Kernel Approximations: Nystrom & Random Fourier Features
Full kernel matrices cost memory and time to factorize. For large-scale GP inference we need structured approximations that preserve the low-rank nature of the kernel.
This notebook showcases six gaussx utilities:
| Function | Returns | Purpose |
|---|---|---|
nystrom_operator | LowRankUpdate | Nystrom low-rank approximation |
rff_operator | LowRankUpdate | Random Fourier features approximation |
center_kernel | MatrixLinearOperator | Center a kernel matrix () |
centering_operator | LowRankUpdate | Centering matrix |
hsic | scalar | Hilbert-Schmidt Independence Criterion |
mmd_squared | scalar | Maximum Mean Discrepancy (squared) |
Background¶
Nystrom approximation. Given data points and inducing (landmark) points , the Nystrom method approximates the full kernel matrix as
This is a rank- positive semi-definite matrix. Internally, gaussx
stores it as a LowRankUpdate with zero base and factor
where is the Cholesky factor of
, so that . Adding noise gives
, a LowRankUpdate that can be solved in
via the Woodbury identity.
Random Fourier Features (RFF). Rahimi & Recht (2007) showed that for shift-invariant kernels the kernel function can be approximated via a finite-dimensional random feature map:
where is drawn from the spectral density of the kernel and
. For the RBF kernel with
lengthscale , the spectral density is
.
The feature matrix gives a rank-
approximation , again stored as a
LowRankUpdate.
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import jax
import jax.numpy as jnp
import lineax as lx
import matplotlib.pyplot as plt
import gaussx
jax.config.update("jax_enable_x64", True)Setup: 1D GP regression data¶
We generate 500 noisy observations from a smooth function and define an RBF kernel helper.
key = jax.random.PRNGKey(42)
k1, k2 = jax.random.split(key)
n_train = 500
noise_std = 0.2
f_true = lambda x: jnp.sin(3 * x) * jnp.exp(-0.5 * x**2)
x_train = jnp.sort(jax.random.uniform(k1, (n_train,), minval=-4.0, maxval=4.0))
y_train = f_true(x_train) + noise_std * jax.random.normal(k2, (n_train,))
x_plot = jnp.linspace(-4.5, 4.5, 300)fig, ax = plt.subplots(figsize=(10, 4))
ax.plot(x_plot, f_true(x_plot), "k--", lw=1.5, label="True function", zorder=4)
ax.scatter(
x_train,
y_train,
s=30,
c="C0",
edgecolors="k",
linewidths=0.5,
alpha=0.5,
label="Training data (N=500)",
zorder=5,
)
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.legend(fontsize=9)
ax.set_title("1D Regression Data")
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.tight_layout()
plt.show()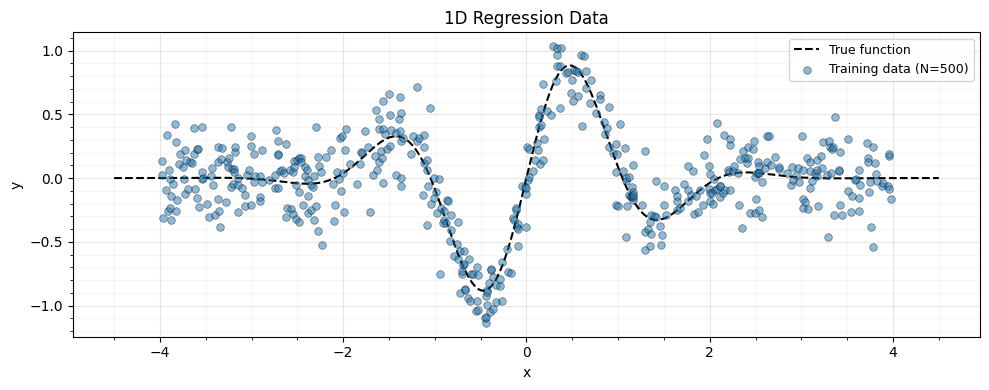
RBF kernel helper¶
def rbf_kernel(x1, x2, lengthscale, variance):
"""Squared-exponential kernel matrix."""
sq_dist = (x1[:, None] - x2[None, :]) ** 2
return variance * jnp.exp(-0.5 * sq_dist / lengthscale**2)
lengthscale = 0.8
variance = 1.0
noise_var = noise_std**2
# Full kernel matrix for comparison
K_full = rbf_kernel(x_train, x_train, lengthscale, variance)
K_full_op = lx.MatrixLinearOperator(K_full)3. Nystrom Approximation¶
We select inducing points on an evenly-spaced grid and build the Nystrom low-rank approximation.
m_inducing = 30
z_inducing = jnp.linspace(-4.0, 4.0, m_inducing)
# Cross-covariance K_XZ (N, M) and inducing kernel K_ZZ (M, M)
K_XZ = rbf_kernel(x_train, z_inducing, lengthscale, variance)
K_ZZ = rbf_kernel(z_inducing, z_inducing, lengthscale, variance)
K_ZZ_op = lx.MatrixLinearOperator(K_ZZ)
# Build the Nystrom operator -> LowRankUpdate
K_nys = gaussx.nystrom_operator(K_XZ, K_ZZ_op)
print(f"Nystrom operator type: {type(K_nys).__name__}")Nystrom operator type: LowRankUpdate
Approximation quality¶
Compare the Nystrom approximation against the full kernel via the Frobenius norm error and a diagonal comparison.
K_nys_dense = K_nys.as_matrix()
frob_error = jnp.linalg.norm(K_full - K_nys_dense, "fro")
frob_relative = frob_error / jnp.linalg.norm(K_full, "fro")
print(f"Frobenius norm error: {frob_error:.4f}")
print(f"Relative Frobenius err: {frob_relative:.4f}")Frobenius norm error: 0.0000
Relative Frobenius err: 0.0000
fig, ax = plt.subplots(figsize=(10, 4))
ax.plot(
x_train,
jnp.diag(K_full),
"k--",
lw=1.5,
label="Full kernel diagonal",
zorder=4,
)
ax.plot(
x_train,
jnp.diag(K_nys_dense),
"--",
color="C1",
lw=2,
label=f"Nystrom diagonal (M={m_inducing})",
zorder=3,
)
ax.set_xlabel("x")
ax.set_ylabel("k(x, x)")
ax.legend(fontsize=9)
ax.set_title("Diagonal Comparison: Full vs Nystrom")
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.tight_layout()
plt.show()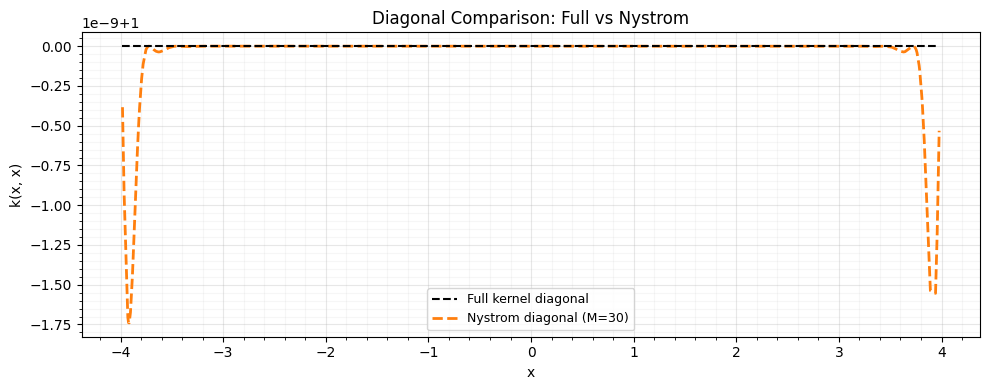
GP regression with the Nystrom approximation¶
The system to solve is .
Because is a LowRankUpdate, gaussx exploits the
Woodbury identity automatically, giving cost instead of
.
# Build K_nys + sigma^2 I as a LowRankUpdate with diagonal base
K_noisy = gaussx.LowRankUpdate(
base=lx.DiagonalLinearOperator(jnp.full(n_train, noise_var)),
U=K_nys.U,
d=K_nys.d,
V=K_nys.V,
)
alpha_nys = gaussx.solve(K_noisy, y_train)
# Predictions at test points
K_star_Z = rbf_kernel(x_plot, z_inducing, lengthscale, variance)
K_star_train = rbf_kernel(x_plot, x_train, lengthscale, variance)
mu_nys = K_star_train @ alpha_nys
# Full GP for reference
K_noisy_full = lx.MatrixLinearOperator(K_full + noise_var * jnp.eye(n_train))
alpha_full = gaussx.solve(K_noisy_full, y_train)
mu_full = K_star_train @ alpha_fullfig, ax = plt.subplots(figsize=(10, 4))
ax.plot(x_plot, f_true(x_plot), "k--", lw=1.5, label="True function", zorder=4)
ax.plot(x_plot, mu_full, "C0-", lw=2, label="Full GP", zorder=3)
ax.plot(x_plot, mu_nys, "C1--", lw=2, label=f"Nystrom GP (M={m_inducing})", zorder=3)
ax.scatter(
x_train,
y_train,
s=30,
c="grey",
edgecolors="k",
linewidths=0.5,
alpha=0.3,
zorder=5,
)
ax.scatter(
z_inducing,
jnp.zeros_like(z_inducing),
marker="|",
s=100,
c="C1",
zorder=5,
label="Inducing points",
)
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.legend(fontsize=9)
ax.set_title("GP Predictions: Full vs Nystrom")
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.tight_layout()
plt.show()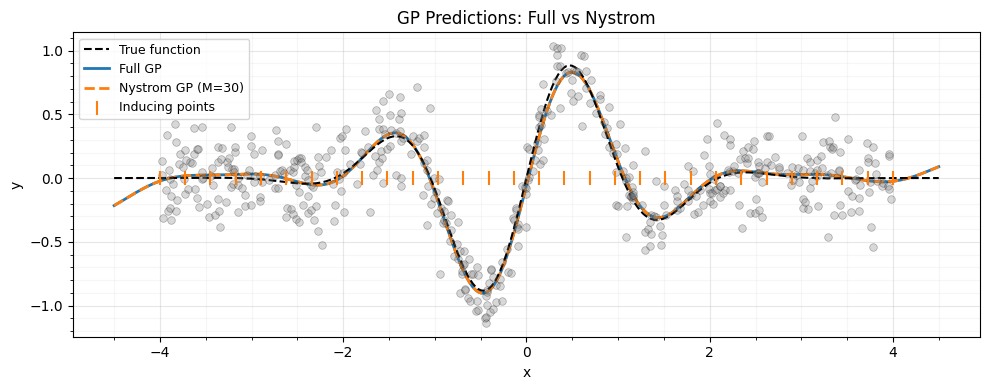
4. Random Fourier Features¶
For the RBF kernel with lengthscale , we sample and .
k_rff = jax.random.PRNGKey(99)
D_rff = 500
k_omega, k_b = jax.random.split(k_rff)
omega = jax.random.normal(k_omega, (D_rff, 1)) / lengthscale # (D_rff, D=1)
b = jax.random.uniform(k_b, (D_rff,), minval=0.0, maxval=2.0 * jnp.pi)
# X must be (N, D) -- reshape 1D inputs
X_2d = x_train[:, None] # (N, 1)
K_rff = gaussx.rff_operator(X_2d, omega, b)
print(f"RFF operator type: {type(K_rff).__name__}")RFF operator type: LowRankUpdate
K_rff_dense = K_rff.as_matrix()
frob_rff = jnp.linalg.norm(K_full - K_rff_dense, "fro")
frob_rff_rel = frob_rff / jnp.linalg.norm(K_full, "fro")
print(f"RFF Frobenius error (D={D_rff}): {frob_rff:.4f}")
print(f"RFF relative error: {frob_rff_rel:.4f}")RFF Frobenius error (D=500): 24.3464
RFF relative error: 0.1189
Convergence with increasing feature dimension¶
As grows the approximation improves at rate.
D_values = [100, 500, 1000]
errors = []
for D in D_values:
k_o, k_bb = jax.random.split(jax.random.PRNGKey(D))
om = jax.random.normal(k_o, (D, 1)) / lengthscale
bb = jax.random.uniform(k_bb, (D,), minval=0.0, maxval=2.0 * jnp.pi)
K_approx = gaussx.rff_operator(X_2d, om, bb).as_matrix()
err = jnp.linalg.norm(K_full - K_approx, "fro") / jnp.linalg.norm(K_full, "fro")
errors.append(float(err))
print(f" D_rff = {D:5d} | relative error = {err:.4f}") D_rff = 100 | relative error = 0.3016
D_rff = 500 | relative error = 0.0947
D_rff = 1000 | relative error = 0.0488
fig, ax = plt.subplots(figsize=(10, 4))
ax.loglog(D_values, errors, "o-", color="C2", lw=2)
ax.set_xlabel("Number of random features (D)")
ax.set_ylabel("Relative Frobenius error")
ax.set_title("RFF Convergence")
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.tight_layout()
plt.show()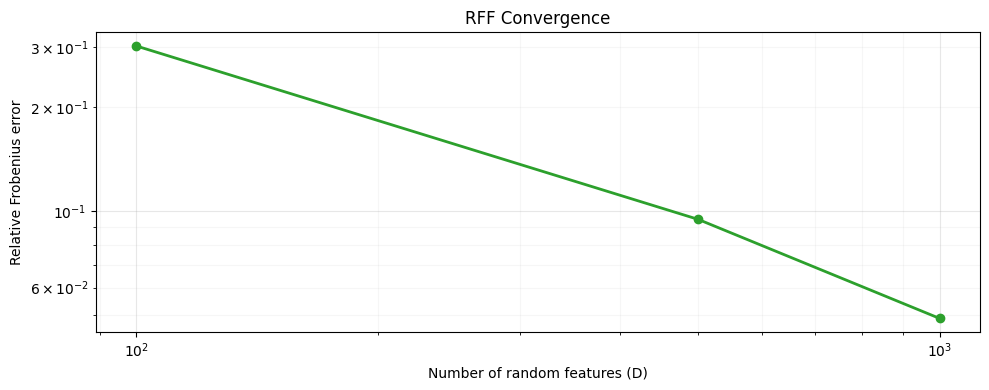
5. Kernel Centering¶
Centering a kernel matrix is required for kernel PCA and many kernel methods. The centered kernel is where is the centering matrix.
n = n_train
# Build the centering operator H = I - (1/n) 11^T
H_op = gaussx.centering_operator(n)
print(f"Centering operator type: {type(H_op).__name__}")
H_dense = H_op.as_matrix()
print(f"H shape: {H_dense.shape}")
# Verify it is I - (1/n) 11^T
H_manual = jnp.eye(n) - jnp.ones((n, n)) / n
print(f"H matches manual: {jnp.allclose(H_dense, H_manual, atol=1e-12)}")Centering operator type: LowRankUpdate
H shape: (500, 500)
H matches manual: True
Center the kernel matrix¶
K_centered_op = gaussx.center_kernel(K_full_op)
K_centered = K_centered_op.as_matrix()
# Manual centering: HKH
K_centered_manual = H_manual @ K_full @ H_manual
print(f"Centered kernel type: {type(K_centered_op).__name__}")
print(f"Matches manual HKH: {jnp.allclose(K_centered, K_centered_manual, atol=1e-10)}")
print(f"Centered row sums ~ 0: {jnp.abs(K_centered.sum(axis=1)).max():.2e}")Centered kernel type: MatrixLinearOperator
Matches manual HKH: True
Centered row sums ~ 0: 8.26e-14
6. HSIC: Independence Testing¶
The Hilbert-Schmidt Independence Criterion measures dependence between two random variables via their kernel embeddings:
HSIC if and only if and are independent (in the population, with characteristic kernels). We compare a correlated pair with an independent pair .
key_hsic = jax.random.PRNGKey(7)
k1, k2, k3 = jax.random.split(key_hsic, 3)
n_hsic = 200
x_hsic = jax.random.normal(k1, (n_hsic,))
# Correlated pair: y depends on x
y_corr = x_hsic + 0.3 * jax.random.normal(k2, (n_hsic,))
# Independent pair: z is independent of x
z_indep = jax.random.normal(k3, (n_hsic,))
# Build kernel matrices (RBF, lengthscale=1)
K_x = rbf_kernel(x_hsic, x_hsic, 1.0, 1.0)
K_y_corr = rbf_kernel(y_corr, y_corr, 1.0, 1.0)
K_z_indep = rbf_kernel(z_indep, z_indep, 1.0, 1.0)
K_x_op = lx.MatrixLinearOperator(K_x)
K_y_op = lx.MatrixLinearOperator(K_y_corr)
K_z_op = lx.MatrixLinearOperator(K_z_indep)hsic_corr = gaussx.hsic(K_x_op, K_y_op)
hsic_indep = gaussx.hsic(K_x_op, K_z_op)
print(f"HSIC (x, y_correlated): {hsic_corr:.6f} (should be > 0)")
print(f"HSIC (x, z_independent): {hsic_indep:.6f} (should be ~ 0)")HSIC (x, y_correlated): 0.055927 (should be > 0)
HSIC (x, z_independent): 0.002026 (should be ~ 0)
7. MMD: Two-Sample Testing¶
The Maximum Mean Discrepancy measures the distance between two distributions in a reproducing kernel Hilbert space:
We compare vs (different) and vs (same).
key_mmd = jax.random.PRNGKey(13)
k1, k2, k3 = jax.random.split(key_mmd, 3)
n_mmd = 200
x_p = jax.random.normal(k1, (n_mmd,)) # P = N(0, 1)
y_q = jax.random.normal(k2, (n_mmd,)) + 1.0 # Q = N(1, 1)
x_p2 = jax.random.normal(k3, (n_mmd,)) # Another sample from P
# Kernel matrices (raw arrays, not operators)
ls_mmd = 1.0
K_xx = rbf_kernel(x_p, x_p, ls_mmd, 1.0)
K_yy = rbf_kernel(y_q, y_q, ls_mmd, 1.0)
K_xy = rbf_kernel(x_p, y_q, ls_mmd, 1.0)
K_xx2 = rbf_kernel(x_p, x_p, ls_mmd, 1.0)
K_pp2 = rbf_kernel(x_p2, x_p2, ls_mmd, 1.0)
K_xp2 = rbf_kernel(x_p, x_p2, ls_mmd, 1.0)mmd_diff = gaussx.mmd_squared(K_xx, K_yy, K_xy)
mmd_same = gaussx.mmd_squared(K_xx2, K_pp2, K_xp2)
print(f"MMD^2(P, Q) [different]: {mmd_diff:.6f} (should be > 0)")
print(f"MMD^2(P, P) [same]: {mmd_same:.6f} (should be ~ 0)")MMD^2(P, Q) [different]: 0.148538 (should be > 0)
MMD^2(P, P) [same]: 0.004874 (should be ~ 0)
8. Plots: Approximation Quality¶
A side-by-side comparison of the kernel matrix approximations.
fig, axes = plt.subplots(1, 3, figsize=(12, 3.5))
vmin, vmax = K_full.min(), K_full.max()
kw = dict(vmin=vmin, vmax=vmax, cmap="viridis", aspect="auto")
axes[0].imshow(K_full, **kw)
axes[0].set_title("Full Kernel")
axes[1].imshow(K_nys_dense, **kw)
axes[1].set_title(f"Nystrom (M={m_inducing})")
axes[2].imshow(K_rff_dense, **kw)
axes[2].set_title(f"RFF (D={D_rff})")
for ax in axes:
ax.set_xlabel("Index")
ax.set_ylabel("Index")
plt.tight_layout()
plt.show()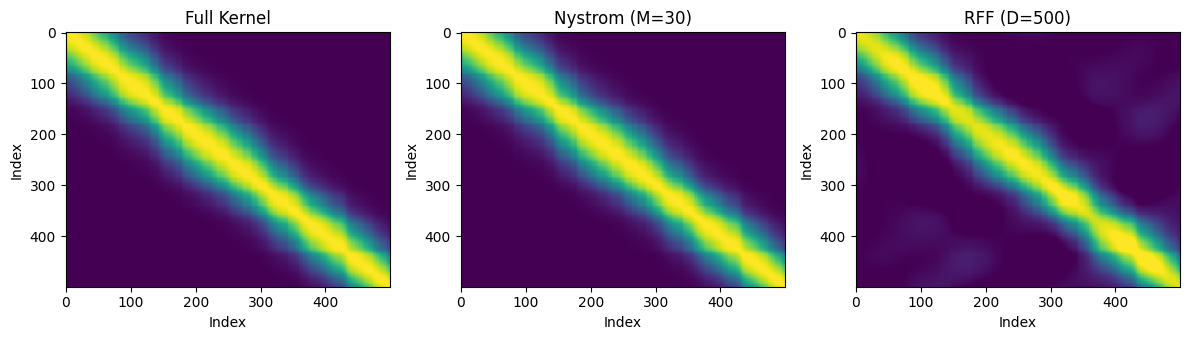
9. Summary¶
| Function | Output | Cost |
|---|---|---|
nystrom_operator(K_XZ, K_ZZ_op) | LowRankUpdate | |
rff_operator(X, omega, b) | LowRankUpdate | |
center_kernel(K) | MatrixLinearOperator | |
centering_operator(n) | LowRankUpdate | |
hsic(K_f, K_q) | scalar | |
mmd_squared(K_xx, K_yy, K_xy) | scalar |
Key takeaways:
- Both Nystrom and RFF produce
LowRankUpdateoperators, so downstream solves and log-determinants exploit the Woodbury identity automatically. center_kernelandcentering_operatorhandle the centering boilerplate needed by kernel PCA and HSIC.hsicandmmd_squaredare one-line kernel statistics for independence and two-sample testing.