Structured GP: Kronecker and Low-Rank
This notebook shows where gaussx shines: exploiting structure in kernel matrices for Gaussian process inference.
We cover two common patterns:
- Kronecker structure — GP on a grid (e.g. spatial data on a regular mesh)
- Low-rank + diagonal — sparse GP with inducing points (Nystrom approximation)
The two most common forms of kernel structure in GP models are (1) Kronecker products from separable kernels on grids, and (2) low-rank approximations from inducing point methods. Both reduce the cost of dense GP inference. gaussx supports both through its operator type system, dispatching to the optimal algorithm automatically.
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import jax
import jax.numpy as jnp
import lineax as lx
import matplotlib.pyplot as plt
import gaussx
jax.config.update("jax_enable_x64", True)Part 1: Kronecker GP on a grid¶
When input data lies on a grid, the kernel matrix factorizes as a Kronecker product: . This turns operations into where .
For a separable kernel on 2D inputs the factorization is:
This includes products of any 1D stationary kernels (e.g. RBF, Matern). See Saatci (2012) for a comprehensive treatment of Kronecker methods in GP models.
Setup: 2D grid data¶
def rbf_kernel_1d(x, lengthscale, variance):
"""1D RBF kernel matrix."""
sq_dist = (x[:, None] - x[None, :]) ** 2
return variance * jnp.exp(-0.5 * sq_dist / lengthscale**2)
# Grid dimensions
n1, n2 = 10, 12
N = n1 * n2 # Total points = 120
x1 = jnp.linspace(0, 5, n1)
x2 = jnp.linspace(0, 5, n2)
print(f"Grid: {n1} x {n2} = {N} points")
print(f"Dense kernel would be {N} x {N} = {N**2:,} entries")
print(f"Kronecker stores {n1}x{n1} + {n2}x{n2} = {n1**2 + n2**2} entries")Grid: 10 x 12 = 120 points
Dense kernel would be 120 x 120 = 14,400 entries
Kronecker stores 10x10 + 12x12 = 244 entries
Build the Kronecker kernel¶
# Per-dimension kernel matrices (lengthscale ~ grid spacing for good conditioning)
K1 = rbf_kernel_1d(x1, lengthscale=1.0, variance=1.0)
K2 = rbf_kernel_1d(x2, lengthscale=1.0, variance=1.0)
# Wrap as PSD lineax operators
K1_op = lx.MatrixLinearOperator(K1, lx.positive_semidefinite_tag)
K2_op = lx.MatrixLinearOperator(K2, lx.positive_semidefinite_tag)
# Kronecker product: K = K1 kron K2
K_kron = gaussx.Kronecker(K1_op, K2_op)
# Add noise: (K1 kron K2) + sigma^2 I
# For now, materialize noise addition — a full structured version
# would use a LowRankUpdate or custom operator.
noise_var = 0.01
print(f"Kronecker operator size: {K_kron.in_size()}")Kronecker operator size: 120
# Structured logdet: O(n1^3 + n2^3)
ld_structured = gaussx.logdet(K_kron)
print(f"logdet (structured): {ld_structured:.6f}")
# Verify against dense: O(N^3)
ld_dense = jnp.linalg.slogdet(K_kron.as_matrix())[1]
print(f"logdet (dense): {ld_dense:.6f}")
print(f"Match: {jnp.allclose(ld_structured, ld_dense)}")logdet (structured): -805.280930
logdet (dense): -805.280929
Match: True
Kronecker solve and Cholesky¶
Solve and Cholesky also decompose per-factor.
# Generate observations on the grid
key = jax.random.PRNGKey(0)
y = jax.random.normal(key, (N,))
# Structured solve of K_kron @ alpha = y
alpha = gaussx.solve(K_kron, y)
print(f"solve residual: {jnp.max(jnp.abs(K_kron.mv(alpha) - y)):.2e}")
# Structured Cholesky: returns Kronecker(chol(K1), chol(K2))
L = gaussx.cholesky(K_kron)
print(f"cholesky type: {type(L).__name__}")
print(f"L factors: {[type(f).__name__ for f in L.operators]}")
# Verify: L L^T = K
recon_err = jnp.max(jnp.abs(L.as_matrix() @ L.as_matrix().T - K_kron.as_matrix()))
print(f"Cholesky reconstruction error: {recon_err:.2e}")solve residual: 2.27e-07
cholesky type: Kronecker
L factors: ['MatrixLinearOperator', 'MatrixLinearOperator']
Cholesky reconstruction error: 4.44e-16
Part 2: Low-rank GP (Nystrom / inducing points)¶
For non-grid data, a common approximation is . The resulting covariance is — a low-rank update.
The Woodbury identity gives an efficient solve:
and the matrix determinant lemma gives an efficient logdet:
Both reduce the cost from to .
Setup: random 1D data with inducing points¶
n_data = 200
n_inducing = 15
key = jax.random.PRNGKey(7)
k1, k2 = jax.random.split(key)
x_data = jax.random.uniform(k1, (n_data,), minval=-3.0, maxval=3.0)
x_inducing = jnp.linspace(-2.5, 2.5, n_inducing)
print(f"Data points: {n_data}")
print(f"Inducing points: {n_inducing}")
print(f"Dense kernel: {n_data}x{n_data} = {n_data**2:,} entries")
print(
f"Low-rank: {n_data}x{n_inducing} + {n_inducing}x{n_inducing} "
f"= {n_data * n_inducing + n_inducing**2:,} entries"
)Data points: 200
Inducing points: 15
Dense kernel: 200x200 = 40,000 entries
Low-rank: 200x15 + 15x15 = 3,225 entries
Build the low-rank approximation¶
ls, var = 1.0, 1.5
noise = 0.1
def rbf_kernel_2d(x1, x2, ls, var):
sq_dist = (x1[:, None] - x2[None, :]) ** 2
return var * jnp.exp(-0.5 * sq_dist / ls**2)
# K_nm: data-to-inducing kernel
K_nm = rbf_kernel_2d(x_data, x_inducing, ls, var)
# K_mm: inducing-to-inducing kernel
K_mm = rbf_kernel_2d(x_inducing, x_inducing, ls, var)
# Nystrom factor: U = K_nm @ chol(K_mm)^{-T} gives K approx U U^T
L_mm = jnp.linalg.cholesky(K_mm + 1e-6 * jnp.eye(n_inducing))
U = jax.scipy.linalg.solve_triangular(L_mm, K_nm.T, lower=True).T
print(f"U shape: {U.shape}") # (200, 15)U shape: (200, 15)
Construct gaussx LowRankUpdate operator¶
diag_noise = noise * jnp.ones(n_data)
sigma = gaussx.low_rank_plus_diag(diag_noise, U)
print(f"LowRankUpdate rank: {sigma.rank}")
print(f"Operator size: {sigma.in_size()}")
print(f"gaussx.is_low_rank: {gaussx.is_low_rank(sigma)}")LowRankUpdate rank: 15
Operator size: 200
gaussx.is_low_rank: True
Solve and logdet via Woodbury¶
gaussx automatically uses the Woodbury identity for solve and the matrix determinant lemma for logdet — O(nk^2 + k^3) instead of O(n^3).
# Generate observations
y_data = jnp.sin(2 * x_data) + noise * jax.random.normal(k2, (n_data,))
# Woodbury solve: (sigma^2 I + U U^T)^{-1} y
alpha = gaussx.solve(sigma, y_data)
print(f"solve residual: {jnp.max(jnp.abs(sigma.mv(alpha) - y_data)):.2e}")
# Matrix determinant lemma logdet
ld = gaussx.logdet(sigma)
ld_dense = jnp.linalg.slogdet(sigma.as_matrix())[1]
print(f"logdet (Woodbury): {ld:.6f}")
print(f"logdet (dense): {ld_dense:.6f}")
print(f"Match: {jnp.allclose(ld, ld_dense, rtol=1e-4)}")solve residual: 4.56e-13
logdet (Woodbury): -421.679302
logdet (dense): -421.679302
Match: True
Log-marginal likelihood¶
def sparse_gp_lml(y, sigma_op):
"""Sparse GP log-marginal likelihood."""
alpha = gaussx.solve(sigma_op, y)
data_fit = -0.5 * jnp.dot(y, alpha)
complexity = -0.5 * gaussx.logdet(sigma_op)
constant = -0.5 * len(y) * jnp.log(2 * jnp.pi)
return data_fit + complexity + constant
lml = sparse_gp_lml(y_data, sigma)
print(f"Sparse GP LML: {lml:.4f}")Sparse GP LML: 12.4216
Visualize the sparse GP fit¶
# Predict on a test grid using the sparse approximation
x_plot = jnp.linspace(-3.5, 3.5, 300)
K_star_m = rbf_kernel_2d(x_plot, x_inducing, ls, var)
L_mm_op = lx.MatrixLinearOperator(
K_mm + 1e-6 * jnp.eye(n_inducing), lx.positive_semidefinite_tag
)
# Nystrom prediction weights
alpha_sparse = gaussx.solve(sigma, y_data)
# Project through inducing points
w = U.T @ alpha_sparse # (k,)
y_plot = K_star_m @ jnp.linalg.solve(
K_mm + 1e-6 * jnp.eye(n_inducing), K_nm.T @ alpha_sparse
)
fig, ax = plt.subplots(figsize=(10, 4))
ax.plot(x_plot, jnp.sin(2 * x_plot), "k--", lw=1.5, label="True", zorder=4)
ax.scatter(
x_data,
y_data,
s=30,
c="C0",
alpha=0.4,
edgecolors="k",
linewidths=0.5,
label="Data",
zorder=5,
)
ax.plot(x_plot, y_plot, "C1-", lw=2, label="Sparse GP mean", zorder=3)
ax.axvline(x_inducing[0], color="gray", lw=0.5, alpha=0.3)
for xi in x_inducing[1:]:
ax.axvline(xi, color="gray", lw=0.5, alpha=0.3)
ax.scatter(
x_inducing,
jnp.zeros_like(x_inducing) - 1.2,
marker="^",
s=40,
c="C2",
zorder=5,
label=f"Inducing pts ({n_inducing})",
)
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.set_title(f"Sparse GP: {n_data} data, {n_inducing} inducing points")
ax.legend(fontsize=9)
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.tight_layout()
plt.show()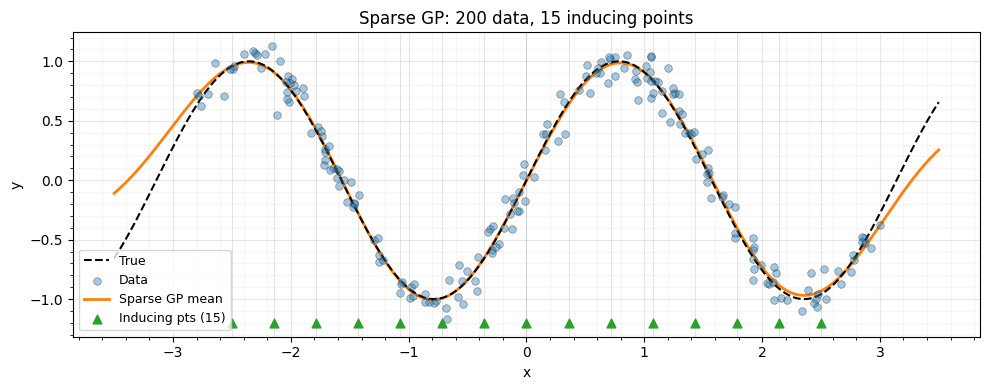
Summary¶
| Structure | When to use | Solve cost | Logdet cost |
|---|---|---|---|
Kronecker(K1, K2) | Grid data | O(n1^3 + n2^3) | O(n1^3 + n2^3) |
low_rank_plus_diag | Inducing points | O(nk^2 + k^3) | O(nk^2 + k^3) |
| Dense | Unstructured | O(n^3) | O(n^3) |
gaussx dispatches to the right algorithm automatically based on the operator type — write the math once, get efficient computation for free.
References¶
- Quinonero-Candela, J. & Rasmussen, C. E. (2005). A unifying view of sparse approximate Gaussian process regression. JMLR, 6, 1939--1959.
- Saatci, Y. (2012). Scalable Inference for Structured Gaussian Process Models. PhD thesis, University of Cambridge.
- Rasmussen, C. E. & Williams, C. K. I. (2006). Gaussian Processes for Machine Learning. MIT Press.