Kernel Approximation with Random Fourier Features
This notebook is a deep dive into the Bochner foundation behind every spectral layer in pyrox.nn. It answers a sequence of empirical questions:
- Does the math actually work? Build via random Fourier features and watch it converge to the exact RBF Gram at the Rahimi-Recht rate .
- Does it generalise across kernels? The same feature map approximates RBF, Matérn-3/2, and Laplace — only the prior on changes.
- Are the two readout flavours equivalent? Compare paired vs phased across all three kernels — equivalence with the kernel, equivalence under the GP prior (sample paths), and head-to-head variance at matched output dimension.
- Can we lower the variance further?
OrthogonalRandomFeatures(Yu et al. 2016) replaces iid Gaussian rows of with negatively-correlated Haar-orthogonal blocks — provably lower variance at fixed .
This is the kernel-approximation half of the spectral story; once the feature map is trusted, the Random Fourier Features regression notebook builds the SSGP / VSSGP / MAP regression hierarchy on top of it. Other adjacent notebooks: RFF as Neural Networks (frozen / ensemble of MAP variants) and Deep Random Feature Expansions (hierarchical extensions).
Background — the math behind Random Fourier Features¶
Bochner’s theorem¶
A continuous, shift-invariant, real-valued positive-definite kernel on is the Fourier transform of a finite non-negative spectral measure μ:
Because is real, μ is symmetric (), so the imaginary part of the integral vanishes and we may write
where is the normalised spectral density. Bochner’s theorem turns the abstract claim “the kernel is positive-definite” into the very concrete claim “the kernel is the characteristic function of a probability distribution on frequencies”.
Monte Carlo feature map (Rahimi & Recht, NeurIPS 2007)¶
Draw from the spectral density. Define the paired random Fourier feature map
The Monte Carlo Gram matrix is an unbiased estimator of , and Rahimi-Recht’s Claim 1 gives uniform convergence on any compact subset of at rate .
All spectral-method NN layers in pyrox.nn are different choices of wrapped around the same feature map. pyrox.nn._layers._rff_forward is literally one line of JAX implementing the equation above. The same Ω also drives the single-cosine cos(\Omega^\top x + b) readout used by RBFCosineFeatures and its Matérn / Laplace siblings — the choice between the two is a variance trade-off we’ll pin down empirically in §1.3.
Spectral densities, by kernel¶
Different stationary kernels correspond to different spectral densities. The core three:
| Kernel | Spectral density | Paired [cos, sin] layer | Phased cos(\cdot + b) layer |
|---|---|---|---|
| RBF | RBFFourierFeatures | RBFCosineFeatures | |
| Matérn-ν | multivariate Student- | MaternFourierFeatures | MaternCosineFeatures |
| Laplace (Matérn-1/2) | multivariate Cauchy | LaplaceFourierFeatures | LaplaceCosineFeatures |
All three kernel families come in two parallel readout flavours. The paired map (Lázaro-Gredilla et al. 2010) outputs features and the kernel identity holds exactly for any Ω. The phased map (Rahimi-Recht 2007 / Random Kitchen Sinks) outputs features and the same identity holds in expectation over — the phase draw kills the cross term . §1.3 below shows the two readouts side-by-side, both numerically and as GP prior samples.
Setup¶
Detect Colab and install pyrox[colab] (which pulls in matplotlib and watermark) only when running there.
import subprocess
import sys
try:
import google.colab # noqa: F401
IN_COLAB = True
except ImportError:
IN_COLAB = False
if IN_COLAB:
subprocess.run(
[
sys.executable,
"-m",
"pip",
"install",
"-q",
"pyrox[colab] @ git+https://github.com/jejjohnson/pyrox@main",
],
check=True,
)import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import equinox as eqx
import jax
import jax.numpy as jnp
import jax.random as jr
import matplotlib.pyplot as plt
import numpy as np
from numpyro import handlers
from pyrox.nn import (
LaplaceCosineFeatures,
LaplaceFourierFeatures,
MaternCosineFeatures,
MaternFourierFeatures,
OrthogonalRandomFeatures,
RBFCosineFeatures,
RBFFourierFeatures,
)
jax.config.update("jax_enable_x64", True)import importlib.util
try:
from IPython import get_ipython
ipython = get_ipython()
except ImportError:
ipython = None
if ipython is not None and importlib.util.find_spec("watermark") is not None:
ipython.run_line_magic("load_ext", "watermark")
ipython.run_line_magic(
"watermark",
"-v -m -p jax,equinox,numpyro,pyrox,matplotlib",
)
else:
print("watermark extension not installed; skipping reproducibility readout.")Python implementation: CPython
Python version : 3.13.5
IPython version : 9.10.0
jax : 0.9.2
equinox : 0.13.6
numpyro : 0.20.1
pyrox : 0.0.7
matplotlib: 3.10.8
Compiler : GCC 11.2.0
OS : Linux
Release : 6.8.0-1044-azure
Machine : x86_64
Processor : x86_64
CPU cores : 16
Architecture: 64bit
1. Bochner kernel approximation — does the math actually work?¶
Before any regression model, we verify the foundation. Bochner says . Rahimi-Recht say the Monte Carlo estimator with samples converges at . Let’s see both.
Build a 1D test grid, compute the exact RBF Gram matrix , then realise the approximate Gram via pyrox.nn.RBFFourierFeatures for and inspect the Frobenius error.
Sampling from RBFFourierFeatures is done by tracing the layer under numpyro.handlers.seed — the layer’s pyrox_sample("W", ...) site fires with the seeded key, and we recover the sampled from the trace. Throughout this notebook we hold the lengthscale fixed via handlers.substitute so the comparison is purely about the feature map.
def trace_rff_features(
rff: RBFFourierFeatures | MaternFourierFeatures | LaplaceFourierFeatures,
x: jax.Array,
seed: int,
lengthscale: float,
) -> jax.Array:
"""Return the realised random feature map at ``x`` for a given seed.
The pyrox RFF layers sample *both* ``W`` and ``lengthscale`` (the latter
from a ``LogNormal`` prior). To compare against a fixed-$\\ell$ exact
Gram matrix in Section 1 we hold the lengthscale fixed via ``substitute``
and only let ``W`` vary across seeds.
"""
name = f"{rff.pyrox_name}.lengthscale"
with (
handlers.substitute(data={name: jnp.asarray(lengthscale)}),
handlers.seed(rng_seed=seed),
):
return rff(x)
def gram_exact_rbf(x: jax.Array, lengthscale: float) -> jax.Array:
"""Exact RBF Gram matrix on a 1D grid."""
diff = x[:, None] - x[None, :]
return jnp.exp(-0.5 * (diff[..., 0] ** 2) / lengthscale**2)
# Test grid + exact target
n_grid = 50
x_grid = jnp.linspace(-2.0, 2.0, n_grid).reshape(-1, 1)
LENGTHSCALE = 0.5
K_exact = gram_exact_rbf(x_grid, LENGTHSCALE)
# Approximate Grams for an increasing budget of features
m_values = [16, 64, 256, 1024]
n_repeats = 20 # average over seeds for the convergence-rate plot
approx_grams = {}
for M in m_values:
rff = eqx.tree_at(
lambda r: r.pyrox_name,
RBFFourierFeatures.init(in_features=1, n_features=M, lengthscale=LENGTHSCALE),
"rff",
)
phi = trace_rff_features(rff, x_grid, seed=0, lengthscale=LENGTHSCALE)
approx_grams[M] = phi @ phi.TVisualise converging to .
fig, axes = plt.subplots(1, len(m_values) + 1, figsize=(15, 3.2))
for ax, M in zip(axes[:-1], m_values, strict=False):
err = float(jnp.linalg.norm(approx_grams[M] - K_exact) / jnp.linalg.norm(K_exact))
ax.imshow(approx_grams[M], vmin=0.0, vmax=1.0, cmap="viridis")
ax.set_title(f"$\\hat{{K}}_M$, $M = {M}$\nrel. err = {err:.3f}")
ax.set_xticks([])
ax.set_yticks([])
axes[-1].imshow(K_exact, vmin=0.0, vmax=1.0, cmap="viridis")
axes[-1].set_title("$K^\\star$ (exact RBF)")
axes[-1].set_xticks([])
axes[-1].set_yticks([])
plt.tight_layout()
plt.show()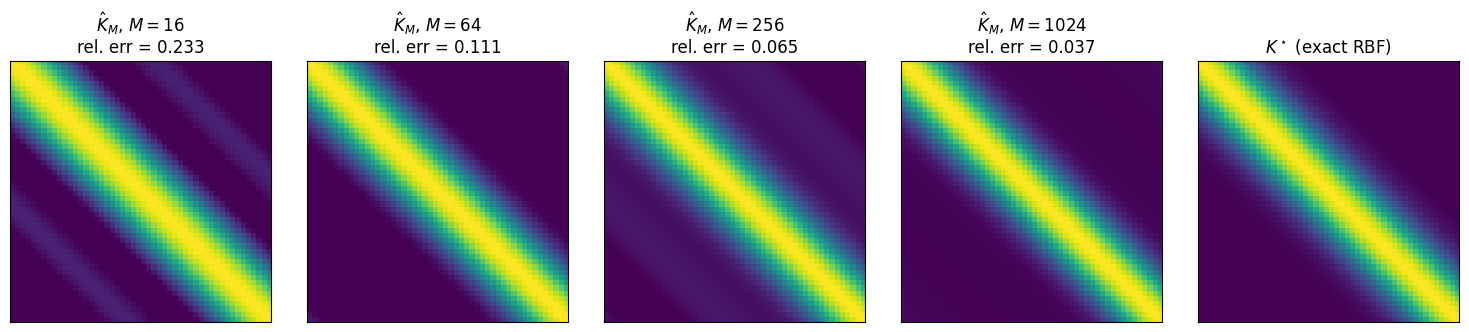
Convergence rate — average over 20 seeds and plot versus on log-log axes. The Rahimi-Recht bound predicts a slope close to .
m_sweep = [8, 16, 32, 64, 128, 256, 512, 1024]
errs = np.zeros((len(m_sweep), n_repeats))
for i, M in enumerate(m_sweep):
rff = eqx.tree_at(
lambda r: r.pyrox_name,
RBFFourierFeatures.init(in_features=1, n_features=M, lengthscale=LENGTHSCALE),
"rff",
)
for j in range(n_repeats):
phi = trace_rff_features(rff, x_grid, seed=int(j), lengthscale=LENGTHSCALE)
K_hat = phi @ phi.T
errs[i, j] = float(jnp.linalg.norm(K_hat - K_exact) / jnp.linalg.norm(K_exact))
fig, ax = plt.subplots(figsize=(7, 4.5))
mean_err = errs.mean(axis=1)
std_err = errs.std(axis=1)
ax.loglog(m_sweep, mean_err, "o-", label="empirical relative error")
ax.fill_between(m_sweep, mean_err - std_err, mean_err + std_err, alpha=0.3)
ref = mean_err[0] * np.sqrt(m_sweep[0]) / np.sqrt(m_sweep)
ax.loglog(m_sweep, ref, "k--", label=r"$\propto 1/\sqrt{M}$ reference")
ax.set_xlabel("number of features $M$")
ax.set_ylabel(r"$\|\hat{K}_M - K^\star\|_F\ /\ \|K^\star\|_F$")
ax.set_title("Bochner-MC kernel approximation: empirical convergence")
ax.legend()
ax.grid(True, which="both", alpha=0.3)
plt.tight_layout()
plt.show()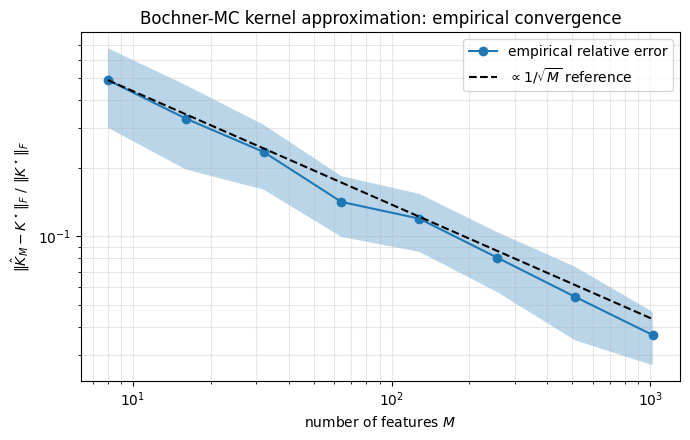
The empirical curve closely tracks the reference, confirming Rahimi-Recht’s bound (the factor is invisible at this scale). At the relative error is below — for most regression problems that is well within noise.
Three kernels, one feature map. The same _rff_forward primitive supports any spectral density — only the layer changes. Show the same diagonal slice of at for RBF, Matérn-3/2, and Laplace (Matérn-1/2). Lengthscale is fixed at across all three so the comparison is purely about regularity.
def gram_exact_matern(x: jax.Array, lengthscale: float, nu: float) -> jax.Array:
"""Exact Matern-1/2, 3/2, 5/2 Gram matrix on a 1D grid."""
r = jnp.abs(x[:, None] - x[None, :])[..., 0]
if nu == 0.5:
return jnp.exp(-r / lengthscale)
if nu == 1.5:
s = jnp.sqrt(3.0) * r / lengthscale
return (1.0 + s) * jnp.exp(-s)
if nu == 2.5:
s = jnp.sqrt(5.0) * r / lengthscale
return (1.0 + s + s**2 / 3.0) * jnp.exp(-s)
raise ValueError(f"Unsupported nu={nu}")
M = 256
rff_rbf = eqx.tree_at(
lambda r: r.pyrox_name,
RBFFourierFeatures.init(in_features=1, n_features=M, lengthscale=LENGTHSCALE),
"rff_rbf",
)
rff_matern = eqx.tree_at(
lambda r: r.pyrox_name,
MaternFourierFeatures.init(
in_features=1, n_features=M, nu=1.5, lengthscale=LENGTHSCALE
),
"rff_matern",
)
rff_laplace = eqx.tree_at(
lambda r: r.pyrox_name,
LaplaceFourierFeatures.init(in_features=1, n_features=M, lengthscale=LENGTHSCALE),
"rff_laplace",
)
phi_rbf = trace_rff_features(rff_rbf, x_grid, seed=0, lengthscale=LENGTHSCALE)
phi_matern = trace_rff_features(rff_matern, x_grid, seed=0, lengthscale=LENGTHSCALE)
phi_laplace = trace_rff_features(rff_laplace, x_grid, seed=0, lengthscale=LENGTHSCALE)
i_centre = n_grid // 2
fig, ax = plt.subplots(figsize=(8, 4.5))
x1 = x_grid[:, 0]
ax.plot(x1, gram_exact_rbf(x_grid, LENGTHSCALE)[i_centre], "C0--", label="RBF (exact)")
ax.plot(x1, (phi_rbf @ phi_rbf.T)[i_centre], "C0", alpha=0.85, label="RBF (RFF)")
ax.plot(
x1,
gram_exact_matern(x_grid, LENGTHSCALE, nu=1.5)[i_centre],
"C1--",
label="Matérn-3/2 (exact)",
)
ax.plot(
x1,
(phi_matern @ phi_matern.T)[i_centre],
"C1",
alpha=0.85,
label="Matérn-3/2 (RFF)",
)
ax.plot(
x1,
gram_exact_matern(x_grid, LENGTHSCALE, nu=0.5)[i_centre],
"C2--",
label="Laplace (exact)",
)
ax.plot(
x1, (phi_laplace @ phi_laplace.T)[i_centre], "C2", alpha=0.85, label="Laplace (RFF)"
)
ax.set_xlabel("$x$")
ax.set_ylabel("$k(x_{\\mathrm{centre}}, x)$")
ax.set_title(
f"Three spectral densities, one feature map — $M={M}$, $\\ell={LENGTHSCALE}$"
)
ax.legend(loc="upper right", fontsize=9)
ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()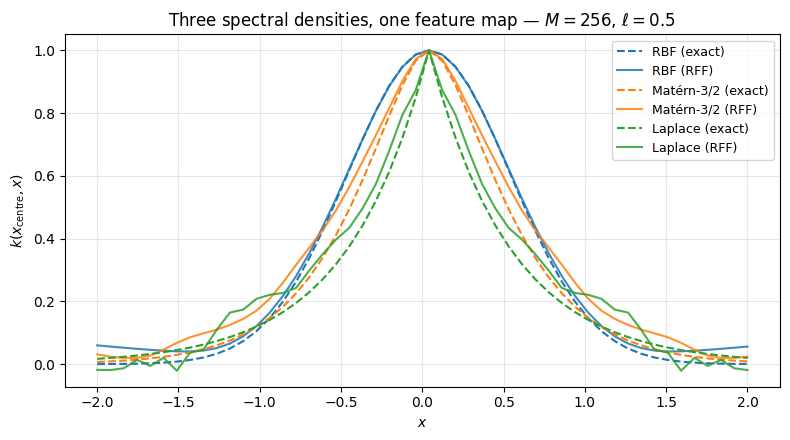
Same scaffold (_rff_forward), three different priors on : → smooth RBF, Student- → less-smooth Matérn-3/2, Cauchy → exponential Laplace. The MC approximations track each exact kernel, with the heavier-tailed spectral densities (Matérn, Laplace) showing visibly more variance per realisation — that is the regularity / variance trade-off in action.
1.3 Paired vs phased readouts — two feature maps, one Bochner identity¶
Both pyrox.nn.RBFFourierFeatures and pyrox.nn.RBFCosineFeatures (and their Matérn / Laplace siblings) draw Ω from the same spectral density. They differ only in the readout that turns Ω into a feature map:
Why both work. Expand the inner products:
- Paired: . Exact for every Ω — no expectation over phase needed.
- Phased: . The cross term has expectation zero under , so the identity holds in expectation only.
Predicted consequence: at matched output dimension, paired has lower variance per draw because it never relies on the phase expectation cancelling. We test all three kernels.
1.3.a Equivalence with the kernel. Sanity check: with a single seed and frequencies, both readouts track each of RBF, Matérn-3/2, Laplace. Lengthscale fixed at .
M_DEMO = 256
i_centre = n_grid // 2 # already defined above; reuse the 1D test grid x_grid
paired_layers = {
"RBF": eqx.tree_at(
lambda r: r.pyrox_name,
RBFFourierFeatures.init(
in_features=1, n_features=M_DEMO, lengthscale=LENGTHSCALE
),
"p_rbf",
),
"Matérn-3/2": eqx.tree_at(
lambda r: r.pyrox_name,
MaternFourierFeatures.init(
in_features=1, n_features=M_DEMO, nu=1.5, lengthscale=LENGTHSCALE
),
"p_matern",
),
"Laplace": eqx.tree_at(
lambda r: r.pyrox_name,
LaplaceFourierFeatures.init(
in_features=1, n_features=M_DEMO, lengthscale=LENGTHSCALE
),
"p_laplace",
),
}
phased_layers = {
"RBF": eqx.tree_at(
lambda r: r.pyrox_name,
RBFCosineFeatures.init(
in_features=1, n_features=M_DEMO, lengthscale=LENGTHSCALE
),
"c_rbf",
),
"Matérn-3/2": eqx.tree_at(
lambda r: r.pyrox_name,
MaternCosineFeatures.init(
in_features=1, n_features=M_DEMO, nu=1.5, lengthscale=LENGTHSCALE
),
"c_matern",
),
"Laplace": eqx.tree_at(
lambda r: r.pyrox_name,
LaplaceCosineFeatures.init(
in_features=1, n_features=M_DEMO, lengthscale=LENGTHSCALE
),
"c_laplace",
),
}
exact_grams = {
"RBF": gram_exact_rbf(x_grid, LENGTHSCALE),
"Matérn-3/2": gram_exact_matern(x_grid, LENGTHSCALE, nu=1.5),
"Laplace": gram_exact_matern(x_grid, LENGTHSCALE, nu=0.5),
}
fig, axes = plt.subplots(1, 3, figsize=(15, 4.0), sharey=True)
x1 = x_grid[:, 0]
for ax, name in zip(axes, ["RBF", "Matérn-3/2", "Laplace"], strict=False):
K_exact_k = exact_grams[name]
phi_p = trace_rff_features(
paired_layers[name], x_grid, seed=0, lengthscale=LENGTHSCALE
)
phi_c = trace_rff_features(
phased_layers[name], x_grid, seed=1, lengthscale=LENGTHSCALE
)
K_p = phi_p @ phi_p.T
K_c = phi_c @ phi_c.T
ax.plot(x1, K_exact_k[i_centre], "k--", linewidth=1.5, label="exact")
ax.plot(
x1,
K_p[i_centre],
"C0",
alpha=0.85,
linewidth=1.4,
label=f"paired ($2M={2 * M_DEMO}$)",
)
ax.plot(
x1,
K_c[i_centre],
"C3",
alpha=0.85,
linewidth=1.4,
label=f"phased ($M={M_DEMO}$)",
)
ax.set_title(name)
ax.set_xlabel("$x$")
ax.grid(True, alpha=0.3)
ax.legend(fontsize=9, loc="upper right")
axes[0].set_ylabel("$k(x_{\\mathrm{centre}}, x)$")
fig.suptitle("Bochner identity holds for both readouts, all three kernels", y=1.02)
plt.tight_layout()
plt.show()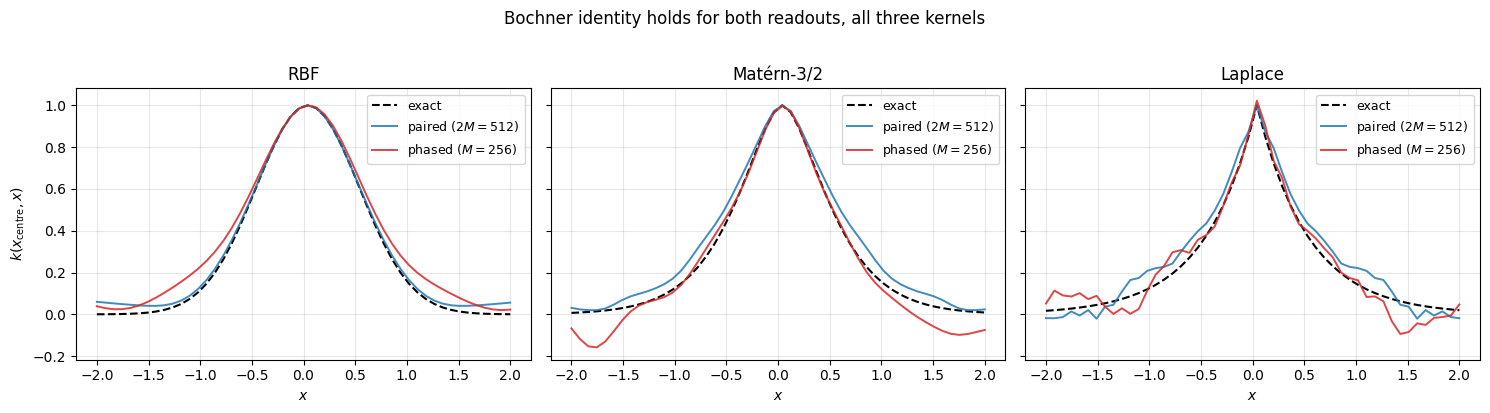
Both readouts hug the exact kernel for all three kernels. The phased curve is visibly noisier (single-seed Monte Carlo over on top of the spectral draw).
1.3.b Equivalence under the prior — sample paths. A linear combination of paired RFFs with is a GP with Monte-Carlo Bochner kernel; the same is true for phased RFFs with . The two parameterisations should produce statistically indistinguishable sample paths from the same GP prior. Draw samples through each readout, recover the empirical Gram , and compare to the exact kernel.
S_PATHS = 200
fig, axes = plt.subplots(3, 3, figsize=(13.5, 10.5), sharex="row")
for row, name in enumerate(["RBF", "Matérn-3/2", "Laplace"]):
K_exact_k = exact_grams[name]
norm = float(jnp.linalg.norm(K_exact_k))
phi_p = trace_rff_features(
paired_layers[name], x_grid, seed=0, lengthscale=LENGTHSCALE
)
phi_c = trace_rff_features(
phased_layers[name], x_grid, seed=1, lengthscale=LENGTHSCALE
)
beta_p = jr.normal(jr.PRNGKey(100 + row), (S_PATHS, phi_p.shape[1]))
alpha_c = jr.normal(jr.PRNGKey(200 + row), (S_PATHS, phi_c.shape[1]))
paths_p = beta_p @ phi_p.T # (S, n_grid)
paths_c = alpha_c @ phi_c.T
K_path_p = paths_p.T @ paths_p / S_PATHS
K_path_c = paths_c.T @ paths_c / S_PATHS
err_p = float(jnp.linalg.norm(K_path_p - K_exact_k) / norm)
err_c = float(jnp.linalg.norm(K_path_c - K_exact_k) / norm)
ax = axes[row, 0]
for s in range(8):
ax.plot(x1, paths_p[s], color="C0", alpha=0.4, linewidth=0.9)
ax.set_title(f"{name} — paired sample paths")
ax.grid(True, alpha=0.3)
ax.set_ylabel("$f(x)$")
ax = axes[row, 1]
for s in range(8):
ax.plot(x1, paths_c[s], color="C3", alpha=0.4, linewidth=0.9)
ax.set_title(f"{name} — phased sample paths")
ax.grid(True, alpha=0.3)
ax = axes[row, 2]
ax.plot(x1, K_exact_k[i_centre], "k--", linewidth=1.5, label="exact")
ax.plot(
x1,
K_path_p[i_centre],
"C0",
alpha=0.85,
linewidth=1.3,
label=f"paired (rel.err = {err_p:.3f})",
)
ax.plot(
x1,
K_path_c[i_centre],
"C3",
alpha=0.85,
linewidth=1.3,
label=f"phased (rel.err = {err_c:.3f})",
)
ax.set_title(f"{name} — empirical Gram from $S = {S_PATHS}$ paths")
ax.grid(True, alpha=0.3)
ax.legend(fontsize=8, loc="upper right")
for ax in axes[-1]:
ax.set_xlabel("$x$")
plt.tight_layout()
plt.show()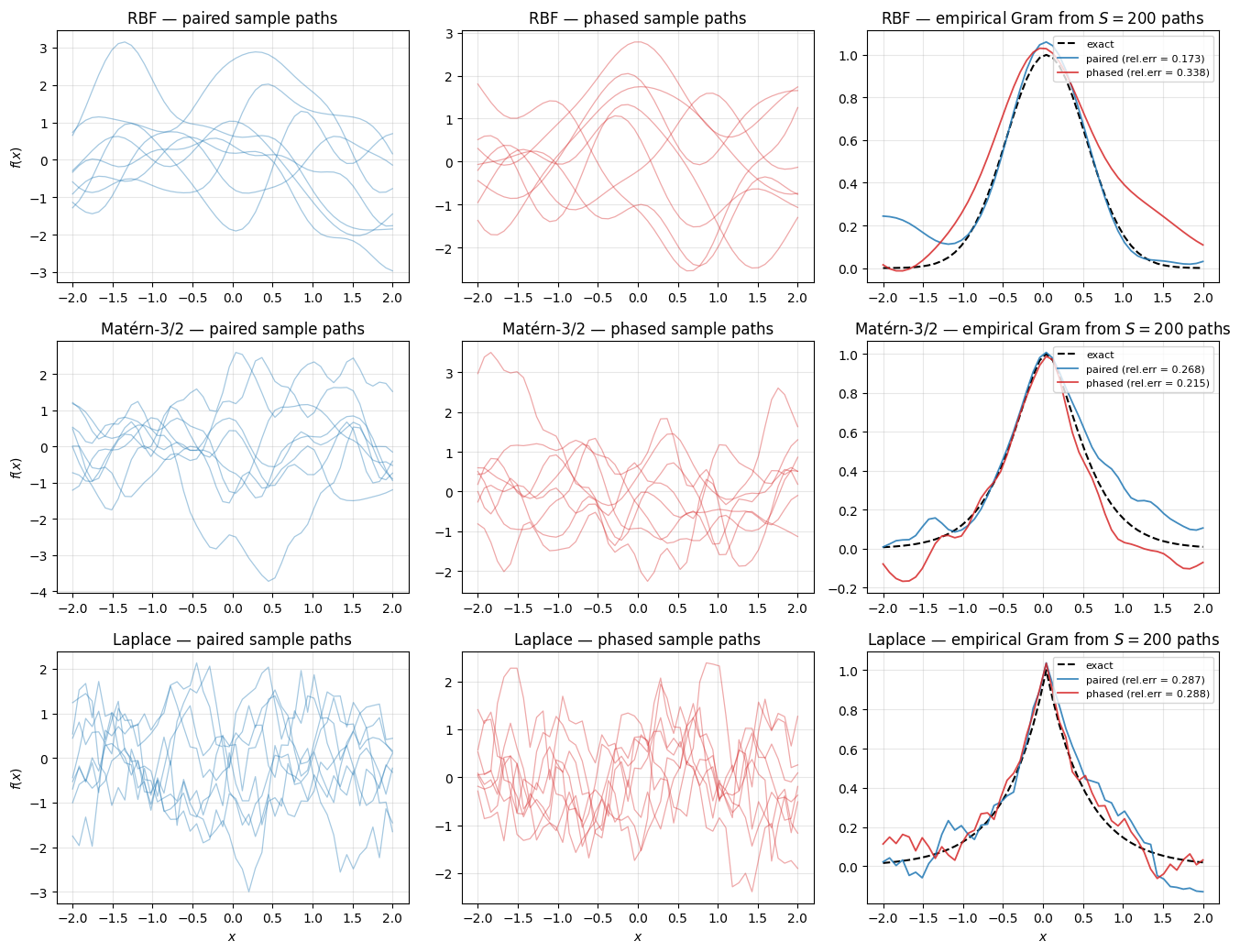
Sample paths from paired vs phased look statistically interchangeable per kernel (same regularity, same lengthscale, same amplitude). The right-column empirical Grams converge to the exact kernel from both readouts at comparable error — the two parameterisations describe the same prior over functions.
1.3.c Variance comparison at matched output dimension. Sweep the output dimension . Paired uses frequencies (output ); phased uses frequencies (output ). Theory: paired has lower variance per draw because it never paid the phase expectation. We expect the gap to be cleanest for RBF (light-tailed spectrum) and to shrink for Matérn / Laplace where the spectral-draw variance dominates the readout variance.
d_sweep = [16, 32, 64, 128, 256, 512]
n_repeats_pp = 12
err_paired = {k: np.zeros((len(d_sweep), n_repeats_pp)) for k in exact_grams}
err_phased = {k: np.zeros((len(d_sweep), n_repeats_pp)) for k in exact_grams}
for kernel_name in exact_grams:
K_exact_k = exact_grams[kernel_name]
norm = float(jnp.linalg.norm(K_exact_k))
for i, d_out in enumerate(d_sweep):
m_paired = d_out // 2
m_phased = d_out
if kernel_name == "RBF":
mk_p = RBFFourierFeatures
mk_c = RBFCosineFeatures
kw = {}
elif kernel_name == "Matérn-3/2":
mk_p = MaternFourierFeatures
mk_c = MaternCosineFeatures
kw = {"nu": 1.5}
else:
mk_p = LaplaceFourierFeatures
mk_c = LaplaceCosineFeatures
kw = {}
for j in range(n_repeats_pp):
l_p = eqx.tree_at(
lambda r: r.pyrox_name,
mk_p.init(
in_features=1, n_features=m_paired, lengthscale=LENGTHSCALE, **kw
),
"swp_p",
)
l_c = eqx.tree_at(
lambda r: r.pyrox_name,
mk_c.init(
in_features=1, n_features=m_phased, lengthscale=LENGTHSCALE, **kw
),
"swp_c",
)
phi_p = trace_rff_features(
l_p, x_grid, seed=int(2 * j), lengthscale=LENGTHSCALE
)
phi_c = trace_rff_features(
l_c, x_grid, seed=int(2 * j + 1), lengthscale=LENGTHSCALE
)
err_paired[kernel_name][i, j] = float(
jnp.linalg.norm(phi_p @ phi_p.T - K_exact_k) / norm
)
err_phased[kernel_name][i, j] = float(
jnp.linalg.norm(phi_c @ phi_c.T - K_exact_k) / norm
)
fig, axes = plt.subplots(1, 3, figsize=(15, 4.0), sharey=True)
for ax, name in zip(axes, ["RBF", "Matérn-3/2", "Laplace"], strict=False):
mp = err_paired[name].mean(axis=1)
sp = err_paired[name].std(axis=1)
mc = err_phased[name].mean(axis=1)
sc = err_phased[name].std(axis=1)
ax.errorbar(
d_sweep,
mp,
yerr=sp,
fmt="o-",
color="C0",
capsize=3,
label="paired $[\\cos, \\sin]$",
)
ax.errorbar(
d_sweep,
mc,
yerr=sc,
fmt="s-",
color="C3",
capsize=3,
label="phased $\\cos(\\cdot + b)$",
)
ref = mp[0] * np.sqrt(d_sweep[0]) / np.sqrt(d_sweep)
ax.plot(
d_sweep, ref, "k--", alpha=0.6, label=r"$\propto 1/\sqrt{d_{\mathrm{out}}}$"
)
ax.set_xscale("log")
ax.set_yscale("log")
ax.set_xlabel("output dimension $d_{\\mathrm{out}}$")
ax.set_title(name)
ax.grid(True, which="both", alpha=0.3)
ax.legend(fontsize=8, loc="lower left")
axes[0].set_ylabel(r"$\|\hat{K} - K^\star\|_F\ /\ \|K^\star\|_F$")
fig.suptitle(
"Variance at matched output dimension — paired vs phased, three kernels", y=1.02
)
plt.tight_layout()
plt.show()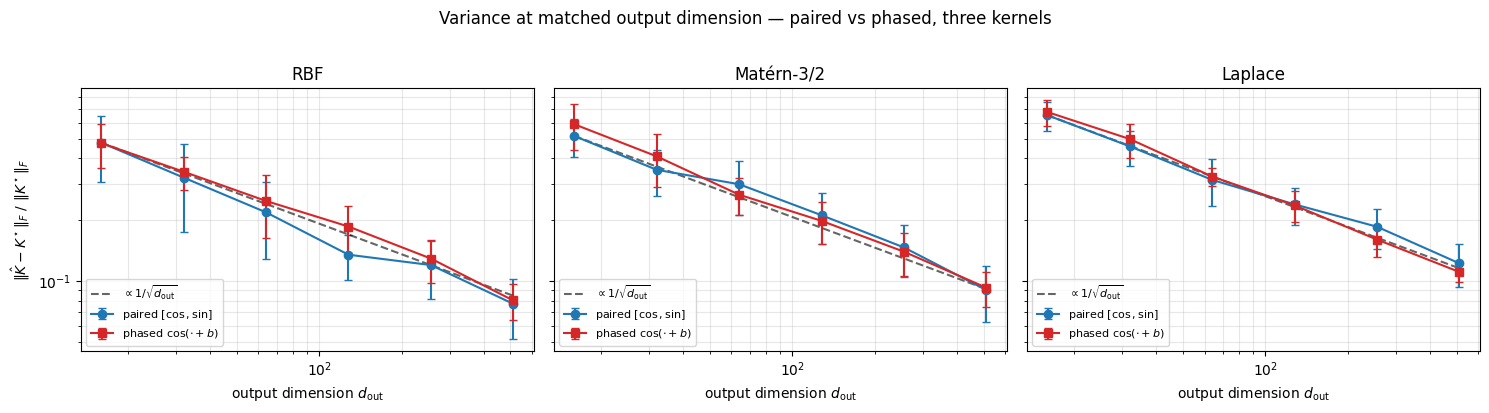
Both curves track the reference. For RBF the paired estimator is consistently below the phased one — paired removes the phase-expectation noise that pays. For Matérn-3/2 and Laplace the gap shrinks: heavy-tailed Student- / Cauchy frequencies dominate the variance, and the choice of readout matters less. Practical take-away: paired is the safer default when you want lower-variance kernel approximations; phased halves the output dimension at matched and is the form Random Kitchen Sinks / pathwise GP samplers naturally use.
Variance reduction with OrthogonalRandomFeatures. A sharper construction within the same family — Yu, Suresh, Choromanski, Felix, Kumar (NeurIPS 2016) — replaces the iid Gaussian rows of with stacked Haar-orthogonal blocks scaled by per-row chi-distributed magnitudes. Each row has the same marginal, but they are negatively correlated, so the variance of at fixed is provably lower. The pyrox.nn.OrthogonalRandomFeatures layer implements this directly. Below: ORF lowers the kernel-approximation error bar at every in .
D_ORF = 4
LENGTHSCALE_ORF = 1.0
n_grid_2d = 100
key_grid = jr.PRNGKey(11)
x_grid_2d = jr.uniform(key_grid, (n_grid_2d, D_ORF), minval=-1.0, maxval=1.0)
diff_2d = x_grid_2d[:, None, :] - x_grid_2d[None, :, :]
K_exact_2d = jnp.exp(-0.5 * jnp.sum(diff_2d**2, axis=-1) / LENGTHSCALE_ORF**2)
m_sweep_orf = [8, 16, 32, 64, 128, 256]
errs_vanilla = np.zeros((len(m_sweep_orf), n_repeats))
errs_orf = np.zeros((len(m_sweep_orf), n_repeats))
for i, M in enumerate(m_sweep_orf):
rff_v = eqx.tree_at(
lambda r: r.pyrox_name,
RBFFourierFeatures.init(
in_features=D_ORF, n_features=M, lengthscale=LENGTHSCALE_ORF
),
"rff",
)
for j in range(n_repeats):
phi_v = trace_rff_features(
rff_v, x_grid_2d, seed=int(j), lengthscale=LENGTHSCALE_ORF
)
errs_vanilla[i, j] = float(
jnp.linalg.norm(phi_v @ phi_v.T - K_exact_2d) / jnp.linalg.norm(K_exact_2d)
)
rff_o = OrthogonalRandomFeatures.init(
in_features=D_ORF,
n_features=M,
key=jr.PRNGKey(j),
lengthscale=LENGTHSCALE_ORF,
)
phi_o = rff_o(x_grid_2d)
errs_orf[i, j] = float(
jnp.linalg.norm(phi_o @ phi_o.T - K_exact_2d) / jnp.linalg.norm(K_exact_2d)
)
fig, ax = plt.subplots(figsize=(7, 4.5))
ax.errorbar(
m_sweep_orf,
errs_vanilla.mean(axis=1),
yerr=errs_vanilla.std(axis=1),
fmt="o-",
label="vanilla RFF",
capsize=3,
)
ax.errorbar(
m_sweep_orf,
errs_orf.mean(axis=1),
yerr=errs_orf.std(axis=1),
fmt="s-",
label="orthogonal RFF",
capsize=3,
)
ax.set_xscale("log")
ax.set_yscale("log")
ax.set_xlabel("number of features $M$")
ax.set_ylabel(r"$\|\hat{K}_M - K^\star\|_F\ /\ \|K^\star\|_F$")
ax.set_title(
f"Vanilla vs orthogonal RFF — variance reduction at matched $M$ ($D = {D_ORF}$)"
)
ax.legend()
ax.grid(True, which="both", alpha=0.3)
plt.tight_layout()
plt.show()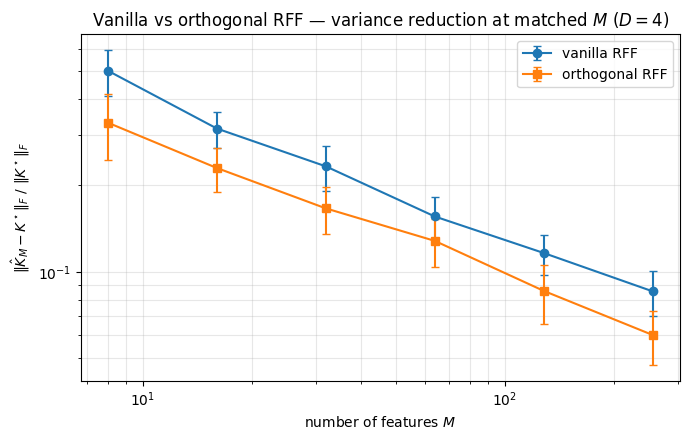
At every , the orthogonal estimator’s error bar sits below the vanilla one — same expected error, lower variance.
Takeaways¶
- Bochner’s theorem is the foundation: any shift-invariant PD kernel is the Fourier transform of a probability density, so admits a Monte Carlo estimator at the Rahimi-Recht rate.
- The same
_rff_forwardprimitive supports RBF, Matérn, Laplace by changing only the prior on .OrthogonalRandomFeatureslowers the variance of at matched via Haar-orthogonal blocks. - Each kernel comes in two readout flavours: paired (
RBFFourierFeatures/MaternFourierFeatures/LaplaceFourierFeatures) and phased (RBFCosineFeatures/MaternCosineFeatures/LaplaceCosineFeatures). Both approximate the same Bochner kernel and produce statistically equivalent GP prior samples; paired has lower variance per draw, phased halves the output dimension.
Where to next.
- With the kernel-approximation foundation trusted, the Random Fourier Features regression notebook builds the MAP / SSGP / VSSGP Bayesian regression hierarchy on top of the same feature map.
- The non-Bayesian / NN flavors of RFF (fixed-Ω with closed-form ridge, ensemble-of-MAP for cheap predictive uncertainty) live in the RFF as Neural Networks notebook.
- The deep / hierarchical versions (deep RFF, deep SSGP, deep VSSGP — Cutajar et al. 2017) live in the Deep Random Feature Expansions notebook.