Parallel Kalman Filter & SSM Natural Parameters
This notebook demonstrates the parallel (associative-scan) Kalman filter and smoother, the Joseph-form covariance update, and the natural/expectation parameter representations of linear-Gaussian state-space models.
What you’ll learn:
- Running
gaussx.parallel_kalman_filterandgaussx.parallel_rts_smoother - Verifying equivalence with the sequential Kalman filter
- The Joseph-form covariance update for numerical stability
- Converting SSM parameters to natural parameters (
BlockTriDiagprecision) - Converting smoother output to expectation parameters and back
Background: why parallelism matters¶
The standard Kalman filter is inherently sequential: each step depends on the previous filtered state. On modern accelerators (GPU/TPU) this sequential bottleneck limits throughput for long time series.
The associative-scan formulation (Sarkka & Garcia-Fernandez, 2021) reformulates the Kalman recursion as a parallel prefix sum over a semigroup of affine transformations. This enables parallel depth instead of sequential steps, while producing identical results up to floating-point precision.
In gaussx, parallel_kalman_filter and parallel_rts_smoother provide
the same interface as their sequential counterparts but use dense array
operations that are faster on GPU/TPU for long sequences.
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import jax
import jax.numpy as jnp
import matplotlib.pyplot as plt
import gaussx
jax.config.update("jax_enable_x64", True)Define the model¶
We reuse the damped oscillator from the sequential Kalman filter notebook. The 2D state follows a discretized spring-mass-damper system, and only position is observed with noise.
- : discretized spring-mass-damper transition
- (observe position only)
- : process noise driving the oscillator
- : observation noise variance
dt = 0.1
T = 200
# Damped oscillator: omega=1.0 rad/s, damping gamma=0.15
omega, gamma = 1.0, 0.15
A = jnp.array([[1.0, dt], [-(omega**2) * dt, 1.0 - gamma * dt]])
# Observation matrix (observe position only)
H = jnp.array([[1.0, 0.0]])
# Process noise covariance
q_var = 0.3
Q = q_var * jnp.array([[dt**3 / 3, dt**2 / 2], [dt**2 / 2, dt]])
# Observation noise covariance
R = jnp.array([[0.5]])
print("A =\n", A)
print("H =", H)
print("Q =\n", Q)
print("R =", R)A =
[[ 1. 0.1 ]
[-0.1 0.985]]
H = [[1. 0.]]
Q =
[[0.0001 0.0015]
[0.0015 0.03 ]]
R = [[0.5]]
Simulate data¶
Generate time steps from the model. The true trajectory is sampled from the generative process so the Kalman filter assumptions are satisfied.
key = jax.random.PRNGKey(42)
def simulate_step(carry, key_t):
x = carry
k1, k2 = jax.random.split(key_t)
q_t = jax.random.multivariate_normal(k1, jnp.zeros(2), Q)
x_new = A @ x + q_t
r_t = jax.random.multivariate_normal(k2, jnp.zeros(1), R)
y_t = H @ x_new + r_t
return x_new, (x_new, y_t)
x0 = jnp.array([3.0, 0.0]) # displaced from equilibrium, at rest
keys = jax.random.split(key, T)
_, (true_states, observations) = jax.lax.scan(simulate_step, x0, keys)
times = jnp.arange(T) * dt
true_position = true_states[:, 0]
print("true_states shape:", true_states.shape)
print("observations shape:", observations.shape)true_states shape: (200, 2)
observations shape: (200, 1)
Run parallel Kalman filter and smoother¶
init_mean = jnp.zeros(2)
init_cov = jnp.eye(2) * 4.0
# Parallel filter
par_filter_state = gaussx.parallel_kalman_filter(
A, H, Q, R, observations, init_mean, init_cov
)
print("Parallel filtered means shape:", par_filter_state.filtered_means.shape)
print("Parallel log-likelihood:", par_filter_state.log_likelihood)Parallel filtered means shape: (200, 2)
Parallel log-likelihood: -223.3188576581507
# Parallel smoother
par_smooth_means, par_smooth_covs = gaussx.parallel_rts_smoother(par_filter_state, A, Q)
print("Parallel smoothed means shape:", par_smooth_means.shape)
print("Parallel smoothed covs shape:", par_smooth_covs.shape)Parallel smoothed means shape: (200, 2)
Parallel smoothed covs shape: (200, 2, 2)
Compare with sequential Kalman filter¶
The parallel and sequential implementations should produce identical results (up to floating-point precision). We verify this by comparing filtered means, covariances, and the log-likelihood.
# Sequential filter
seq_filter_state = gaussx.kalman_filter(A, H, Q, R, observations, init_mean, init_cov)
# Sequential smoother
seq_smooth_means, seq_smooth_covs = gaussx.rts_smoother(seq_filter_state, A, Q)
# Compare filtered means
mean_diff = jnp.max(
jnp.abs(par_filter_state.filtered_means - seq_filter_state.filtered_means)
)
print(f"Max abs diff (filtered means): {mean_diff:.2e}")
# Compare filtered covariances
cov_diff = jnp.max(
jnp.abs(par_filter_state.filtered_covs - seq_filter_state.filtered_covs)
)
print(f"Max abs diff (filtered covs): {cov_diff:.2e}")
# Compare log-likelihoods
ll_diff = jnp.abs(par_filter_state.log_likelihood - seq_filter_state.log_likelihood)
print(f"Abs diff (log-likelihood): {ll_diff:.2e}")
# Compare smoothed means
smooth_mean_diff = jnp.max(jnp.abs(par_smooth_means - seq_smooth_means))
print(f"Max abs diff (smoothed means): {smooth_mean_diff:.2e}")
# Compare smoothed covariances
smooth_cov_diff = jnp.max(jnp.abs(par_smooth_covs - seq_smooth_covs))
print(f"Max abs diff (smoothed covs): {smooth_cov_diff:.2e}")Max abs diff (filtered means): 1.05e-15
Max abs diff (filtered covs): 4.44e-16
Abs diff (log-likelihood): 0.00e+00
Max abs diff (smoothed means): 4.00e-15
Max abs diff (smoothed covs): 8.88e-16
Joseph-form covariance update¶
The standard Kalman update computes the posterior covariance as , but this can lose symmetry or positive definiteness due to round-off. The Joseph form is more robust:
We manually compute the Kalman gain at one time step and compare both formulas.
# Pick a time step to illustrate
t_idx = 50
P_pred = par_filter_state.predicted_covs[t_idx] # (2, 2)
x_pred = par_filter_state.predicted_means[t_idx] # (2,)
# Compute Kalman gain K = P_pred H^T (H P_pred H^T + R)^{-1}
S = H @ P_pred @ H.T + R
K = P_pred @ H.T @ jnp.linalg.inv(S)
print(f"Time step {t_idx}:")
print(f"Kalman gain K =\n{K}")
# Standard update: (I - KH) P_pred
d_state = P_pred.shape[0]
P_standard = (jnp.eye(d_state) - K @ H) @ P_pred
# Joseph-form update
P_joseph = gaussx.joseph_update(P_pred, K, H, R)
print(f"\nStandard update P =\n{P_standard}")
print(f"\nJoseph update P =\n{P_joseph}")
print(f"\nMax abs difference: {jnp.max(jnp.abs(P_standard - P_joseph)):.2e}")
# The Joseph form guarantees symmetry
print(f"\nStandard symmetry error: {jnp.max(jnp.abs(P_standard - P_standard.T)):.2e}")
print(f"Joseph symmetry error: {jnp.max(jnp.abs(P_joseph - P_joseph.T)):.2e}")Time step 50:
Kalman gain K =
[[0.16284017]
[0.13551443]]
Standard update P =
[[0.08142008 0.06775721]
[0.06775721 0.21846408]]
Joseph update P =
[[0.08142008 0.06775721]
[0.06775721 0.21846408]]
Max abs difference: 9.71e-17
Standard symmetry error: 1.94e-16
Joseph symmetry error: 0.00e+00
SSM natural parameters¶
A linear-Gaussian state-space model defines a joint Gaussian
whose precision matrix is block-tridiagonal.
The function ssm_to_naturals extracts the natural parameters
where is
stored as a BlockTriDiag operator.
The API expects Q to have shape (N, d, d) where Q[0] is the initial
covariance and Q[1:] contains the process noise at each step.
# Build the full Q array: Q[0] = P_0, Q[1:] = process noise
d_state = A.shape[0]
Q_full = jnp.concatenate([init_cov[None], jnp.tile(Q, (T - 1, 1, 1))], axis=0)
A_full = jnp.tile(A, (T - 1, 1, 1))
print("A_full shape:", A_full.shape)
print("Q_full shape:", Q_full.shape)
print("Q_full[0] (= P_0):\n", Q_full[0])A_full shape: (199, 2, 2)
Q_full shape: (200, 2, 2)
Q_full[0] (= P_0):
[[4. 0.]
[0. 4.]]
# Convert to natural parameters
theta_linear, theta_precision = gaussx.ssm_to_naturals(
A_full, Q_full, init_mean, Q_full[0]
)
print("theta_linear shape:", theta_linear.shape)
print("theta_precision type:", type(theta_precision).__name__)
print("theta_precision diagonal shape:", theta_precision.diagonal.shape)
print("theta_precision sub_diagonal shape:", theta_precision.sub_diagonal.shape)theta_linear shape: (400,)
theta_precision type: BlockTriDiag
theta_precision diagonal shape: (200, 2, 2)
theta_precision sub_diagonal shape: (199, 2, 2)
Inspect the BlockTriDiag structure¶
The BlockTriDiag operator stores (N, d, d) diagonal blocks and
(N-1, d, d) sub-diagonal blocks. Its dense representation is a
matrix with the characteristic banded sparsity
pattern of a Gauss-Markov chain.
# Materialize the dense precision matrix (small enough for T=200, d=2)
dense_precision = theta_precision.as_matrix()
print("Dense precision shape:", dense_precision.shape)Dense precision shape: (400, 400)
# Round-trip: convert back to SSM parameters
A_rt, Q_rt, mu0_rt, P0_rt = gaussx.naturals_to_ssm(theta_linear, theta_precision)
print("Round-trip A max error:", jnp.max(jnp.abs(A_rt - A_full)).item())
print("Round-trip Q max error:", jnp.max(jnp.abs(Q_rt - Q_full)).item())
print("Round-trip mu_0 max error:", jnp.max(jnp.abs(mu0_rt - init_mean)).item())
print("Round-trip P_0 max error:", jnp.max(jnp.abs(P0_rt - init_cov)).item())Round-trip A max error: 1.8197554574328478e-11
Round-trip Q max error: 8.177721610991284e-09
Round-trip mu_0 max error: 0.0
Round-trip P_0 max error: 8.177721610991284e-09
SSM expectation parameters¶
Given smoothed marginals and cross-covariances , we can compute the expectation parameters of the joint Gaussian:
- (concatenated means)
- : a
BlockTriDiagwith diagonal blocks and sub-diagonal blocks
The cross-covariance is computed from the smoother gain: where .
# Compute cross-covariances from filter/smoother outputs
# G_t = P_{t|t} A^T P_{t+1|t}^{-1}
# C_t = G_t @ P_{t+1|T}
def compute_cross_covs(filter_state, smoothed_covs, A):
"""Compute cross-covariances P_{t+1,t|T} = G_t @ P_{t+1|T}."""
P_filt = filter_state.filtered_covs[:-1] # (T-1, d, d)
P_pred = filter_state.predicted_covs[1:] # (T-1, d, d)
P_smooth_next = smoothed_covs[1:] # (T-1, d, d)
def _one_step(P_f, P_p, P_s_next):
G = jnp.linalg.solve(P_p.T, (P_f @ A.T).T).T
return G @ P_s_next
return jax.vmap(_one_step)(P_filt, P_pred, P_smooth_next)
cross_covs = compute_cross_covs(par_filter_state, par_smooth_covs, A)
print("Cross-covariances shape:", cross_covs.shape)Cross-covariances shape: (199, 2, 2)
# Convert to expectation parameters
eta1, eta2 = gaussx.ssm_to_expectations(par_smooth_means, par_smooth_covs, cross_covs)
print("eta1 shape:", eta1.shape)
print("eta2 type:", type(eta2).__name__)
print("eta2 diagonal shape:", eta2.diagonal.shape)
print("eta2 sub_diagonal shape:", eta2.sub_diagonal.shape)eta1 shape: (400,)
eta2 type: BlockTriDiag
eta2 diagonal shape: (200, 2, 2)
eta2 sub_diagonal shape: (199, 2, 2)
# Round-trip: convert back to SSM marginals
means_rt, covs_rt, cross_covs_rt = gaussx.expectations_to_ssm(eta1, eta2)
err_m = jnp.max(jnp.abs(means_rt - par_smooth_means)).item()
err_c = jnp.max(jnp.abs(covs_rt - par_smooth_covs)).item()
err_cc = jnp.max(jnp.abs(cross_covs_rt - cross_covs)).item()
print("Round-trip means max error:", err_m)
print("Round-trip covs max error:", err_c)
print("Round-trip cross_covs max error:", err_cc)Round-trip means max error: 0.0
Round-trip covs max error: 8.604228440844963e-16
Round-trip cross_covs max error: 8.604228440844963e-16
Visualizations¶
BlockTriDiag sparsity pattern¶
The precision matrix of a Gauss-Markov chain has a characteristic block-tridiagonal sparsity pattern. We visualize the first few blocks to see this structure clearly.
fig, axes = plt.subplots(1, 2, figsize=(14, 5))
# Sparsity pattern of the precision (show first 20 blocks = 40x40)
n_show = min(20, T)
d_show = n_show * d_state
prec_sub = dense_precision[:d_show, :d_show]
ax = axes[0]
ax.spy(jnp.abs(prec_sub) > 1e-12, markersize=1, color="C0")
ax.set_title(f"BlockTriDiag sparsity (first {n_show} blocks)")
ax.set_xlabel("Column index")
ax.set_ylabel("Row index")
# Zoom into a 3-block region to see the structure
n_zoom = 3
d_zoom = n_zoom * d_state
prec_zoom = dense_precision[:d_zoom, :d_zoom]
ax = axes[1]
im = ax.imshow(prec_zoom, cmap="RdBu_r", aspect="equal")
ax.set_title(f"Precision blocks (first {n_zoom} blocks)")
ax.set_xlabel("Column index")
ax.set_ylabel("Row index")
# Draw block boundaries
for i in range(1, n_zoom):
ax.axhline(i * d_state - 0.5, color="k", lw=0.5, ls="--")
ax.axvline(i * d_state - 0.5, color="k", lw=0.5, ls="--")
plt.colorbar(im, ax=ax, shrink=0.8)
plt.tight_layout()
plt.show()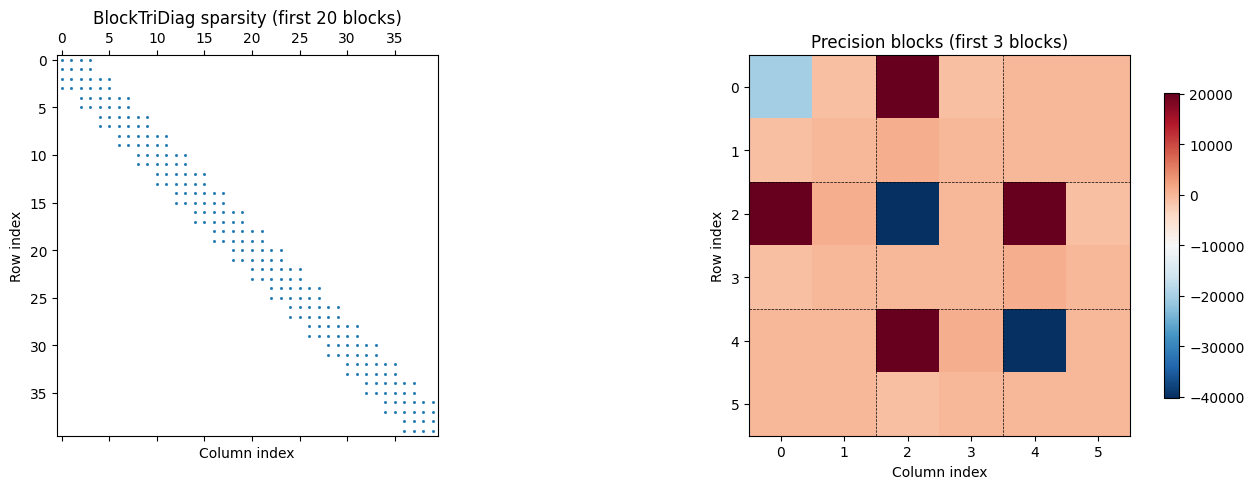
Filtered vs smoothed estimates¶
filt_pos_mean = par_filter_state.filtered_means[:, 0]
filt_pos_std = jnp.sqrt(par_filter_state.filtered_covs[:, 0, 0])
smooth_pos_mean = par_smooth_means[:, 0]
smooth_pos_std = jnp.sqrt(par_smooth_covs[:, 0, 0])
filt_vel_mean = par_filter_state.filtered_means[:, 1]
filt_vel_std = jnp.sqrt(par_filter_state.filtered_covs[:, 1, 1])
smooth_vel_mean = par_smooth_means[:, 1]
smooth_vel_std = jnp.sqrt(par_smooth_covs[:, 1, 1])
fig, axes = plt.subplots(2, 1, figsize=(12, 7), sharex=True)
# Position
ax = axes[0]
ax.scatter(
times,
observations[:, 0],
s=30,
c="C0",
edgecolors="k",
linewidths=0.5,
alpha=0.25,
label="Observations",
zorder=5,
)
ax.plot(times, true_position, "k--", lw=1.5, label="True", zorder=4)
ax.fill_between(
times,
filt_pos_mean - 2 * filt_pos_std,
filt_pos_mean + 2 * filt_pos_std,
color="C1",
alpha=0.15,
label=r"Filtered $\pm 2\sigma$",
)
ax.plot(times, filt_pos_mean, "C1-", lw=2, alpha=0.6, label="Filtered", zorder=3)
ax.fill_between(
times,
smooth_pos_mean - 2 * smooth_pos_std,
smooth_pos_mean + 2 * smooth_pos_std,
color="C2",
alpha=0.25,
label=r"Smoothed $\pm 2\sigma$",
)
ax.plot(times, smooth_pos_mean, "C2-", lw=2, label="Smoothed", zorder=3)
ax.set_ylabel("Position")
ax.set_title("Parallel Kalman: Filtered vs Smoothed (Position)")
ax.legend(loc="upper right", fontsize=9, ncol=2)
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
# Velocity
ax = axes[1]
ax.plot(times, true_states[:, 1], "k--", lw=1.5, label="True", zorder=4)
ax.fill_between(
times,
filt_vel_mean - 2 * filt_vel_std,
filt_vel_mean + 2 * filt_vel_std,
color="C1",
alpha=0.15,
label=r"Filtered $\pm 2\sigma$",
)
ax.plot(times, filt_vel_mean, "C1-", lw=2, alpha=0.6, label="Filtered", zorder=3)
ax.fill_between(
times,
smooth_vel_mean - 2 * smooth_vel_std,
smooth_vel_mean + 2 * smooth_vel_std,
color="C3",
alpha=0.25,
label=r"Smoothed $\pm 2\sigma$",
)
ax.plot(times, smooth_vel_mean, "C3-", lw=2, label="Smoothed", zorder=3)
ax.set_xlabel("Time")
ax.set_ylabel("Velocity (latent)")
ax.set_title("Parallel Kalman: Filtered vs Smoothed (Velocity)")
ax.legend(loc="upper right", fontsize=9, ncol=2)
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.tight_layout()
plt.show()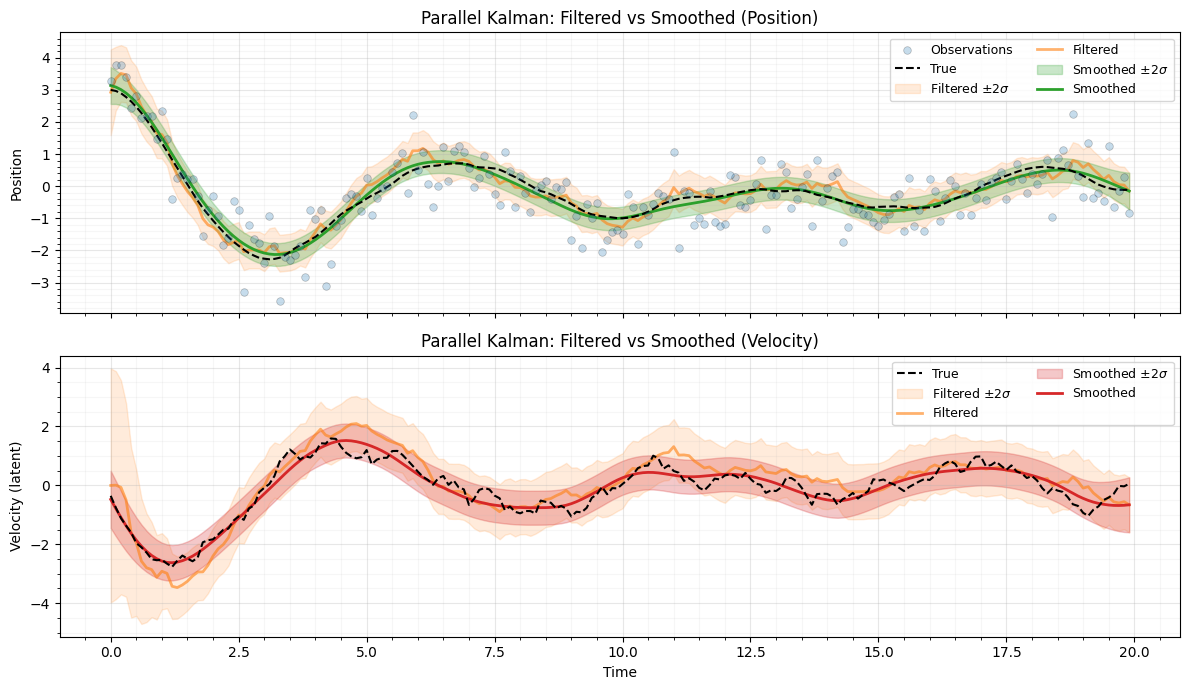
Summary¶
- Parallel Kalman filter:
gaussx.parallel_kalman_filterproduces identical results to the sequentialgaussx.kalman_filterbut uses dense array operations suited for GPU/TPU acceleration. - Parallel RTS smoother:
gaussx.parallel_rts_smootherlikewise matches the sequential smoother. - Joseph-form update:
gaussx.joseph_updatecomputes the Kalman covariance update in a numerically stable form that preserves symmetry and positive-definiteness. - Natural parameters:
gaussx.ssm_to_naturalsextracts the block-tridiagonal precision (as aBlockTriDiagoperator) and the linear natural parameter from SSM matrices.gaussx.naturals_to_ssminverts this exactly. - Expectation parameters:
gaussx.ssm_to_expectationsconverts smoother marginals (means, covariances, cross-covariances) to expectation parameters withBlockTriDiagsecond moments.gaussx.expectations_to_ssminverts this exactly. - The
BlockTriDiagoperator exploits Gauss-Markov sparsity for operations instead of dense cost.
References¶
- Sarkka, S. & Garcia-Fernandez, A. F. (2021). Temporal parallelization of Bayesian smoothers. IEEE Trans. Automatic Control, 66(1), 299--306.
- Kalman, R. E. (1960). A new approach to linear filtering and prediction problems. J. Basic Engineering, 82(1), 35--45.
- Rauch, H. E., Tung, F., & Striebel, C. T. (1965). Maximum likelihood estimates of linear dynamic systems. AIAA Journal, 3(8), 1445--1450.
- Sarkka, S. (2013). Bayesian Filtering and Smoothing. Cambridge University Press.