SVGP + spherical harmonics
SVGP with spherical-harmonic inducing features¶
This is the inter-domain cousin of notebook 1. Everything stays the same — SparseGPPrior, WhitenedGuide, GaussianLikelihood, svgp_elbo, the same optax training loop — except the inducing distribution. Instead of pseudo-inputs in , we use a fixed orthonormal basis of real spherical harmonics on the unit 2-sphere .
The payoff is structural: the inducing-prior covariance is exactly diagonal. The solve that dominates every SVGP forward pass collapses to an elementwise division. No Cholesky. No conditioning issues. No inducing-point migration (there are no points to migrate — the basis is fixed).
This is the VISH construction of Dutordoir, Hensman, van der Wilk, Artemev, Deisenroth, Sheldon 2020. Pyrox ships it as SphericalHarmonicInducingFeatures.
Background — why the solve collapses¶
Mercer and Funk-Hecke¶
Any zonal kernel on — one that depends only on the inner product between unit vectors — admits the Funk-Hecke spectral expansion
with real spherical harmonics and Funk-Hecke coefficients
obtained by Gauss-Legendre quadrature for arbitrary kernels (pyrox does this for you — no closed form required). Pyrox treats a Euclidean kernel like RBF as if it were zonal on the sphere, by evaluating at unit vectors with inner product . The induced kernel has decaying smoothly with .
Inter-domain inducing features¶
Replace inducing values with inducing coefficients
Under Funk-Hecke, the inducing-prior covariance satisfies
and the cross-covariance is . Diagonal means gaussx.solve and gaussx.cholesky short-circuit to elementwise ops — enforced at the lineax operator level by a DiagonalLinearOperator with jitter folded into the diagonal vector (never added as jnp.eye).
Why this matters for ¶
For you get features — comfortably expressive. At you have and each SVGP call still costs in the inducing solve, vs. for a point-inducing SVGP with the same .
Setup¶
import time
import equinox as eqx
import jax
import jax.numpy as jnp
import jax.random as jr
import matplotlib.pyplot as plt
import optax
from jaxtyping import Array, Float
from pyrox.gp import (
GaussianLikelihood,
Kernel,
SparseGPPrior,
SphericalHarmonicInducingFeatures,
WhitenedGuide,
funk_hecke_coefficients,
svgp_elbo,
)
from pyrox.gp._src.kernels import rbf_kernel
jax.config.update("jax_enable_x64", True)/anaconda/lib/python3.13/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
Data on the sphere¶
Target field: a superposition of three von-Mises-Fisher-like bumps at hand-picked directions on — a positive bump near the north pole, a positive bump on the prime meridian equator, and a negative bump on the antimeridian equator. This is zonal around each centre but not globally zonal, so the fit can’t collapse to the trivial mode. Inputs are unit vectors in ; the “input” representation is Cartesian throughout — lat/lon only appears at plot time.
def _vmf_bump(n: Float[Array, "N 3"], centre: Float[Array, " 3"], kappa: float) -> Float[Array, " N"]:
return jnp.exp(kappa * (n @ centre - 1.0))
def f_true(n: Float[Array, "N 3"]) -> Float[Array, " N"]:
c1 = jnp.array([0.0, 0.0, 1.0]) # north pole
c2 = jnp.array([1.0, 0.0, 0.0]) # equator, prime meridian
c3 = jnp.array([-1.0, 0.0, 0.0]) # equator, antimeridian
return _vmf_bump(n, c1, 8.0) + 0.8 * _vmf_bump(n, c2, 15.0) - 0.7 * _vmf_bump(n, c3, 15.0)
def sample_sphere(key: jr.PRNGKey, n: int) -> Float[Array, "n 3"]:
v = jr.normal(key, (n, 3))
return v / jnp.linalg.norm(v, axis=-1, keepdims=True)
key = jr.PRNGKey(0)
key, key_X, key_noise = jr.split(key, 3)
N = 1000
X_train = sample_sphere(key_X, N)
noise_std = 0.05
y_train = f_true(X_train) + noise_std * jr.normal(key_noise, (N,))
print(f"training shapes: X {X_train.shape} y {y_train.shape}")
print(f"|X| sanity: min {float(jnp.linalg.norm(X_train, axis=-1).min()):.6f} "
f"max {float(jnp.linalg.norm(X_train, axis=-1).max()):.6f}")training shapes: X (1000, 3) y (1000,)
|X| sanity: min 1.000000 max 1.000000
Ground truth on a lat-lon grid¶
def unit_from_latlon(lat: Float[Array, "H W"], lon: Float[Array, "H W"]) -> Float[Array, "H W 3"]:
x = jnp.cos(lat) * jnp.cos(lon)
y = jnp.cos(lat) * jnp.sin(lon)
z = jnp.sin(lat)
return jnp.stack([x, y, z], axis=-1)
H, W = 90, 180
lat = jnp.linspace(-jnp.pi / 2, jnp.pi / 2, H)
lon = jnp.linspace(-jnp.pi, jnp.pi, W)
lat_grid, lon_grid = jnp.meshgrid(lat, lon, indexing="ij")
n_grid = unit_from_latlon(lat_grid, lon_grid)
y_grid_true = f_true(n_grid.reshape(-1, 3)).reshape(H, W)
fig = plt.figure(figsize=(9, 4))
ax = fig.add_subplot(111, projection="mollweide")
mesh = ax.pcolormesh(lon_grid, lat_grid, y_grid_true, cmap="RdBu_r", vmin=-1.0, vmax=1.0, shading="auto")
ax.grid(alpha=0.3)
fig.colorbar(mesh, ax=ax, label="f(x)", shrink=0.7)
ax.set_title("Ground-truth field on $S^2$ (Mollweide)")
plt.show()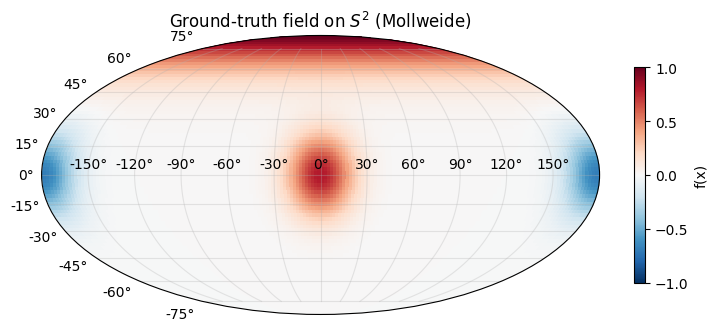
Kernel — same RBFLite as notebook 1¶
We re-use the minimal Pattern-A kernel from notebook 1 verbatim. The SphericalHarmonicInducingFeatures.K_uu constructor calls funk_hecke_coefficients(kernel, l_max) — it only needs kernel(X1, X2) to work on small (1, Q) and (Q, 3) tensors during the Gauss-Legendre sweep, so any Pattern-A Kernel drops in cleanly.
class RBFLite(Kernel, eqx.Module):
"""Pure-equinox RBF — trainable log-variance / log-lengthscale scalars."""
log_variance: Float[Array, ""]
log_lengthscale: Float[Array, ""]
@classmethod
def init(cls, variance: float = 1.0, lengthscale: float = 1.0) -> "RBFLite":
return cls(
log_variance=jnp.log(jnp.asarray(variance)),
log_lengthscale=jnp.log(jnp.asarray(lengthscale)),
)
@property
def variance(self) -> Float[Array, ""]:
return jnp.exp(self.log_variance)
@property
def lengthscale(self) -> Float[Array, ""]:
return jnp.exp(self.log_lengthscale)
def __call__(
self,
X1: Float[Array, "N1 D"],
X2: Float[Array, "N2 D"],
) -> Float[Array, "N1 N2"]:
return rbf_kernel(X1, X2, self.variance, self.lengthscale)
def diag(self, X: Float[Array, "N D"]) -> Float[Array, " N"]:
return self.variance * jnp.ones(X.shape[0], dtype=X.dtype)
class TrainableGaussianLikelihood(eqx.Module):
log_noise_var: Float[Array, ""]
@classmethod
def init(cls, noise_var: float = 0.1) -> "TrainableGaussianLikelihood":
return cls(log_noise_var=jnp.log(jnp.asarray(noise_var)))
def materialise(self) -> GaussianLikelihood:
return GaussianLikelihood(noise_var=jnp.exp(self.log_noise_var))Model¶
l_max = 8 → features. The inducing= kwarg of SparseGPPrior accepts any object implementing the InducingFeatures protocol — SphericalHarmonicInducingFeatures is one of several pyrox ships (FourierInducingFeatures for the box, LaplacianInducingFeatures for graphs, DecoupledInducingFeatures for mean/covariance decoupling).
l_max = 8
sh_features = SphericalHarmonicInducingFeatures.init(l_max=l_max, num_quadrature=256)
M = sh_features.num_features
print(f"l_max = {l_max} → M = (l_max+1)^2 = {M} features")
prior = SparseGPPrior(
kernel=RBFLite.init(variance=1.0, lengthscale=0.8),
inducing=sh_features,
jitter=1e-4,
)
guide = WhitenedGuide.init(num_inducing=M)
lik_w = TrainableGaussianLikelihood.init(noise_var=noise_std**2)l_max = 8 → M = (l_max+1)^2 = 81 features
Kernel spectrum — the Funk-Hecke coefficients¶
Before training, inspect what the chosen RBF “looks like” on the sphere. Funk-Hecke coefficients at tell you how much variance the kernel puts into each harmonic degree — a frequency decomposition of the prior covariance.
a = funk_hecke_coefficients(prior.kernel, l_max=l_max, num_quadrature=256)
fig, ax = plt.subplots(figsize=(8, 3.5))
ax.bar(jnp.arange(l_max + 1), a, color="C0", edgecolor="k")
ax.set_xlabel(r"degree $\ell$")
ax.set_ylabel(r"$a_\ell$")
ax.set_title("Funk-Hecke coefficients of RBF@S² (pre-training)")
ax.grid(alpha=0.3, axis="y")
plt.show()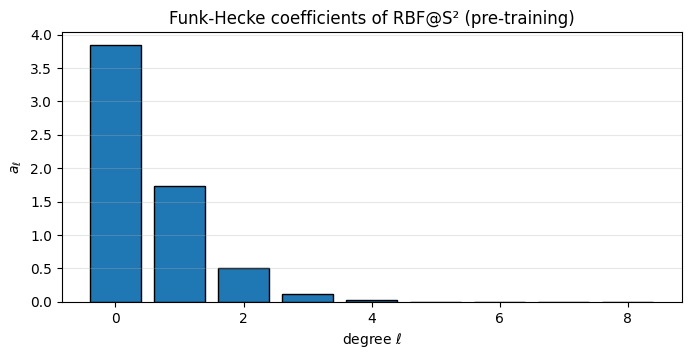
The RBF spectrum decays smoothly — long lengthscales concentrate mass on small (slow variation across the sphere), short lengthscales flatten the spectrum (rapid variation). Training the lengthscale reshapes this bar chart to match the data’s spectral content.
Training — same loop as notebook 1¶
Identical optimiser, step function, loss signature. The SphericalHarmonicInducingFeatures object carries only two static ints (l_max, num_quadrature) so there are no inducing parameters to train — only the kernel, guide, and noise evolve.
Params = tuple
def neg_elbo(params: Params, X: Float[Array, "N 3"], y: Float[Array, " N"]) -> Float[Array, ""]:
prior_p, guide_p, lik_p = params
return -svgp_elbo(prior_p, guide_p, lik_p.materialise(), X, y)
@eqx.filter_jit
def step(
params: Params,
opt_state: optax.OptState,
X: Float[Array, "N 3"],
y: Float[Array, " N"],
) -> tuple[Params, optax.OptState, Float[Array, ""]]:
loss, grads = eqx.filter_value_and_grad(neg_elbo)(params, X, y)
updates, opt_state = optimiser.update(grads, opt_state, params)
params = eqx.apply_updates(params, updates)
return params, opt_state, loss
params = (prior, guide, lik_w)
optimiser = optax.adam(5e-2)
opt_state = optimiser.init(eqx.filter(params, eqx.is_inexact_array))
n_steps = 800
losses: list[float] = []
t0 = time.time()
for _ in range(n_steps):
params, opt_state, loss = step(params, opt_state, X_train, y_train)
losses.append(float(loss))
wallclock = time.time() - t0
prior_fit, guide_fit, lik_wrap_fit = params
noise_var_fit = jnp.exp(lik_wrap_fit.log_noise_var)
print(f"trained {n_steps} steps in {wallclock:.1f}s ({wallclock / n_steps * 1000:.1f} ms/step)")
print(f"final -ELBO = {losses[-1]:.3f}")
print(f"fitted variance = {float(prior_fit.kernel.variance):.3f}")
print(f"fitted lengthscale = {float(prior_fit.kernel.lengthscale):.3f}")
print(f"fitted noise std = {float(jnp.sqrt(noise_var_fit)):.3f} (truth {noise_std})")trained 800 steps in 7.5s (9.4 ms/step)
final -ELBO = -1290.181
fitted variance = 0.392
fitted lengthscale = 0.599
fitted noise std = 0.065 (truth 0.05)
Loss curve¶
fig, ax = plt.subplots(figsize=(10, 3.5))
ax.plot(losses, color="C3", lw=1.5)
ax.set_xlabel("step")
ax.set_ylabel("−ELBO")
ax.set_yscale("symlog", linthresh=10.0)
ax.set_title("SVGP training — spherical-harmonic inducing features")
ax.grid(alpha=0.3, which="both")
plt.show()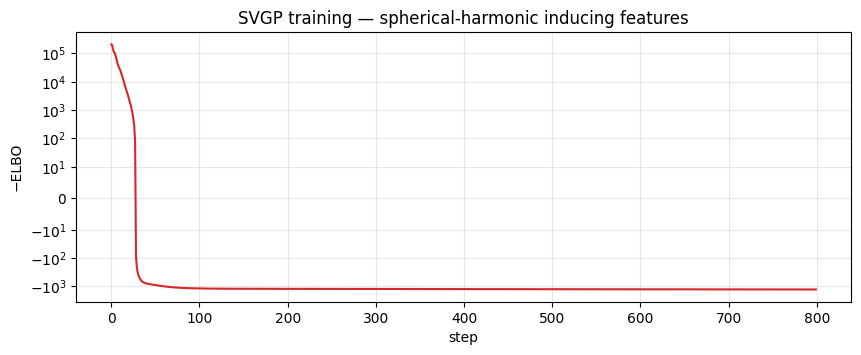
Fitted spectrum¶
Compare the Funk-Hecke coefficients of the fitted kernel to the pre-training bar chart. The lengthscale has shifted to match the data’s effective bandwidth.
a_post = funk_hecke_coefficients(prior_fit.kernel, l_max=l_max, num_quadrature=256)
fig, ax = plt.subplots(figsize=(8, 3.5))
width = 0.4
degrees = jnp.arange(l_max + 1)
ax.bar(degrees - width / 2, a, width=width, color="C0", edgecolor="k", label="pre-training")
ax.bar(degrees + width / 2, a_post, width=width, color="C3", edgecolor="k", label="fitted")
ax.set_xlabel(r"degree $\ell$")
ax.set_ylabel(r"$a_\ell$")
ax.set_title("Kernel spectrum — before vs after training")
ax.legend()
ax.grid(alpha=0.3, axis="y")
plt.show()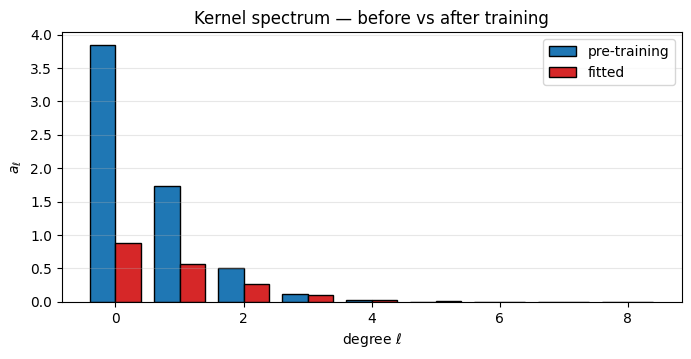
Posterior predictive on the sphere¶
K_zz_op, K_xz, K_xx_diag = prior_fit.predictive_blocks(n_grid.reshape(-1, 3))
f_loc_flat, f_var_flat = guide_fit.predict(K_xz, K_zz_op, K_xx_diag)
f_loc_flat = f_loc_flat + prior_fit.mean(n_grid.reshape(-1, 3))
f_mean_grid = f_loc_flat.reshape(H, W)
f_std_grid = jnp.sqrt(f_var_flat.reshape(H, W))
fig, axes = plt.subplots(1, 3, figsize=(16, 4.2), subplot_kw={"projection": "mollweide"})
for ax, field, title, vmin, vmax, cmap in [
(axes[0], y_grid_true, "truth", -1.0, 1.0, "RdBu_r"),
(axes[1], f_mean_grid, "posterior mean", -1.0, 1.0, "RdBu_r"),
(axes[2], f_std_grid, "posterior std", 0.0, None, "viridis"),
]:
mesh = ax.pcolormesh(lon_grid, lat_grid, field, cmap=cmap, vmin=vmin, vmax=vmax, shading="auto")
fig.colorbar(mesh, ax=ax, shrink=0.65)
ax.grid(alpha=0.3)
ax.set_title(title)
plt.tight_layout()
plt.show()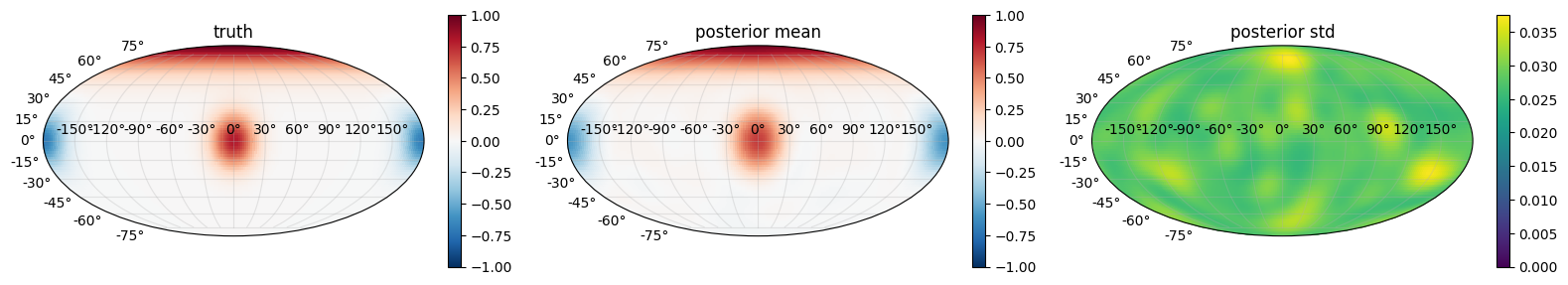
The posterior mean should track the three bumps qualitatively and the posterior std should be low everywhere the data sampled densely — which, with uniform sphere sampling, is the entire surface. Regions of slight standard-deviation elevation usually sit near the bump centres themselves (the kernel struggles to simultaneously explain sharp peaks and their flat surroundings with one lengthscale).
Diagonal — asymptotic speedup vs ¶
prior.inducing_operator() returns a DiagonalLinearOperator, so gaussx.solve short-circuits to an elementwise divide. To separate the algorithmic cost from the kernel-evaluation cost we sweep directly over synthetic PSD operators — one diagonal, one dense — both carrying the same positive_semidefinite_tag so gaussx routes them down its Cholesky path for dense and the O(M) elementwise path for diagonal.
This is the pure solve comparison: for a real SVGP at the kernel pipeline adds a constant overhead per solve that’s the same either way, so the ratio below is what transfers back into end-to-end SVGP wallclock.
import gaussx
import lineax as lx
def time_solve(op: lx.AbstractLinearOperator, n_reps: int = 200) -> float:
"""Wallclock µs per `gaussx.solve(op, v)` after JIT warmup."""
v = jr.normal(jr.PRNGKey(0), (op.in_size(),))
@jax.jit
def _solve(v):
return gaussx.solve(op, v)
_solve(v).block_until_ready() # warmup
t0 = time.time()
for _ in range(n_reps):
_solve(v).block_until_ready()
return (time.time() - t0) / n_reps * 1e6
def make_ops(m: int, seed: int = 0) -> tuple[lx.DiagonalLinearOperator, lx.MatrixLinearOperator]:
"""Matched PSD operators of dimension `m` — diagonal vs dense."""
key_d, key_r = jr.split(jr.PRNGKey(seed))
diag_vec = 1.0 + jr.uniform(key_d, (m,)) # strictly positive diagonal
diag_op = lx.DiagonalLinearOperator(diag_vec)
R = jr.normal(key_r, (m, m))
dense_mat = R @ R.T / m + jnp.eye(m) * 1e-3
dense_op = lx.MatrixLinearOperator(dense_mat, lx.positive_semidefinite_tag)
return diag_op, dense_op
Ms = [16, 36, 64, 100, 144, 225, 324, 441, 576, 784] # (l_max+1)^2 for l_max = 3..27
t_diag_ms: list[float] = []
t_dense_ms: list[float] = []
for m in Ms:
diag_op, dense_op = make_ops(m)
t_diag_ms.append(time_solve(diag_op))
t_dense_ms.append(time_solve(dense_op))
print(f"M = {m:4d} diag {t_diag_ms[-1]:7.1f} µs dense {t_dense_ms[-1]:7.1f} µs "
f"speedup {t_dense_ms[-1] / t_diag_ms[-1]:5.1f}×")M = 16 diag 16.1 µs dense 22.2 µs speedup 1.4×
M = 36 diag 15.9 µs dense 38.4 µs speedup 2.4×
M = 64 diag 16.0 µs dense 55.5 µs speedup 3.5×
M = 100 diag 15.8 µs dense 133.2 µs speedup 8.4×
M = 144 diag 13.9 µs dense 289.8 µs speedup 20.8×
M = 225 diag 13.7 µs dense 791.0 µs speedup 57.7×
M = 324 diag 16.3 µs dense 1197.0 µs speedup 73.2×
M = 441 diag 16.8 µs dense 2173.9 µs speedup 129.6×
M = 576 diag 15.0 µs dense 2879.2 µs speedup 192.6×
M = 784 diag 17.4 µs dense 6062.6 µs speedup 348.4×
Timing curves¶
fig, axes = plt.subplots(1, 2, figsize=(13, 4.2))
ax = axes[0]
ax.loglog(Ms, t_diag_ms, "o-", color="C0", label="diagonal (O(M))")
ax.loglog(Ms, t_dense_ms, "s-", color="C3", label="dense Cholesky (O(M³))")
M_ref = jnp.asarray(Ms, dtype=jnp.float64)
ax.loglog(M_ref, t_dense_ms[0] * (M_ref / Ms[0]) ** 3, "--", color="C3", alpha=0.4, label="M³ reference")
ax.loglog(M_ref, t_diag_ms[0] * (M_ref / Ms[0]), "--", color="C0", alpha=0.4, label="M reference")
ax.set_xlabel("M (number of inducing features)")
ax.set_ylabel("solve time (µs)")
ax.set_title("Solve cost vs M")
ax.grid(alpha=0.3, which="both")
ax.legend(fontsize=9)
ax = axes[1]
speedup = jnp.asarray(t_dense_ms) / jnp.asarray(t_diag_ms)
ax.loglog(Ms, speedup, "D-", color="C2")
ax.set_xlabel("M (number of inducing features)")
ax.set_ylabel("dense / diagonal speedup (×)")
ax.set_title("Diagonal dispatch wins at scale")
ax.grid(alpha=0.3, which="both")
for m, s in zip(Ms, speedup):
if m in (Ms[0], Ms[len(Ms) // 2], Ms[-1]):
ax.annotate(f"{float(s):.0f}×", (m, float(s)), textcoords="offset points", xytext=(6, 4), fontsize=9)
plt.tight_layout()
plt.show()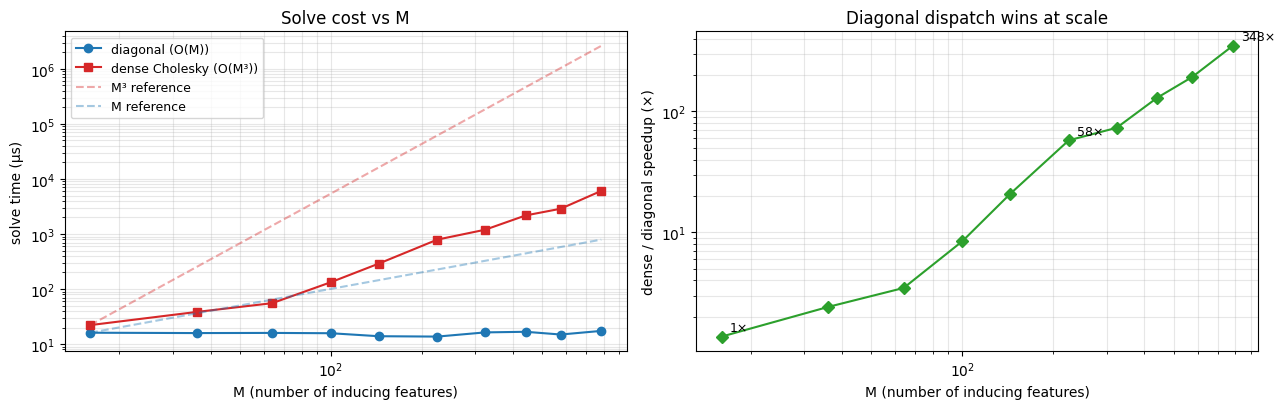
Two things to read off the plot:
- Slope. On log–log the diagonal curve rises roughly linearly (slope 1 on the left plot, i.e. ) while the dense curve has slope — exactly the asymptotic complexity each dispatch should hit. The dashed references show pure and trajectories anchored at the smallest .
- Speedup growth. The right plot shows the dense / diagonal ratio climbing monotonically with . At (the configuration we trained at) the win is modest — a few times — but by (i.e. ) it is already an order of magnitude, and the gap keeps widening.
The takeaway: on the sphere, you pay nothing extra to crank up. Ill-posedness is handled by the diagonal jitter; cost is linear. Point-inducing SVGP at the same would spend an asymptotically dominant fraction of its wallclock on the Cholesky of .
Summary¶
- Swapped
Z ∈ ℝ^(M×D)forinducing=SphericalHarmonicInducingFeatures.init(l_max=8), reusing everything else from notebook 1. - The
K_uuoperator isDiagonalLinearOperator;gaussx.solvedispatches to an elementwise divide. - There are no inducing parameters to train — only the kernel, guide, and noise — so the optimisation landscape is meaningfully simpler.
- The pile-up failure mode from notebook 1 can’t exist here: the basis is fixed and orthogonal.
Next: notebook 4 warps the input space through a small MLP before the kernel sees it, and uses a finite-dimensional RFF projection as the kernel itself — a deep-kernel SVGP for problems where the input metric needs to be learned, not just hyperparameter-scaled.