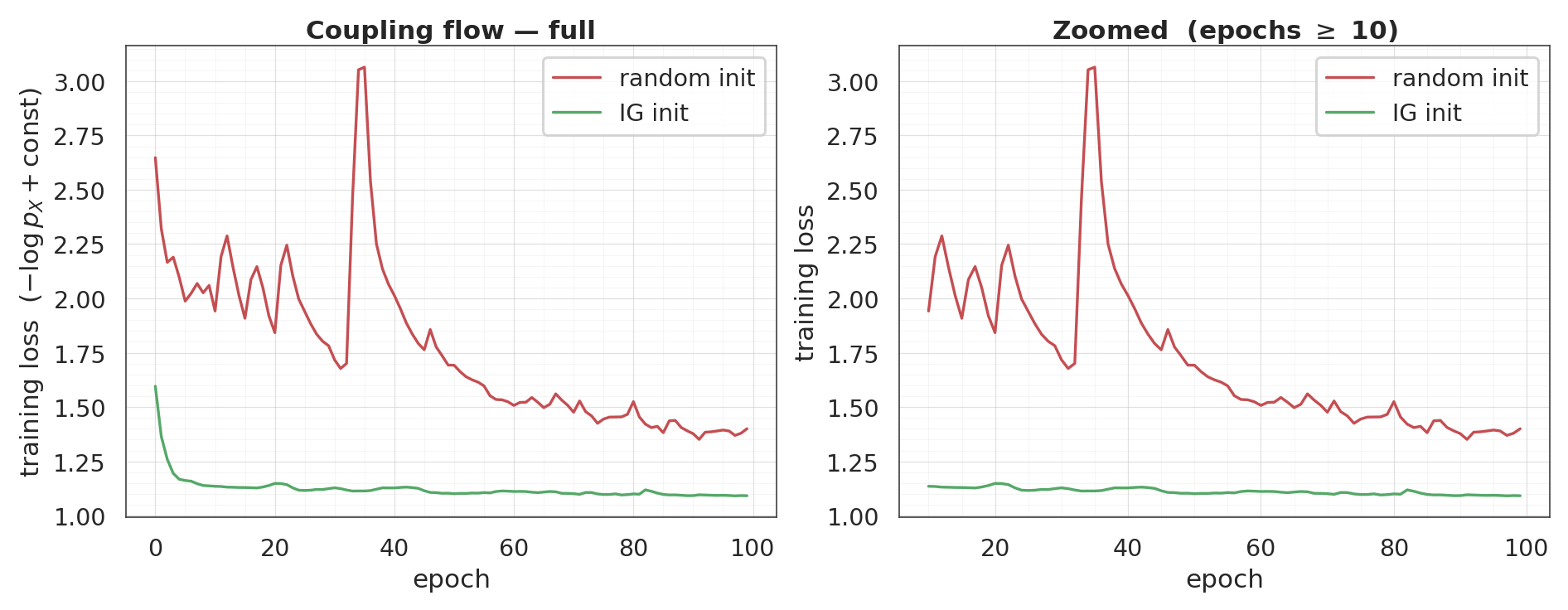
final loss: random init = +1.400 IG init = +1.092
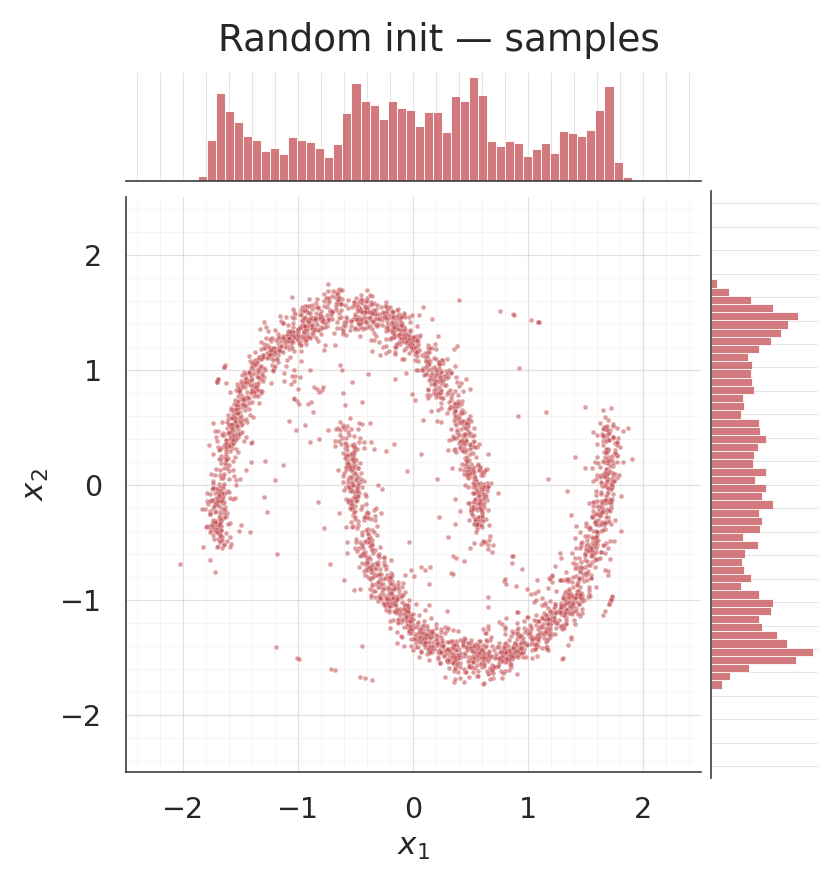
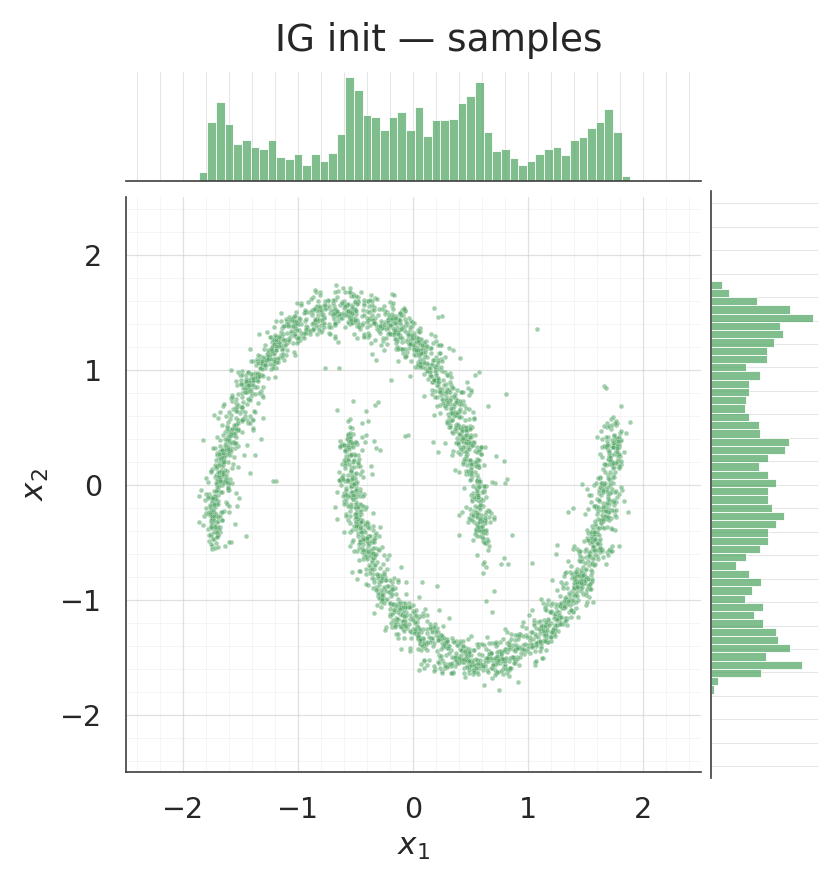
Coupling density and samples after training¶
log_px_crnd = ops.convert_to_numpy(
cpl_random.log_prob(ops.convert_to_tensor(pts))
).reshape(grid, grid)
log_px_cig = ops.convert_to_numpy(
cpl_ig.log_prob(ops.convert_to_tensor(pts))
).reshape(grid, grid)
fig, axes = plt.subplots(1, 2, figsize=(17, 7.5))
for ax, lp, title in zip(
axes, [log_px_crnd, log_px_cig], ["Coupling — random init", "Coupling — IG init"]
):
pcm = ax.pcolormesh(xx, yy, np.exp(lp), cmap="magma", shading="auto")
ax.scatter(X[:800, 0], X[:800, 1], s=2, alpha=0.25, color="white")
ax.set_title(title)
ax.set_xlabel("$x_1$")
ax.set_ylabel("$x_2$")
style_axes(ax, aspect="equal")
fig.colorbar(pcm, ax=ax, shrink=0.85)
plt.show()
csamples_rnd = ops.convert_to_numpy(cpl_random.sample(num_samples=3000, seed=4))
csamples_ig = ops.convert_to_numpy(cpl_ig.sample(num_samples=3000, seed=4))
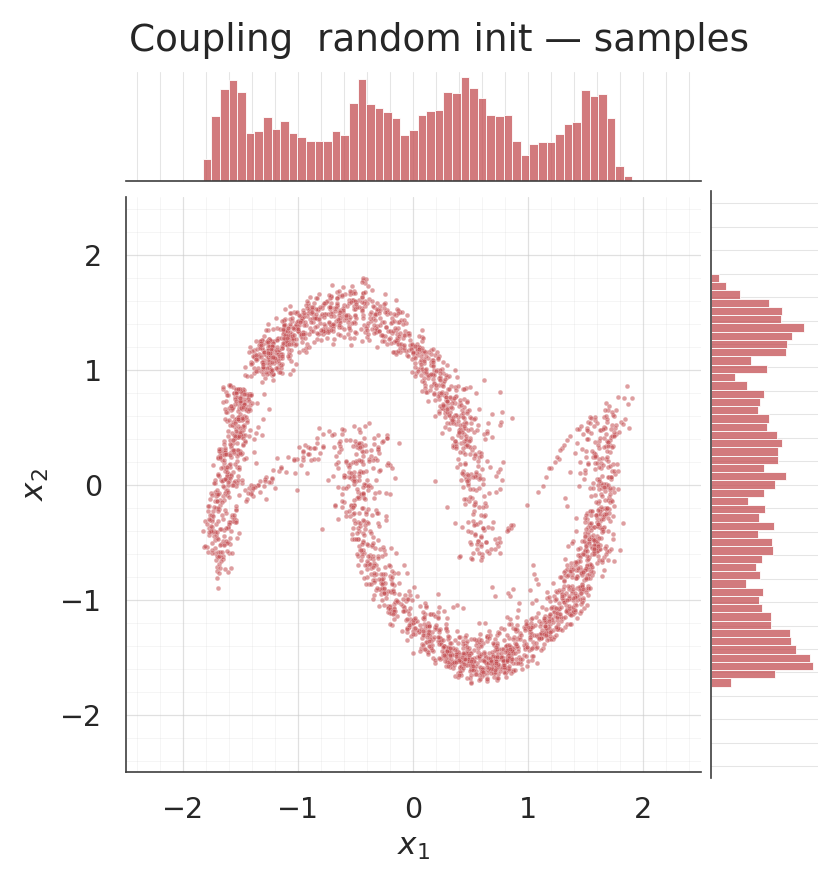
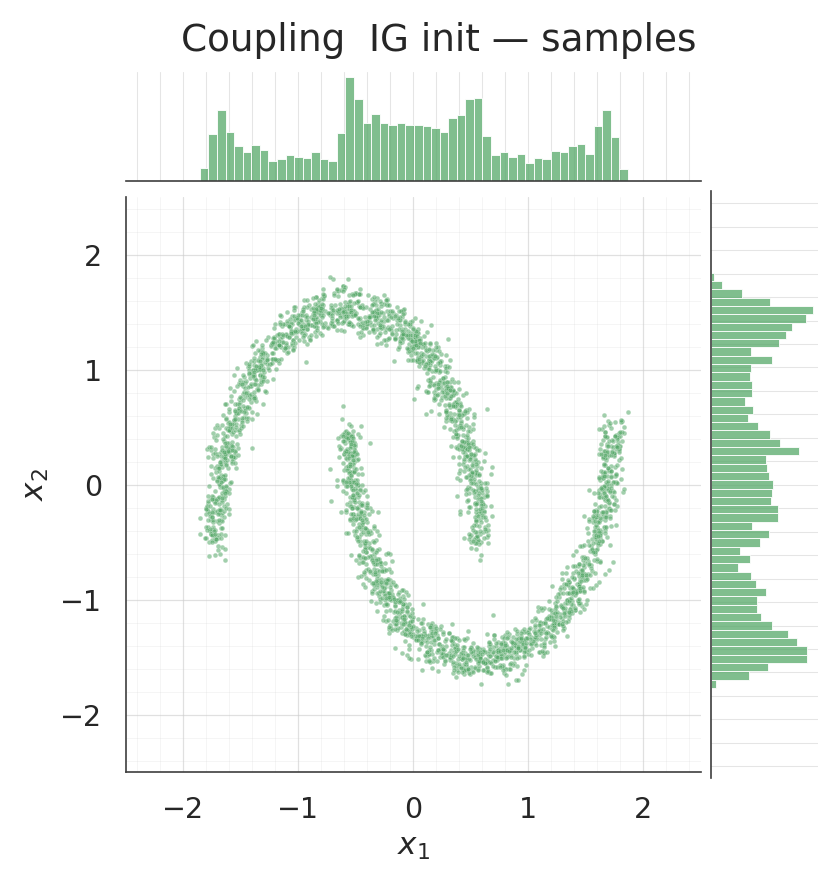
for s, title, col in [
(csamples_rnd, "Coupling random init — samples", COLOR_RANDOM),
(csamples_ig, "Coupling IG init — samples", COLOR_IG),
]:
g = sns.jointplot(
x=s[:, 0], y=s[:, 1], kind="scatter", color=col,
height=7.5, ratio=5, space=0.1,
joint_kws={"s": 10, "alpha": 0.55},
marginal_kws={"bins": 50, "color": col, "alpha": 0.75, "edgecolor": "white"},
xlim=(-2.5, 2.5), ylim=(-2.5, 2.5),
)
g.set_axis_labels("$x_1$", "$x_2$")
g.figure.suptitle(title, y=1.02)
style_jointgrid(g)
plt.show()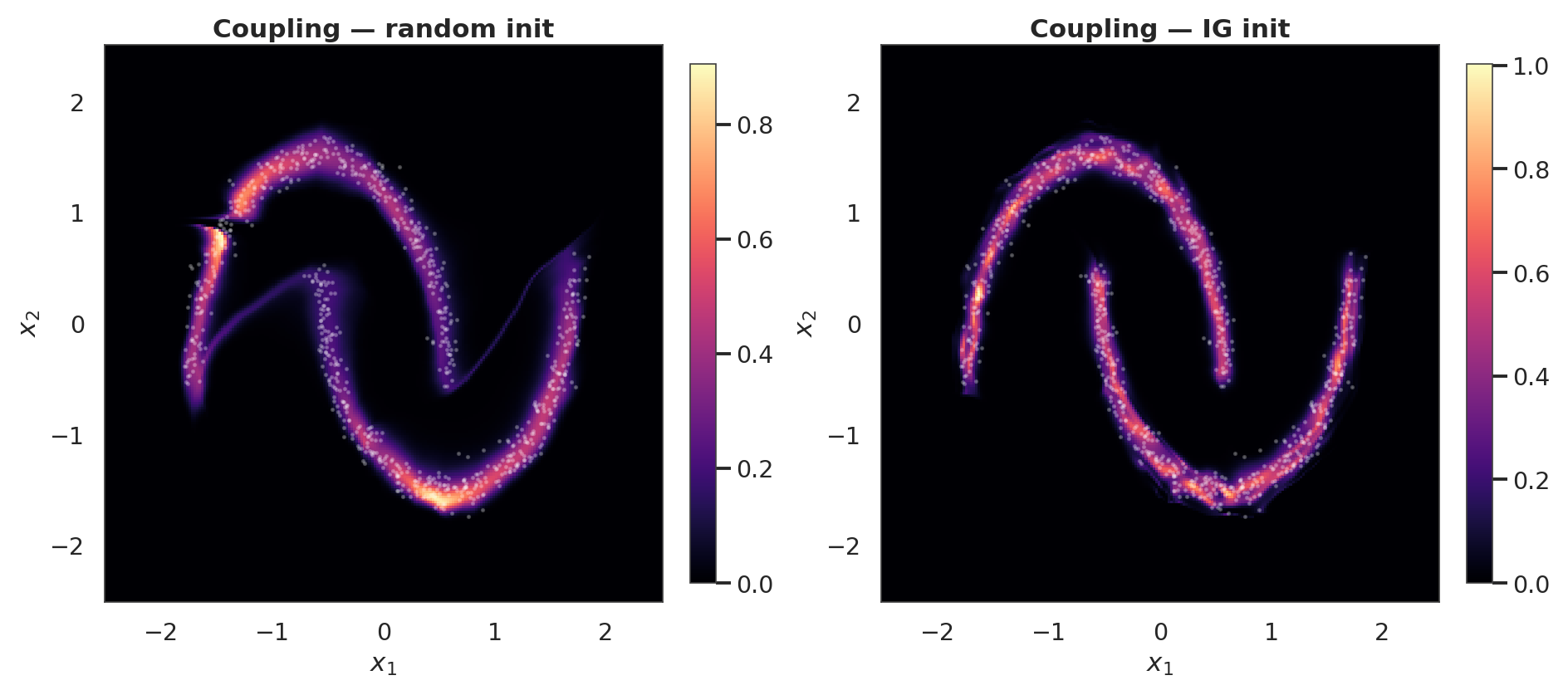
/home/azureuser/.cache/uv/archive-v0/aYSeLZUlluhRY4DCBNG7F/lib/python3.13/site-packages/seaborn/axisgrid.py:1766: UserWarning: The figure layout has changed to tight
f.tight_layout()
/home/azureuser/.cache/uv/archive-v0/aYSeLZUlluhRY4DCBNG7F/lib/python3.13/site-packages/seaborn/axisgrid.py:1766: UserWarning: The figure layout has changed to tight
f.tight_layout()
4. Verifying the zero-kernel contract¶
Immediately after initialize_flow_from_ig every MixtureCDFCoupling’s final Dense has a zero kernel (so the conditioner emits its bias for any input — the coupling layer behaves as a diagonal marginal). During gradient training those kernels become non-zero as the conditioner learns to modulate on .
Below: build a fresh coupling flow, IG-init it, check the kernels before any training, then train briefly and check again.
from gaussianization.gauss_keras.bijectors import MixtureCDFCoupling
def max_final_kernel(flow):
out = []
for b in flow.bijector_layers:
if isinstance(b, MixtureCDFCoupling):
last = None
for sub in b.conditioner.layers:
if hasattr(sub, "kernel") and hasattr(sub, "bias"):
last = sub
out.append(float(np.max(np.abs(ops.convert_to_numpy(last.kernel)))))
return out
keras.utils.set_random_seed(42)
cpl_probe = build_coupling_flow()
_ = cpl_probe(ops.convert_to_tensor(X[:4]))
kernels_before = max_final_kernel(cpl_probe)
print(f"fresh flow (before IG init): max|W| per coupling = {[f'{k:.3f}' for k in kernels_before]}")
initialize_flow_from_ig(cpl_probe, X)
kernels_post_ig = max_final_kernel(cpl_probe)
print(f"right after IG init (no train): max|W| per coupling = {[f'{k:.1e}' for k in kernels_post_ig]}")
print(f"all zero after IG init : {all(k < 1e-6 for k in kernels_post_ig)}")
cpl_probe.compile(optimizer=keras.optimizers.Adam(3e-3), loss=base_nll_loss)
_ = cpl_probe.fit(X, X, batch_size=512, epochs=20, verbose=0)
kernels_after_train = max_final_kernel(cpl_probe)
print(f"after 20 epochs of training: max|W| per coupling = {[f'{k:.3f}' for k in kernels_after_train]}")
print("(non-zero is expected — the conditioner has started to modulate on x_a)")fresh flow (before IG init): max|W| per coupling = ['0.000', '0.000', '0.000', '0.000', '0.000', '0.000', '0.000', '0.000']
right after IG init (no train): max|W| per coupling = ['0.0e+00', '0.0e+00', '0.0e+00', '0.0e+00', '0.0e+00', '0.0e+00', '0.0e+00', '0.0e+00']
all zero after IG init : True
after 20 epochs of training: max|W| per coupling = ['0.283', '0.394', '0.373', '0.376', '0.556', '0.181', '0.145', '0.226']
(non-zero is expected — the conditioner has started to modulate on x_a)
Recap¶
- RBIG (Laparra & Malo 2011) is a non-gradient greedy warm-start that fits each block’s parameters in sequence (PCA for rotation, sklearn
GaussianMixtureper dim for marginal/coupling) and propagates the data forward. - For the diagonal flow the init is exact: the layer’s parameters are exactly the GMM fit, and the pre-training pushforward is already near .
- For the coupling flow the conditioner’s inner Dense weights stay random, but the final Dense is forced to kernel=0, bias = fitted mixture params. So at init the coupling layer behaves like a diagonal marginal layer with the IG-fitted mixture; gradient training then lets the conditioner become genuinely conditional.
- End result in both cases: lower starting NLL, faster convergence, and fewer chances for the optimiser to get stuck in a bad local minimum.