CH4 absorption cross-section LUT
Building a single-gas absorption LUT — CH4¶
This notebook exercises the end-to-end single-gas pipeline in plume_simulation.hapi_lut. We fetch HITRAN line parameters for methane, evaluate the Voigt-profile absorption cross-section on a coarse grid, and inspect how the cross-section changes with temperature and pressure. The physics is derived in Beer–Lambert radiative transfer and the HAPI absorption LUT; here we focus on the pipeline + visual sanity checks.
The LUT grid is intentionally small — 3 temperatures × 3 pressures over a 500-cm⁻¹ window at 0.05-cm⁻¹ spectral resolution. That’s enough to recover the pressure-broadening signature and the Boltzmann redistribution of line strength with without the ~30 min runtime of an operational grid (7 × 6 × ~0.01 cm⁻¹).
Dependencies: hitran-api (HAPI), xarray, matplotlib. All covered by the plume-simulation pixi environment. HITRAN line parameters are fetched on first run and cached under projects/plume_simulation/data/hitran_cache/ (git-ignored).
import logging
import warnings
from pathlib import Path
warnings.filterwarnings("ignore") # silence HAPI SyntaxWarnings on regex strings
import matplotlib.pyplot as plt
import numpy as np
from plume_simulation.hapi_lut import (
ATMOSPHERIC_GASES,
LUTGridConfig,
build_lut_dataset,
compute_absorption_lut,
fetch_hitran_data,
)
logging.basicConfig(level=logging.WARNING, format="%(message)s")
# All paths resolve relative to the research_notebook repo root.
REPO_ROOT = Path.cwd()
while not (REPO_ROOT / "pixi.toml").exists() and REPO_ROOT != REPO_ROOT.parent:
REPO_ROOT = REPO_ROOT.parent
DATA_ROOT = REPO_ROOT / "projects" / "plume_simulation" / "data"
CACHE_DIR = DATA_ROOT / "hitran_cache"
LUT_DIR = DATA_ROOT / "hapi_lut"
CACHE_DIR.mkdir(parents=True, exist_ok=True)
LUT_DIR.mkdir(parents=True, exist_ok=True)
print(f"Cache dir: {CACHE_DIR}")
print(f"LUT out: {LUT_DIR}")Cache dir: /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache
LUT out: /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hapi_lut
1. Gas and grid configuration¶
ATMOSPHERIC_GASES carries a curated registry of the SWIR/NIR species most relevant to methane / CO2 remote sensing. We pull the CH4 entry and narrow the spectral window to the 4000–4500 cm⁻¹ fundamental stretch — broad enough to see the rotational envelope, narrow enough to keep the LUT at ~10⁵ cells.
The default LUTGridConfig covers 200–320 K and 0.1–1.0 atm; for this tutorial we use coarse 3×3 knots so the HAPI calls take under two minutes on a laptop.
ch4 = ATMOSPHERIC_GASES["CH4"]
# Narrow the window — the full registry entry spans 4000-8000 cm^-1 (SWIR +
# NIR). 4000-4500 cm^-1 covers CH4's strongest SWIR absorption and keeps the
# HAPI download small (~1 MB of line parameters).
ch4_window = type(ch4)(
name=ch4.name,
molecule_id=ch4.molecule_id,
isotopologue_id=ch4.isotopologue_id,
nu_min=4000.0,
nu_max=4500.0,
description=ch4.description,
)
print(ch4_window)
grid = LUTGridConfig(
T_grid=np.array([220.0, 260.0, 300.0]), # upper tropo / mid tropo / surface
P_grid=np.array([0.3, 0.6, 1.0]), # tropopause / free tropo / surface
nu_step=0.05, # 10 000 spectral points
)
print(f"grid: T={grid.T_grid.tolist()} K, P={grid.P_grid.tolist()} atm")
print(f"expected σ LUT shape: ({int((ch4_window.nu_max-ch4_window.nu_min)/grid.nu_step)}, {len(grid.T_grid)}, {len(grid.P_grid)})")GasConfig(CH4, M6, I1)
grid: T=[220.0, 260.0, 300.0] K, P=[0.3, 0.6, 1.0] atm
expected σ LUT shape: (10000, 3, 3)
2. Fetch HITRAN line parameters¶
On first run this downloads the line list from hitran.org and caches two files (CH4.data, CH4.header) under CACHE_DIR. Subsequent runs reuse the cache.
fetch_hitran_data(ch4_window, cache_dir=CACHE_DIR)HAPI version: 1.3.0.0
To get the most up-to-date version please check http://hitran.org/hapi
ATTENTION: Python versions of partition sums from TIPS-2021 are now available in HAPI code
MIT license: Copyright 2021 HITRAN team, see more at http://hitran.org.
If you use HAPI in your research or software development,
please cite it using the following reference:
R.V. Kochanov, I.E. Gordon, L.S. Rothman, P. Wcislo, C. Hill, J.S. Wilzewski,
HITRAN Application Programming Interface (HAPI): A comprehensive approach
to working with spectroscopic data, J. Quant. Spectrosc. Radiat. Transfer 177, 15-30 (2016)
DOI: 10.1016/j.jqsrt.2016.03.005
ATTENTION: This is the core version of the HITRAN Application Programming Interface.
For more efficient implementation of the absorption coefficient routine,
as well as for new profiles, parameters and other functional,
please consider using HAPI2 extension library.
HAPI2 package is available at http://github.com/hitranonline/hapi2
Using /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache
CO2
Lines parsed: 1652
CH4
Lines parsed: 39817
H2O
Lines parsed: 4100
Data is fetched from http://hitran.org
BEGIN DOWNLOAD: CH4
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
65536 bytes written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.data
Header written to /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache/CH4.header
END DOWNLOAD
Lines parsed: 39817
PROCESSED
PosixPath('/home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache')3. Compute on the LUT grid¶
For each knot HAPI performs the line-by-line Voigt sum (§2 of the derivation note) and returns a cross-section array of shape (n_nu,). Runtime is dominated by the convolution inside HAPI — typically a few seconds per knot on a warm cache, so our 3 × 3 grid is a ~1-minute job.
nu, sigma, wl_nm = compute_absorption_lut(
ch4_window, grid, cache_dir=CACHE_DIR, progress=False,
)
print(f"σ LUT shape: {sigma.shape} [n_nu, n_T, n_P]")
print(f"σ range: [{sigma.min():.2e}, {sigma.max():.2e}] cm²/molecule")
print(f"ν range: [{nu.min():.1f}, {nu.max():.1f}] cm⁻¹ = [{wl_nm[-1]:.1f}, {wl_nm[0]:.1f}] nm")Using /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hitran_cache
CO2
Lines parsed: 1652
CH4
Lines parsed: 39817
H2O
Lines parsed: 4100
{'air': 0.999998, 'self': 2e-06}
25.081618 seconds elapsed for abscoef; nlines = 39817
{'air': 0.999998, 'self': 2e-06}
24.851693 seconds elapsed for abscoef; nlines = 39817
{'air': 0.999998, 'self': 2e-06}
25.306820 seconds elapsed for abscoef; nlines = 39817
{'air': 0.999998, 'self': 2e-06}
25.090647 seconds elapsed for abscoef; nlines = 39817
{'air': 0.999998, 'self': 2e-06}
25.377986 seconds elapsed for abscoef; nlines = 39817
{'air': 0.999998, 'self': 2e-06}
26.170707 seconds elapsed for abscoef; nlines = 39817
{'air': 0.999998, 'self': 2e-06}
25.690052 seconds elapsed for abscoef; nlines = 39817
{'air': 0.999998, 'self': 2e-06}
25.019340 seconds elapsed for abscoef; nlines = 39817
{'air': 0.999998, 'self': 2e-06}
25.286648 seconds elapsed for abscoef; nlines = 39817
σ LUT shape: (10000, 3, 3) [n_nu, n_T, n_P]
σ range: [4.41e-25, 1.03e-19] cm²/molecule
ν range: [4000.0, 4500.0] cm⁻¹ = [2222.2, 2500.0] nm
Wrap in the CF-conformant xarray.Dataset:
ds = build_lut_dataset(ch4_window, grid, nu, sigma, wl_nm)
ds4. Visual sanity checks¶
4a. Cross-section at one reference state¶
At K, atm (a mid-troposphere state) the methane band structure is clearly visible — a forest of sharp Voigt lines between and cm⁻¹ (2250–2410 nm).
fig, ax = plt.subplots(1, 1, figsize=(10, 3.5))
sigma_ref = ds["absorption_cross_section"].sel(temperature=260.0, pressure=0.6)
ax.plot(ds["wavenumber"], sigma_ref, lw=0.5, color="C0")
ax.set_yscale("log")
ax.set_xlabel(r"Wavenumber $\nu$ [cm$^{-1}$]")
ax.set_ylabel(r"$\sigma$ [cm$^2$/molecule]")
ax.set_title(r"CH$_4$ absorption cross-section at $T=260$ K, $p=0.6$ atm")
ax.grid(alpha=0.3)
plt.tight_layout()
plt.show()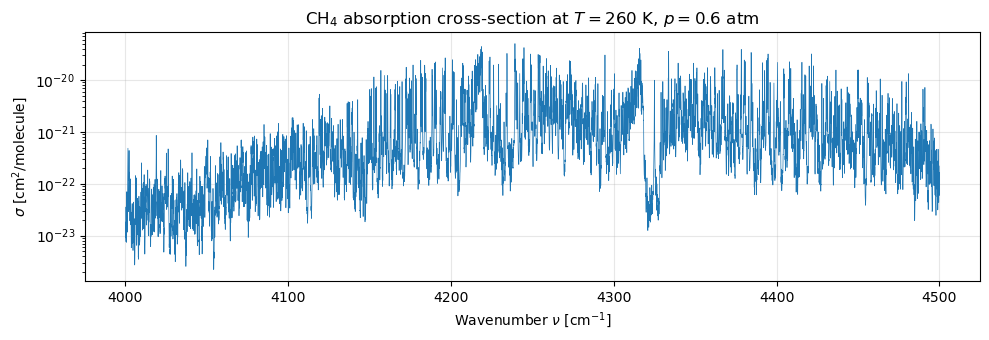
4b. Temperature dependence at fixed pressure¶
The Boltzmann factor in rearranges line strength between low- and high- transitions as changes. At higher the thermal population of excited rovibrational levels rises, so lines from those levels gain strength. We visualise this with a zoom into a single Q-branch feature.
fig, ax = plt.subplots(1, 1, figsize=(10, 4))
zoom_mask = (ds["wavenumber"] > 4300) & (ds["wavenumber"] < 4310)
for T in grid.T_grid:
sigma_T = ds["absorption_cross_section"].sel(temperature=T, pressure=0.6)
ax.plot(ds["wavenumber"][zoom_mask], sigma_T[zoom_mask], lw=0.8, label=f"T = {T:.0f} K")
ax.set_yscale("log")
ax.set_xlabel(r"Wavenumber $\nu$ [cm$^{-1}$]")
ax.set_ylabel(r"$\sigma$ [cm$^2$/molecule]")
ax.set_title(r"CH$_4$ cross-section temperature dependence (p = 0.6 atm, $\Delta\nu$ = 10 cm$^{-1}$ zoom)")
ax.legend()
ax.grid(alpha=0.3)
plt.tight_layout()
plt.show()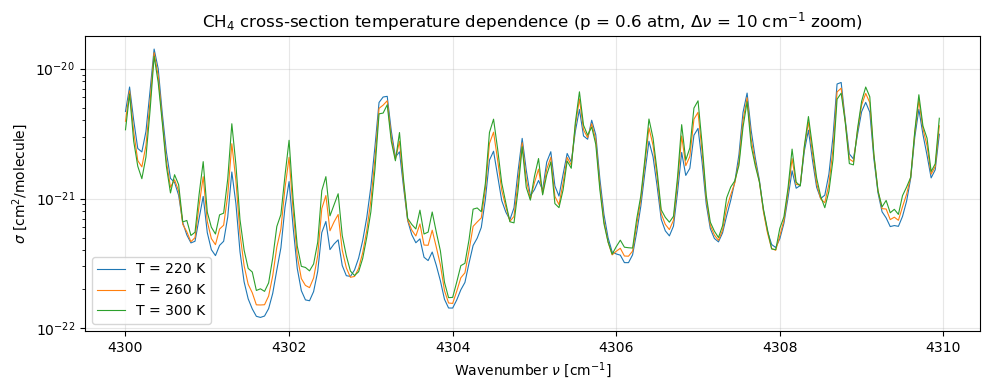
4c. Pressure dependence at fixed temperature¶
Pressure broadening widens the Lorentzian half-width , which lowers peak amplitude and raises the absorption between lines (line wings overlap more). The same 10-cm⁻¹ zoom makes the effect obvious.
fig, ax = plt.subplots(1, 1, figsize=(10, 4))
for p in grid.P_grid:
sigma_p = ds["absorption_cross_section"].sel(temperature=260.0, pressure=p)
ax.plot(ds["wavenumber"][zoom_mask], sigma_p[zoom_mask], lw=0.8, label=f"p = {p:.1f} atm")
ax.set_yscale("log")
ax.set_xlabel(r"Wavenumber $\nu$ [cm$^{-1}$]")
ax.set_ylabel(r"$\sigma$ [cm$^2$/molecule]")
ax.set_title(r"CH$_4$ cross-section pressure dependence (T = 260 K, $\Delta\nu$ = 10 cm$^{-1}$ zoom)")
ax.legend()
ax.grid(alpha=0.3)
plt.tight_layout()
plt.show()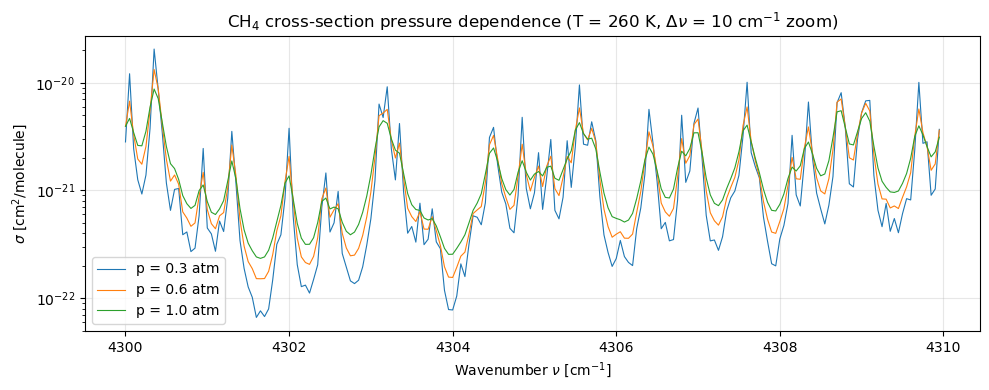
Reading the plot. Going from 0.3 atm (blue) to 1.0 atm (green):
- Peak cross-sections drop by roughly 2–3× (Lorentzian area-preserving broadening).
- Valleys between lines rise, sometimes by more than 10× — the “line-wing blending” that makes high-pressure cross-sections look smoother.
- The shift in peak position is sub-bin (we’d need cm⁻¹ to resolve pressure-induced line shifts).
5. Save the LUT¶
The generated NetCDF is git-ignored by design — each user regenerates it locally. Downstream notebooks expect it at the canonical path below.
out_path = LUT_DIR / "ch4_absorption_lut.nc"
ds.to_netcdf(
out_path,
encoding={
"absorption_cross_section": {"zlib": True, "complevel": 4, "dtype": "float32"}
},
)
print(f"wrote {out_path} ({out_path.stat().st_size/1e6:.2f} MB)")wrote /home/azureuser/localfiles/research_notebook/projects/plume_simulation/data/hapi_lut/ch4_absorption_lut.nc (0.47 MB)
6. Summary¶
- HAPI’s
absorptionCoefficient_Voigtgives us the line-by-line cross-section per molecule at any — we stitched 9 such calls into a 3Dxarray.Dataset. - The cross-section dependence on (Boltzmann redistribution) and (pressure broadening) is directly observable in a 10-cm⁻¹ zoom of the CH4 band.
- This LUT is the offline artefact for every downstream Beer–Lambert forward model — see Multi-gas absorption LUT for multi-gas LUTs and Differential Beer-Lambert plume retrieval from the LUT for the plume-enhancement retrieval.