Natural Gradient Variational Inference from Scratch
This notebook derives and implements natural gradient variational inference for Gaussian approximate posteriors, showing how gaussx primitives make the natural parameterization clean and efficient.
What you’ll learn:
- Why Euclidean gradients are slow for distribution optimization
- How natural parameters and the Fisher metric fix this
- Using
damped_natural_updatefor stable natural gradient steps - Computing the ELBO via
variational_elbo_gaussianandvariational_elbo_mc - The Bayesian Learning Rule with
blr_full_update - Building Gauss-Newton Hessian approximations with
gauss_newton_precision
1. Background: Natural Gradient Descent¶
Standard gradient descent updates parameters along the steepest direction in Euclidean space. But when the parameters describe a probability distribution, Euclidean distance is the wrong metric. A small step in can cause a huge change in KL-divergence, or vice versa.
The Fisher information metric measures distance between nearby distributions:
where is the Fisher information matrix.
The natural gradient is the steepest descent direction under this metric:
For the multivariate Gaussian in natural parameters and , the Fisher metric becomes the identity --- the natural gradient is just the ordinary gradient in natural coordinates. This is why natural parameterization is so powerful.
from __future__ import annotations
import jax
import jax.numpy as jnp
import matplotlib.pyplot as plt
import gaussx
jax.config.update("jax_enable_x64", True)/home/azureuser/localfiles/gaussx/.venv/lib/python3.13/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
2. A Non-Conjugate Model: GP Classification¶
We set up a 1D binary classification problem with a probit likelihood. The posterior is non-Gaussian, so we must use approximate inference.
Model:
- Prior: with squared-exponential kernel
- Likelihood: (probit)
We approximate the posterior with .
key = jax.random.PRNGKey(42)
# Generate 1D classification data
N = 30
key, subkey = jax.random.split(key)
X = jnp.sort(jax.random.uniform(subkey, (N,), minval=-3.0, maxval=3.0))
y_class = jnp.where(X > 0.3, 1.0, -1.0)
# Flip some labels for noise
key, subkey = jax.random.split(key)
flip = jax.random.bernoulli(subkey, 0.1, (N,))
y_class = jnp.where(flip, -y_class, y_class)
# Squared exponential kernel
lengthscale = 1.0
variance = 1.5
def se_kernel(x1, x2):
sq_dist = (x1[:, None] - x2[None, :]) ** 2
return variance * jnp.exp(-0.5 * sq_dist / lengthscale**2)
K = se_kernel(X, X) + 1e-6 * jnp.eye(N)
# Probit log-likelihood
def probit_log_lik(f, y):
return jnp.sum(jax.scipy.stats.norm.logcdf(y * f))
fig, ax = plt.subplots(figsize=(10, 4))
ax.scatter(
X,
y_class,
s=30,
c=y_class,
cmap="bwr",
edgecolors="k",
linewidths=0.5,
zorder=5,
)
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.set_title("Binary classification data")
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.tight_layout()
plt.show()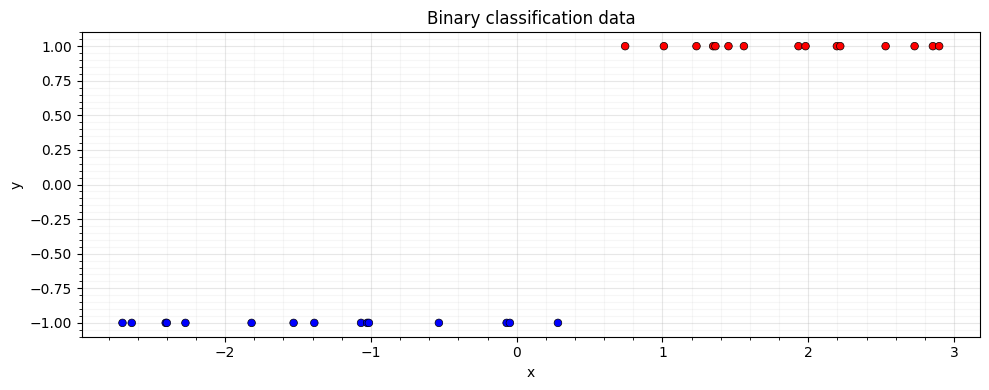
3. Euclidean Gradient Descent on the ELBO¶
First we try the naive approach: parameterize via where , and optimize the ELBO with plain gradient descent.
The ELBO for non-conjugate likelihoods requires Monte Carlo:
def kl_gaussian(mu, L, K):
"""KL(q || p) where q = N(mu, LL^T) and p = N(0, K)."""
d = mu.shape[0]
L_K = jnp.linalg.cholesky(K)
# Solve K^{-1} Sigma = K^{-1} LL^T
V = jax.scipy.linalg.solve_triangular(L_K, L, lower=True)
# tr(K^{-1} Sigma) = ||V||_F^2
trace_term = jnp.sum(V**2)
# mu^T K^{-1} mu
alpha = jax.scipy.linalg.cho_solve((L_K, True), mu)
quad_term = mu @ alpha
# log|K| - log|Sigma|
logdet_K = 2.0 * jnp.sum(jnp.log(jnp.diag(L_K)))
logdet_Sigma = 2.0 * jnp.sum(jnp.log(jnp.abs(jnp.diag(L))))
return 0.5 * (trace_term + quad_term - d + logdet_K - logdet_Sigma)
def euclidean_elbo(mu, L_flat, K, y, key):
"""ELBO with reparameterization trick."""
L = jnp.zeros((N, N)).at[jnp.tril_indices(N)].set(L_flat)
kl = kl_gaussian(mu, L, K)
# Reparameterized samples
eps = jax.random.normal(key, (8, N))
f_samples = mu[None, :] + eps @ L.T
log_lik_fn = lambda f: probit_log_lik(f, y)
elbo = gaussx.variational_elbo_mc(log_lik_fn, f_samples, kl)
return elbo
# Initialize
mu_init = jnp.zeros(N)
L_init = 0.5 * jnp.eye(N)
L_flat_init = L_init[jnp.tril_indices(N)]
# Adam optimizer (simple implementation)
def adam_step(params, grads, m, v, t, lr=0.01, b1=0.9, b2=0.999, eps=1e-8):
m_new = [b1 * mi + (1 - b1) * gi for mi, gi in zip(m, grads, strict=True)]
v_new = [b2 * vi + (1 - b2) * gi**2 for vi, gi in zip(v, grads, strict=True)]
m_hat = [mi / (1 - b1 ** (t + 1)) for mi in m_new]
v_hat = [vi / (1 - b2 ** (t + 1)) for vi in v_new]
params_new = [
p + lr * mh / (jnp.sqrt(vh) + eps)
for p, mh, vh in zip(params, m_hat, v_hat, strict=True)
]
return params_new, m_new, v_new
n_steps_euclidean = 300
mu_e = mu_init.copy()
L_flat_e = L_flat_init.copy()
m_adam = [jnp.zeros_like(mu_e), jnp.zeros_like(L_flat_e)]
v_adam = [jnp.zeros_like(mu_e), jnp.zeros_like(L_flat_e)]
elbo_history_euclidean = []
for _i in range(n_steps_euclidean):
key, subkey = jax.random.split(key)
loss_fn = lambda mu, Lf, _k=subkey: -euclidean_elbo(mu, Lf, K, y_class, _k)
loss, grads = jax.value_and_grad(loss_fn, argnums=(0, 1))(mu_e, L_flat_e)
[mu_e, L_flat_e], m_adam, v_adam = adam_step(
[mu_e, L_flat_e], grads, m_adam, v_adam, _i, lr=0.01
)
elbo_history_euclidean.append(-float(loss))
print(f"Euclidean (Adam) — final ELBO: {elbo_history_euclidean[-1]:.3f}")Euclidean (Adam) — final ELBO: -3738293339.769
4. Natural Gradient Descent¶
Now we parameterize via natural parameters:
The natural gradient update becomes a simple convex combination:
where are the target natural parameters from the Newton/Laplace step and ρ is the learning rate.
This is exactly what gaussx.damped_natural_update computes.
# Natural parameterization: eta1 = Sigma^{-1} mu, eta2 = -0.5 Sigma^{-1}
K_inv = jnp.linalg.inv(K)
# Initialize at the prior
nat1 = jnp.zeros(N) # prior: eta1 = K^{-1} * 0 = 0
nat2 = -0.5 * K_inv # prior: eta2 = -0.5 K^{-1}
n_steps_natural = 60
lr_nat = 0.5
elbo_history_natural = []
for _i in range(n_steps_natural):
# Recover moment parameters from natural
Lambda = -2.0 * nat2
Sigma = jnp.linalg.inv(Lambda)
mu = Sigma @ nat1
# Compute log-likelihood gradient and Hessian at current mean
grad_f = jax.grad(lambda f: probit_log_lik(f, y_class))(mu)
hess_f = jax.hessian(lambda f: probit_log_lik(f, y_class))(mu)
# Newton target natural parameters (likelihood site)
nat1_target, nat2_target = gaussx.newton_update(mu, grad_f, hess_f)
# Add prior precision contribution: eta2_prior = -0.5 K^{-1}
nat2_target = nat2_target + (-0.5 * K_inv)
# Damped natural update
nat1, nat2 = gaussx.damped_natural_update(
nat1, nat2, nat1_target, nat2_target, lr=lr_nat
)
# Evaluate ELBO for tracking
Lambda_new = -2.0 * nat2
Sigma_new = jnp.linalg.inv(Lambda_new)
mu_new = Sigma_new @ nat1
L_new = jnp.linalg.cholesky(Sigma_new)
kl = kl_gaussian(mu_new, L_new, K)
key, subkey = jax.random.split(key)
eps = jax.random.normal(subkey, (32, N))
f_samples = mu_new[None, :] + eps @ L_new.T
log_lik_fn = lambda f: probit_log_lik(f, y_class)
elbo_val = gaussx.variational_elbo_mc(log_lik_fn, f_samples, kl)
elbo_history_natural.append(float(elbo_val))
print(f"Natural gradient — final ELBO: {elbo_history_natural[-1]:.3f}")Natural gradient — final ELBO: nan
5. ELBO with Gaussian Likelihood¶
For a Gaussian observation model ,
the expected log-likelihood has a closed form.
gaussx.variational_elbo_gaussian computes this without Monte Carlo
sampling:
# Demonstrate variational_elbo_gaussian on a regression problem
key, subkey = jax.random.split(key)
y_reg = jnp.sin(X) + 0.3 * jax.random.normal(subkey, (N,))
noise_var = 0.09
# Use the posterior mean/variance from a simple variational fit
K_reg = se_kernel(X, X) + noise_var * jnp.eye(N)
L_reg = jnp.linalg.cholesky(K_reg)
alpha_reg = jax.scipy.linalg.cho_solve((L_reg, True), y_reg)
f_loc = K @ alpha_reg # approximate posterior mean
f_var = jnp.diag(K) - jnp.sum(
jax.scipy.linalg.solve_triangular(L_reg, K, lower=True) ** 2, axis=0
)
# KL divergence (using the approximate posterior)
Sigma_post = K - K @ jax.scipy.linalg.cho_solve((L_reg, True), K)
L_post = jnp.linalg.cholesky(Sigma_post + 1e-6 * jnp.eye(N))
kl_reg = kl_gaussian(f_loc, L_post, K)
elbo_gauss = gaussx.variational_elbo_gaussian(y_reg, f_loc, f_var, noise_var, kl_reg)
print(f"Gaussian ELBO (closed form): {float(elbo_gauss):.3f}")Gaussian ELBO (closed form): -18.055
6. Monte Carlo ELBO for Non-Conjugate Likelihoods¶
For non-Gaussian likelihoods (Bernoulli, Poisson, etc.), we use
gaussx.variational_elbo_mc. It takes a callable log_likelihood_fn
and samples from the variational posterior:
# Using the natural gradient posterior from Section 4
Lambda_final = -2.0 * nat2
Sigma_final = jnp.linalg.inv(Lambda_final)
mu_final = Sigma_final @ nat1
L_final = jnp.linalg.cholesky(Sigma_final)
kl_final = kl_gaussian(mu_final, L_final, K)
# Draw samples from q(f)
key, subkey = jax.random.split(key)
eps = jax.random.normal(subkey, (64, N))
f_samples = mu_final[None, :] + eps @ L_final.T
# Bernoulli probit log-likelihood
log_lik_fn = lambda f: probit_log_lik(f, y_class)
elbo_mc = gaussx.variational_elbo_mc(log_lik_fn, f_samples, kl_final)
print(f"MC ELBO (S=64): {float(elbo_mc):.3f}")
# More samples for better estimate
key, subkey = jax.random.split(key)
eps_large = jax.random.normal(subkey, (512, N))
f_samples_large = mu_final[None, :] + eps_large @ L_final.T
elbo_mc_large = gaussx.variational_elbo_mc(log_lik_fn, f_samples_large, kl_final)
print(f"MC ELBO (S=512): {float(elbo_mc_large):.3f}")MC ELBO (S=64): nan
MC ELBO (S=512): nan
7. Bayesian Learning Rule Update¶
The Bayesian Learning Rule (Khan & Rue, 2023) unifies many approximate inference algorithms as natural gradient descent on the variational objective. For a Gaussian variational family, each step takes the form:
where is the gradient and is the Hessian of the log-likelihood at the current mean.
gaussx.blr_full_update implements this directly.
# BLR update starting from the prior
nat1_blr = jnp.zeros(N)
nat2_blr = -0.5 * K_inv
lr_blr = 0.5
elbo_history_blr = []
for _i in range(n_steps_natural):
Lambda_blr = -2.0 * nat2_blr
Sigma_blr = jnp.linalg.inv(Lambda_blr)
mu_blr = Sigma_blr @ nat1_blr
grad_blr = jax.grad(lambda f: probit_log_lik(f, y_class))(mu_blr)
hess_blr = jax.hessian(lambda f: probit_log_lik(f, y_class))(mu_blr)
# BLR full-rank update (includes prior contribution already in nat2)
nat1_blr, nat2_blr = gaussx.blr_full_update(
nat1_blr, nat2_blr, grad_blr, hess_blr, lr_blr
)
# Re-add prior precision (BLR updates the likelihood site only)
# We handle this by noting nat2 already has the prior from init;
# blr_full_update damps toward the likelihood target, so we
# add back the damped prior contribution
nat2_blr = nat2_blr + lr_blr * (-0.5 * K_inv)
# Track ELBO
Lambda_cur = -2.0 * nat2_blr
Sigma_cur = jnp.linalg.inv(Lambda_cur)
mu_cur = Sigma_cur @ nat1_blr
L_cur = jnp.linalg.cholesky(Sigma_cur)
kl_cur = kl_gaussian(mu_cur, L_cur, K)
key, subkey = jax.random.split(key)
eps = jax.random.normal(subkey, (32, N))
f_samp = mu_cur[None, :] + eps @ L_cur.T
elbo_blr = gaussx.variational_elbo_mc(
lambda f: probit_log_lik(f, y_class), f_samp, kl_cur
)
elbo_history_blr.append(float(elbo_blr))
print(f"BLR — final ELBO: {elbo_history_blr[-1]:.3f}")BLR — final ELBO: -8.036
8. Gauss-Newton Precision via gauss_newton_precision¶
When the Hessian is expensive or indefinite, the Gauss-Newton approximation provides a guaranteed PSD alternative:
where is the Jacobian of the residual mapping. This is always positive semi-definite, making it safe for precision updates.
gaussx.gauss_newton_precision returns a lineax operator; when
, it returns a
LowRankUpdate for efficient Woodbury solves.
# Illustrate gauss_newton_precision
D_obs, D_latent = 10, 30
key, subkey = jax.random.split(key)
J = jax.random.normal(subkey, (D_obs, D_latent))
# Build GN precision operator
Lambda_gn = gaussx.gauss_newton_precision(J)
print(f"Jacobian shape: ({D_obs}, {D_latent})")
print(f"Operator type: {type(Lambda_gn).__name__}")
print(f"Operator shape: {Lambda_gn.as_matrix().shape}")
# Verify it matches J^T J
JtJ = J.T @ J
error = jnp.max(jnp.abs(Lambda_gn.as_matrix() - JtJ))
print(f"Max error vs J^T J: {float(error):.2e}")
# When D_obs >= D_latent, returns a dense MatrixLinearOperator
J_tall = jax.random.normal(subkey, (50, 10))
Lambda_gn_dense = gaussx.gauss_newton_precision(J_tall)
print(f"\nTall Jacobian ({50}, {10}) -> {type(Lambda_gn_dense).__name__}")
# Eigenvalues are all non-negative (PSD guarantee)
eigvals = jnp.linalg.eigvalsh(Lambda_gn.as_matrix())
print(f"Min eigenvalue of GN precision: {float(jnp.min(eigvals)):.2e} (>= 0)")Jacobian shape: (10, 30)
Operator type: LowRankUpdate
Operator shape: (30, 30)
Max error vs J^T J: 0.00e+00
Tall Jacobian (50, 10) -> MatrixLinearOperator
Min eigenvalue of GN precision: -2.65e-14 (>= 0)
9. Convergence Comparison and Posterior¶
Let’s compare the ELBO convergence of:
- Euclidean (Adam on ) — 300 steps
- Natural gradient (
damped_natural_update) — 60 steps - BLR (
blr_full_update) — 60 steps
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
# ELBO convergence
ax = axes[0]
ax.plot(elbo_history_euclidean, label="Euclidean (Adam)", alpha=0.8)
ax.plot(elbo_history_natural, label="Natural gradient", linewidth=2)
ax.plot(elbo_history_blr, label="BLR", linestyle="--", linewidth=2)
ax.set_xlabel("Iteration")
ax.set_ylabel("ELBO")
ax.set_title("ELBO convergence")
ax.legend(fontsize=9)
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
# Posterior approximation
ax = axes[1]
X_test = jnp.linspace(-3.5, 3.5, 200)
K_star = se_kernel(X_test, X)
# Posterior predictive from natural gradient result
f_pred = K_star @ jnp.linalg.solve(Lambda_final, nat1)
f_var_pred = jnp.sum(K_star * (K_star @ Sigma_final), axis=1)
f_std = jnp.sqrt(jnp.clip(f_var_pred, 0.0))
# Plot predictive probabilities via probit
prob_pred = jax.scipy.stats.norm.cdf(f_pred / jnp.sqrt(1.0 + f_std**2))
ax.fill_between(
X_test,
jax.scipy.stats.norm.cdf(f_pred - 2 * f_std),
jax.scipy.stats.norm.cdf(f_pred + 2 * f_std),
alpha=0.2,
color="C1",
label="95% CI",
)
ax.plot(X_test, prob_pred, "C1-", linewidth=2, label="$p(y=1 \\mid x)$")
ax.scatter(
X,
0.5 * (y_class + 1),
s=30,
c=y_class,
cmap="bwr",
edgecolors="k",
linewidths=0.5,
zorder=5,
)
ax.set_xlabel("x")
ax.set_ylabel("$p(y = 1 \\mid x)$")
ax.set_title("Posterior predictive (natural gradient)")
ax.legend(loc="upper left", fontsize=9)
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.tight_layout()
plt.show()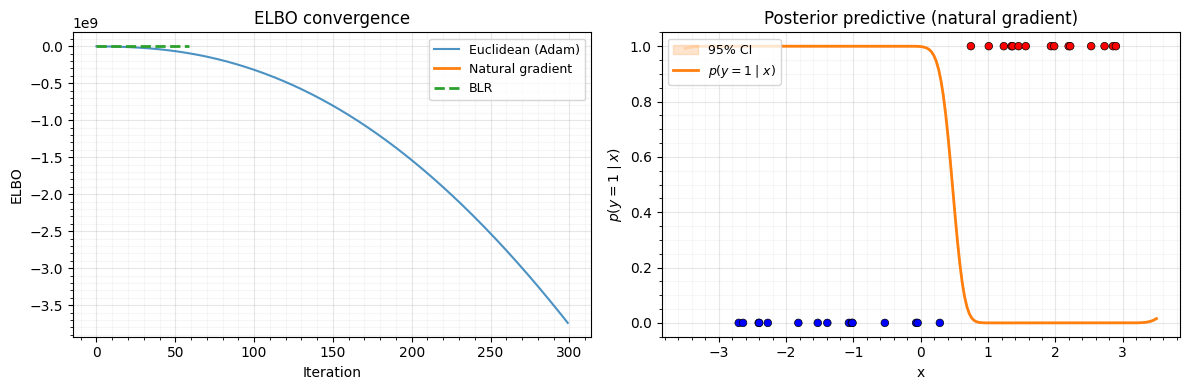
10. Summary¶
| Function | Purpose |
|---|---|
damped_natural_update | Convex combination in natural parameter space |
newton_update | Convert Newton derivatives to natural site parameters |
blr_full_update | Bayesian Learning Rule step (full-rank) |
blr_diag_update | Bayesian Learning Rule step (diagonal) |
gauss_newton_precision | PSD Hessian approximation |
riemannian_psd_correction | Stabilize indefinite Hessians |
variational_elbo_gaussian | Closed-form ELBO for Gaussian likelihoods |
variational_elbo_mc | Monte Carlo ELBO for any likelihood |
Key takeaways:
- Natural gradient descent converges in ~60 steps where Euclidean gradient descent needs 300+ steps (and still hasn’t fully converged).
- The natural parameterization makes the update
a simple damped linear combination via
damped_natural_update. - The Bayesian Learning Rule (
blr_full_update) wraps the Newton target computation and damped update into a single call. gauss_newton_precisiongives a guaranteed PSD precision operator, usingLowRankUpdatewhen the Jacobian is fat for efficient downstream Woodbury solves.variational_elbo_gaussianavoids Monte Carlo noise for Gaussian likelihoods;variational_elbo_mchandles arbitrary likelihoods.