Pathwise GP Posterior Sampling
This notebook demonstrates pyrox.gp.PathwiseSampler and pyrox.gp.DecoupledPathwiseSampler — callable posterior function draws via Matheron’s rule. Each sampled path is a single deterministic function that can be evaluated at arbitrary inputs without refactorizing a test-set covariance, which is the enabler for Thompson sampling, Bayesian optimization, and posterior visualization.
What you’ll learn:
- The Matheron-rule identity behind
PathwiseSamplerand why the RFF prior draw matters. - Drawing and evaluating posterior function samples from an exact
ConditionedGP. - The sparse / decoupled variant over an SVGP-style inducing set.
- A minimal Thompson-sampling loop on a 1D toy objective.
Background¶
Matheron’s rule¶
For a GP prior with Gaussian observations , , the posterior sample path admits the Matheron decomposition
where is drawn from the prior and is an independent noise draw at the training inputs. The key insight: if the prior draw is represented by a closed-form basis (e.g. random Fourier features), then is a single callable function — evaluate it at any in without touching .
Random Fourier features for the prior draw¶
For a stationary kernel , Bochner’s theorem gives a spectral density such that
Sampling frequencies and phases ,
defines a function whose covariance converges to as . pyrox._basis.draw_rff_cosine_basis is the pure shared primitive that implements this for RBF and Matern kernels; the pathwise samplers call into it.
Sparse / decoupled form¶
For a sparse SparseGPPrior + variational guide over inducing values , the decoupled Matheron sample is
DecoupledPathwiseSampler handles WhitenedGuide automatically by unwhitening into before forming the correction.
Setup¶
import subprocess
import sys
try:
import google.colab # noqa: F401
IN_COLAB = True
except ImportError:
IN_COLAB = False
if IN_COLAB:
subprocess.run(
[
sys.executable,
"-m",
"pip",
"install",
"-q",
"pyrox[colab] @ git+https://github.com/jejjohnson/pyrox@main",
],
check=True,
)import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import jax
import jax.numpy as jnp
import jax.random as jr
import matplotlib.pyplot as plt
import numpyro
from pyrox.gp import (
RBF,
DecoupledPathwiseSampler,
FullRankGuide,
GPPrior,
PathwiseSampler,
SparseGPPrior,
)
jax.config.update("jax_enable_x64", True)Toy dataset¶
key = jr.PRNGKey(0)
f_true = lambda x: jnp.sin(2.0 * x) + 0.3 * jnp.cos(5.0 * x)
X_train = jnp.concatenate(
[jnp.linspace(-3.0, -0.5, 10), jnp.linspace(1.5, 3.0, 10)]
).reshape(-1, 1)
noise_std = 0.15
y_train = f_true(X_train.squeeze(-1)) + noise_std * jr.normal(key, (X_train.shape[0],))
X_test = jnp.linspace(-3.5, 3.5, 200).reshape(-1, 1)
noise_var = jnp.array(noise_std**2)Exact pathwise samples¶
Fit a plain ConditionedGP, then draw 32 callable posterior paths and evaluate them on the dense test grid.
posterior = GPPrior(
kernel=RBF(init_variance=1.0, init_lengthscale=0.6),
X=X_train,
jitter=1e-6,
).condition(y_train, noise_var)
sampler = PathwiseSampler(posterior, n_features=1024)
with numpyro.handlers.seed(rng_seed=0):
paths = sampler.sample_paths(jr.PRNGKey(1), n_paths=32)
draws = paths(X_test) # (32, N_star) — same callable evaluated on the full grid
mean_star, var_star = posterior.predict(X_test)
std_star = jnp.sqrt(jnp.clip(var_star, min=0.0))fig, ax = plt.subplots(figsize=(12, 5))
ax.plot(
X_test.squeeze(),
f_true(X_test.squeeze()),
"k--",
lw=1.5,
label="True function",
zorder=5,
)
for draw in draws:
ax.plot(X_test.squeeze(), draw, color="C0", alpha=0.15, lw=1.0)
ax.plot(
X_test.squeeze(), mean_star, "C3-", lw=2, label="Analytic posterior mean", zorder=4
)
ax.fill_between(
X_test.squeeze(),
mean_star - 2 * std_star,
mean_star + 2 * std_star,
color="C3",
alpha=0.15,
label=r"$\pm 2\sigma$ (analytic)",
)
ax.scatter(
X_train.squeeze(),
y_train,
s=30,
c="C1",
edgecolors="k",
linewidths=0.5,
label="Training data",
zorder=6,
)
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.set_title(
"Exact GP: 32 pathwise posterior samples (blue) vs. analytic moments (red)"
)
ax.legend(loc="lower left", fontsize=9)
plt.show()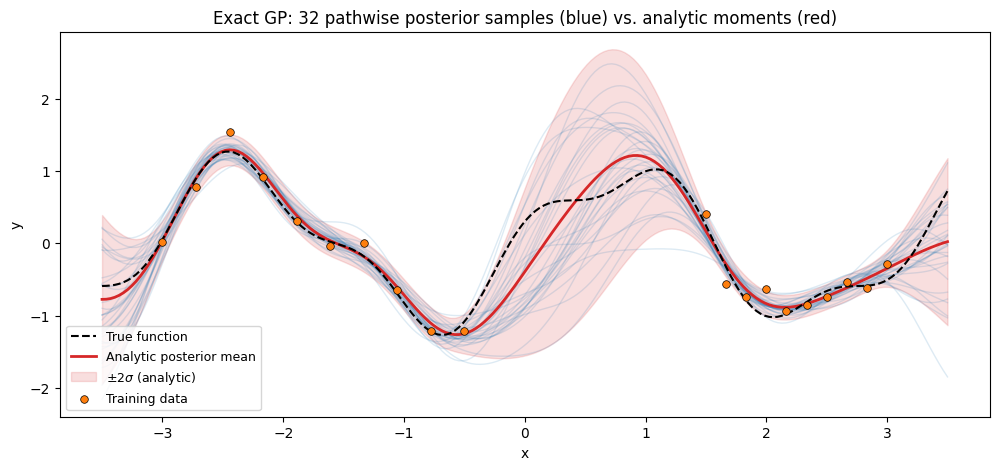
The blue paths are independent callable function draws. Reevaluating the same path on a different input batch gives numerically identical values — a property the analytic predictive moments alone cannot provide without refactorizing a joint test-set covariance.
Sanity check: empirical moments from 512 draws match the analytic posterior.
with numpyro.handlers.seed(rng_seed=0):
big = PathwiseSampler(posterior, n_features=2048)(
jr.PRNGKey(2), X_test, n_paths=512
)
empirical_mean = jnp.mean(big, axis=0)
empirical_std = jnp.std(big, axis=0)
fig, axes = plt.subplots(1, 2, figsize=(14, 4), sharex=True)
axes[0].plot(X_test.squeeze(), empirical_mean, "C0-", lw=2, label="Empirical")
axes[0].plot(X_test.squeeze(), mean_star, "k--", lw=1.5, label="Analytic")
axes[0].set_title("Mean")
axes[0].set_xlabel("x")
axes[0].legend()
axes[1].plot(X_test.squeeze(), empirical_std, "C0-", lw=2, label="Empirical")
axes[1].plot(X_test.squeeze(), std_star, "k--", lw=1.5, label="Analytic")
axes[1].set_title("Standard deviation")
axes[1].set_xlabel("x")
axes[1].legend()
plt.tight_layout()
plt.show()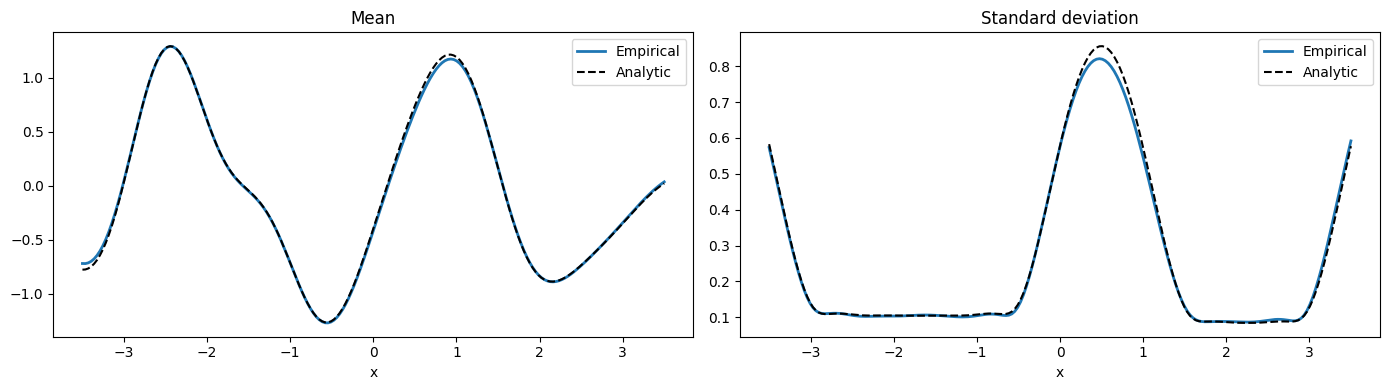
Sparse / decoupled pathwise samples¶
Same story with a sparse GP: a point-inducing SparseGPPrior, a variational guide , and the decoupled Matheron update. The prior draw reuses the RFF basis; the correction is computed in the inducing-point basis, so the resulting function is callable at arbitrary inputs after a one-time inducing solve.
To give the sampler a realistic guide — instead of a hand-picked mean/covariance with no relation to the data — we set to the exact conditional of the inducing values given the training data. This is the closed-form Bayes-optimal Gaussian over , and it’s what a fully-converged SVGP guide would match. The resulting decoupled paths should reproduce the exact GP posterior samples up to the RFF + inducing-set approximation.
Z = jnp.linspace(-3.5, 3.5, 20).reshape(-1, 1)
kernel_sparse = RBF(init_variance=1.0, init_lengthscale=0.6)
sparse_prior = SparseGPPrior(kernel=kernel_sparse, Z=Z, jitter=1e-6)
# Closed-form posterior over inducing values u = f(Z) given training data:
# q(u) = p(u | y) = N(K_zx (K_xx + σ² I)⁻¹ y, K_zz − K_zx (K_xx + σ² I)⁻¹ K_xz).
K_xx = kernel_sparse(X_train, X_train) + noise_var * jnp.eye(X_train.shape[0])
K_xx_inv = jnp.linalg.inv(K_xx)
K_zx = kernel_sparse(Z, X_train)
K_zz = kernel_sparse(Z, Z) + 1e-6 * jnp.eye(Z.shape[0])
m_u = K_zx @ K_xx_inv @ y_train
S_u = K_zz - K_zx @ K_xx_inv @ K_zx.T
S_u = 0.5 * (S_u + S_u.T) + 1e-6 * jnp.eye(Z.shape[0])
L_u = jnp.linalg.cholesky(S_u)
guide = FullRankGuide(mean=m_u, scale_tril=L_u)
with numpyro.handlers.seed(rng_seed=0):
sparse_paths = DecoupledPathwiseSampler(
sparse_prior, guide, n_features=1024
).sample_paths(jr.PRNGKey(3), n_paths=32)
sparse_draws = sparse_paths(X_test)fig, ax = plt.subplots(figsize=(12, 5))
for draw in sparse_draws:
ax.plot(X_test.squeeze(), draw, color="C2", alpha=0.15, lw=1.0)
ax.plot(
X_test.squeeze(),
f_true(X_test.squeeze()),
"k--",
lw=1.5,
label="True function",
zorder=5,
)
ax.plot(
X_test.squeeze(),
mean_star,
"C3-",
lw=2,
label="Exact-GP posterior mean (reference)",
zorder=4,
)
ax.scatter(
X_train.squeeze(),
y_train,
s=30,
c="C1",
edgecolors="k",
linewidths=0.5,
label="Training data",
zorder=6,
)
ax.scatter(
Z.squeeze(),
m_u,
s=40,
c="C2",
marker="^",
edgecolors="k",
linewidths=0.5,
label=r"Inducing posterior mean $m_u$",
zorder=6,
)
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.set_title("DecoupledPathwiseSampler: 32 SVGP posterior samples")
ax.legend(loc="lower left", fontsize=9)
plt.show()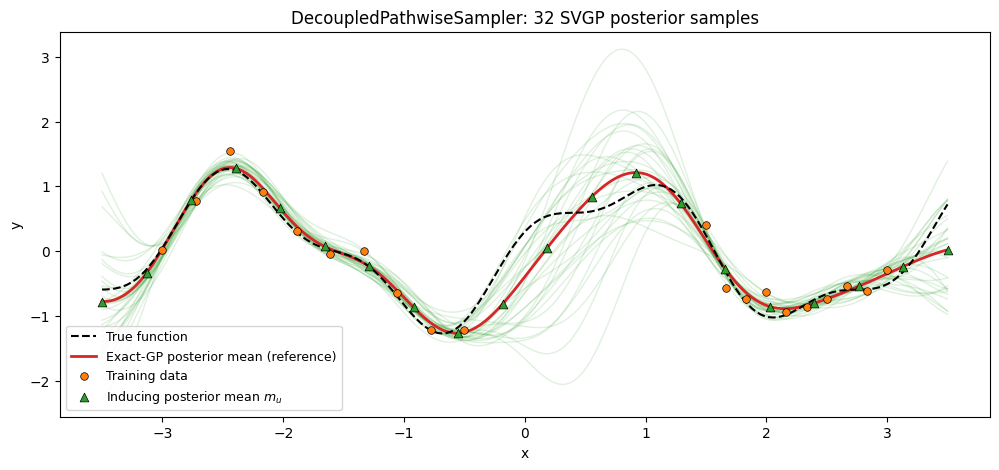
Thompson sampling on a 1D toy¶
The canonical pathwise-sampling use case. At each round:
- Draw a single posterior path from the current
ConditionedGP. - Pick the argmin of that path (Thompson’s “sample and act” rule for minimization).
- Observe the true objective plus noise and condition the GP on the augmented dataset.
Because a sampled path is a single callable function, step 2 is a trivial grid / argmin evaluation — no repeated full-covariance predictives.
def objective(x):
return (x - 1.7) ** 2 * jnp.cos(3.0 * x) + 0.1 * x
domain = jnp.linspace(-3.0, 3.0, 400).reshape(-1, 1)
# Seed observations.
ts_key = jr.PRNGKey(123)
X_obs = jnp.array([[-2.0], [0.0], [2.5]])
y_obs = objective(X_obs.squeeze(-1)) + 0.05 * jr.normal(ts_key, (X_obs.shape[0],))
def ts_step(X_obs, y_obs, round_key):
ts_posterior = GPPrior(
kernel=RBF(init_variance=1.5, init_lengthscale=0.6),
X=X_obs,
jitter=1e-6,
).condition(y_obs, jnp.array(0.05**2))
path = PathwiseSampler(ts_posterior, n_features=1024).sample_paths(
round_key, n_paths=1
)
draw = path(domain)[0] # (N_domain,)
x_next = domain[jnp.argmin(draw)] # (1,) — one row of domain
y_next = objective(x_next) + 0.05 * jr.normal(round_key, (1,))
return (
jnp.concatenate([X_obs, x_next[None, :]], axis=0),
jnp.concatenate([y_obs, y_next], axis=0),
draw,
)
rounds = 8
recorded = [(X_obs.copy(), y_obs.copy(), None)]
for r in range(rounds):
X_obs, y_obs, draw = ts_step(X_obs, y_obs, jr.fold_in(jr.PRNGKey(999), r))
recorded.append((X_obs.copy(), y_obs.copy(), draw))fig, axes = plt.subplots(2, 4, figsize=(18, 8), sharex=True, sharey=True)
for ax, (X_step, y_step, draw) in zip(axes.flat, recorded[1:]):
ax.plot(
domain.squeeze(),
objective(domain.squeeze()),
"k--",
lw=1.2,
label="True objective",
)
if draw is not None:
ax.plot(domain.squeeze(), draw, color="C0", lw=1.2, label="Sampled path")
ax.scatter(
X_step.squeeze(),
y_step,
s=40,
c="C1",
edgecolors="k",
linewidths=0.5,
label="Observed",
)
ax.scatter(
X_step[-1:].squeeze(),
y_step[-1:],
s=120,
facecolors="none",
edgecolors="C3",
linewidths=2.0,
label="Just picked",
)
ax.set_xlabel("x")
for ax in axes[:, 0]:
ax.set_ylabel("y")
axes[0, 0].legend(loc="upper left", fontsize=8)
plt.suptitle("Thompson sampling: 8 rounds")
plt.tight_layout()
plt.show()
domain_obj = objective(domain.squeeze(-1))
print(f"Best observed value: {float(jnp.min(y_obs)):.3f}")
print(
f"Argmin (true): x* = {float(domain.squeeze(-1)[jnp.argmin(domain_obj)]):.3f}"
)
print(f"Argmin (observed): x = {float(X_obs.squeeze(-1)[jnp.argmin(y_obs)]):.3f}")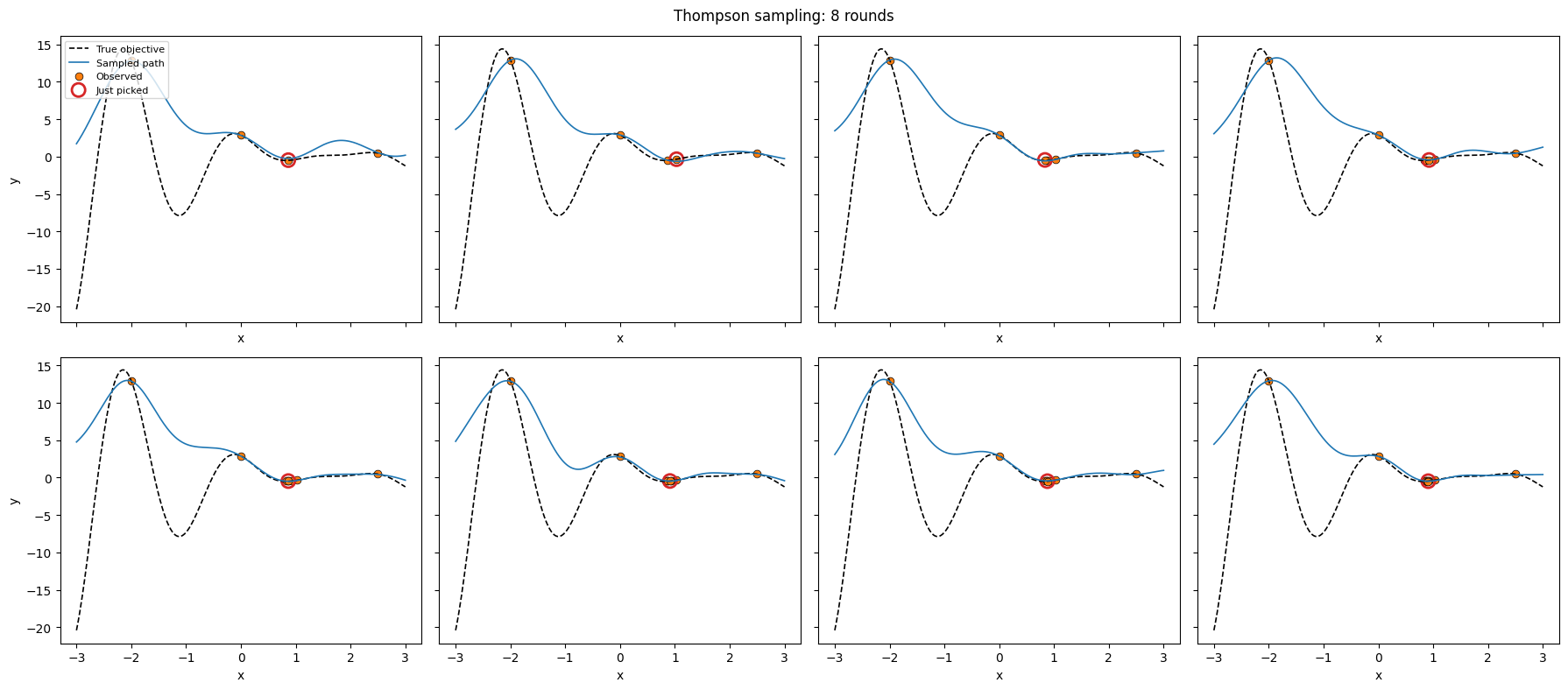
Best observed value: -0.484
Argmin (true): x* = -3.000
Argmin (observed): x = 0.910
Takeaways¶
- A pathwise sample is a callable function, not a vector of values — reusable across arbitrary test sets.
PathwiseSampleruses the cached noisy operator fromConditionedGP, so the training Cholesky is shared with the standard predictive path.DecoupledPathwiseSamplercomposes with any of theFullRankGuide,MeanFieldGuide,WhitenedGuide,NaturalGuide,DeltaGuideguides; whitened guides are unwhitened automatically.- The shared RFF prior draw lives in
pyrox._basis.draw_rff_cosine_basis+evaluate_rff_cosine_paths— the same primitive other posterior-function workflows (custom BO acquisitions, ensembled spectral estimators) can build on directly.