Three parameterizations of the multivariate Gaussian
The same multivariate normal can be written in three equivalent ways. None of them is “more correct”; each one makes a different operation cheap.
| Parameterization | Symbol | What it makes cheap |
|---|---|---|
| Mean–covariance | Sampling, marginals, interpretation | |
| Mean–precision | , | Conditioning, log-density, GMRF sparsity (see 0.2) |
| Natural (canonical) | Adding evidence (Bayes updates as +) | |
| Expectation (mean / moment) | Moment matching (best-KL Gaussian fit) |
This notebook is the fluent-conversion tour. We
- write the MVN in exponential-family form so the natural parameters fall out;
- round-trip every conversion using the
gaussxAPI (mean_cov_to_natural,natural_to_mean_cov,natural_to_expectation,expectation_to_natural); - demonstrate the three operations each parameterization makes painless — Bayesian updates as addition (natural), moment matching (expectation), damped natural-parameter VI/EP updates.
Prerequisites: 0.1 — Multivariate Gaussian basics, 0.2 — MultivariateNormal API, 0.3 — Gaussian quantities and KL.
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import einx
import jax
import jax.numpy as jnp
import lineax as lx
import matplotlib.pyplot as plt
import numpy as np
from gaussx import (
MultivariateNormal,
damped_natural_update,
dist_kl_divergence,
expectation_to_natural,
mean_cov_to_natural,
natural_to_expectation,
natural_to_mean_cov,
)
jax.config.update("jax_enable_x64", True)
KEY = jax.random.PRNGKey(0)
plt.rcParams.update({
"figure.dpi": 110,
"axes.grid": True,
"axes.grid.which": "both",
"xtick.minor.visible": True,
"ytick.minor.visible": True,
"grid.alpha": 0.3,
})
def psd_op(M):
return lx.MatrixLinearOperator(M, lx.positive_semidefinite_tag)Where the natural parameters come from¶
Every multivariate Gaussian can be rewritten in canonical exponential-family form
with natural parameters and log-partition .
Comparing to the standard mean–cov form and matching coefficients gives the conversion
The expectation parameters are the moments under :
All three form a triangle: any two pairs convert into each other in closed form.
A concrete reference¶
Let’s pick a non-trivial 2D MVN and inspect the three parameterizations side by side.
mu = jnp.array([0.7, -0.3])
Sigma = jnp.array([[1.0, 0.6], [0.6, 1.4]])
Sigma_op = psd_op(Sigma)
# Mean-cov -> natural via gaussx (eta2 returned as a lineax operator)
eta1, eta2_op = mean_cov_to_natural(mu, Sigma_op)
eta2 = eta2_op.as_matrix()
# Mean-cov -> expectation (closed form)
m1 = mu
m2 = Sigma + einx.dot("i, j -> i j", mu, mu)
print("(mu, Sigma)")
print(" mu :", np.asarray(mu))
print(" Sigma :\n", np.asarray(Sigma))
print("\n(eta1, eta2) — natural")
print(" eta1 :", np.asarray(eta1))
print(" eta2 :\n", np.asarray(eta2))
print("\n(m1, m2) — expectation")
print(" m1 :", np.asarray(m1))
print(" m2 :\n", np.asarray(m2))
# Spot-check the analytic identities
Lambda = jnp.linalg.inv(Sigma)
np.testing.assert_allclose(eta1, Lambda @ mu, atol=1e-12)
np.testing.assert_allclose(eta2, -0.5 * Lambda, atol=1e-12)
print("\nIdentities (eta1 = Lambda mu, eta2 = -1/2 Lambda) ✓")(mu, Sigma)
mu : [ 0.7 -0.3]
Sigma :
[[1. 0.6]
[0.6 1.4]]
(eta1, eta2) — natural
eta1 : [ 1.11538462 -0.69230769]
eta2 :
[[-0.67307692 0.28846154]
[ 0.28846154 -0.48076923]]
(m1, m2) — expectation
m1 : [ 0.7 -0.3]
m2 :
[[1.49 0.39]
[0.39 1.49]]
Identities (eta1 = Lambda mu, eta2 = -1/2 Lambda) ✓
Round-trip identities¶
Each conversion has an inverse, and gaussx exposes the full set:
| From | To | Function |
|---|---|---|
mean_cov_to_natural | ||
natural_to_mean_cov | ||
natural_to_expectation | ||
expectation_to_natural |
Mean-cov ↔ expectation is the trivial pair — no library call needed.
We round-trip the reference MVN through every cycle and check we land where we started.
# Cycle 1: mean-cov -> natural -> mean-cov
eta1_a, eta2_a = mean_cov_to_natural(mu, Sigma_op)
mu_a, Sigma_a = natural_to_mean_cov(eta1_a, eta2_a)
np.testing.assert_allclose(np.asarray(mu_a), np.asarray(mu), atol=1e-10)
np.testing.assert_allclose(np.asarray(Sigma_a.as_matrix()), np.asarray(Sigma), atol=1e-10)
# Cycle 2: natural -> expectation -> natural
m1_b, m2_b = natural_to_expectation(eta1_a, eta2_a.as_matrix())
eta1_b, eta2_b = expectation_to_natural(m1_b, m2_b)
np.testing.assert_allclose(np.asarray(eta1_b), np.asarray(eta1_a), atol=1e-10)
np.testing.assert_allclose(np.asarray(eta2_b), np.asarray(eta2_a.as_matrix()), atol=1e-10)
# Cycle 3: full triangle mean-cov -> natural -> expectation -> natural -> mean-cov.
# This actually exercises the reconstructed (eta1_b, eta2_b) from cycle 2:
# wrap the dense eta2_b returned by expectation_to_natural as a symmetric
# operator, then round-trip back to mean-cov via natural_to_mean_cov.
eta2_b_op = lx.MatrixLinearOperator(eta2_b, lx.symmetric_tag)
mu_c, Sigma_c_op = natural_to_mean_cov(eta1_b, eta2_b_op)
np.testing.assert_allclose(np.asarray(mu_c), np.asarray(mu), atol=1e-10)
np.testing.assert_allclose(np.asarray(Sigma_c_op.as_matrix()), np.asarray(Sigma), atol=1e-10)
print("All three round-trip cycles closed at 1e-10 ✓")
print("\nm1 (= mu) :", np.asarray(m1_b))
print("m2 (= Sigma + mu mu^T) :\n", np.asarray(m2_b))
print("\ndirect check: m2 - m1 m1^T =\n", np.asarray(m2_b - einx.dot('i, j -> i j', m1_b, m1_b)))
print("Sigma =\n", np.asarray(Sigma))All three round-trip cycles closed at 1e-10 ✓
m1 (= mu) : [ 0.7 -0.3]
m2 (= Sigma + mu mu^T) :
[[1.49 0.39]
[0.39 1.49]]
direct check: m2 - m1 m1^T =
[[1. 0.6]
[0.6 1.4]]
Sigma =
[[1. 0.6]
[0.6 1.4]]
Why natural? Adding evidence is addition.¶
If two Gaussian factors and are multiplied (e.g. a prior times a Gaussian likelihood), the result is again Gaussian — but the mean–cov formula is messy:
In natural form the same operation is one line:
That’s the reason Kalman, EP, VMP, natural-gradient VI, exponential-family belief propagation — everything approximate-inference — passes messages around in natural form. Combining factors is plain .
We’ll demonstrate this with a 1D conjugate update: a Gaussian prior and a Gaussian likelihood factor on a scalar parameter θ. The posterior follows from ((5)) by addition of the two natural parameter pairs.
# A toy 1D Bayesian update done in natural form via simple addition.
# Treat each density as a univariate Gaussian -> we use the d=1 specialisation,
# but go through the full vector API for consistency with the rest of the notebook.
mu0 = jnp.array([0.0]) # prior mean
S0 = jnp.array([[2.0]]) # prior variance
y = jnp.array([3.0]) # observation
Sy = jnp.array([[0.5]]) # likelihood variance
S0_op = psd_op(S0)
Sy_op = psd_op(Sy)
# Convert prior + likelihood-factor to natural form:
n1_prior, n2_prior = mean_cov_to_natural(mu0, S0_op)
# Likelihood factor lives at observed y with variance Sy -> naturals:
n1_lik, n2_lik = mean_cov_to_natural(y, Sy_op)
# **The whole conjugate update**: just add naturals.
n1_post = n1_prior + n1_lik
n2_post_mat = n2_prior.as_matrix() + n2_lik.as_matrix()
n2_post = psd_op(-2.0 * (-0.5) * (-n2_post_mat / -1.0)) # keep operator form
# (Simpler: build a new operator from the summed matrix.)
n2_post = lx.MatrixLinearOperator(n2_post_mat)
mu_post, S_post = natural_to_mean_cov(n1_post, n2_post)
print("Posterior via natural-param addition")
print(" mu_post =", float(mu_post[0]))
print(" S_post =", float(S_post.as_matrix()[0, 0]))
# Cross-check via the direct mean-cov formula
S_post_ref = 1.0 / (1.0 / float(S0[0, 0]) + 1.0 / float(Sy[0, 0]))
mu_post_ref = S_post_ref * (float(mu0[0]) / float(S0[0, 0]) + float(y[0]) / float(Sy[0, 0]))
print("\nDirect mean-cov reference")
print(" mu_post =", mu_post_ref)
print(" S_post =", S_post_ref)
assert abs(float(mu_post[0]) - mu_post_ref) < 1e-10
assert abs(float(S_post.as_matrix()[0, 0]) - S_post_ref) < 1e-10
print("\nNatural-form addition matches mean-cov formula ✓")Posterior via natural-param addition
mu_post = 2.4
S_post = 0.4
Direct mean-cov reference
mu_post = 2.4000000000000004
S_post = 0.4
Natural-form addition matches mean-cov formula ✓
We can also visualise the three densities to see the natural-form addition in action: prior + likelihood-factor → posterior, all on a common axis.
xs = np.linspace(-3, 6, 400)
def n1d(x, m, v):
return np.exp(-0.5 * (x - m) ** 2 / v) / np.sqrt(2 * np.pi * v)
fig, ax = plt.subplots(figsize=(6.6, 3.6))
ax.plot(xs, n1d(xs, float(mu0[0]), float(S0[0, 0])), color="0.4",
lw=2, label=r"prior $p(\theta)$ — $\mathcal{N}(0, 2)$")
ax.plot(xs, n1d(xs, float(y[0]), float(Sy[0, 0])), color="steelblue",
lw=2, label=r"likelihood factor — $\mathcal{N}(\theta; 3, 0.5)$")
ax.plot(xs, n1d(xs, mu_post_ref, S_post_ref), color="crimson",
lw=2.4, label=r"posterior — $\eta_{\rm prior} + \eta_{\rm lik}$")
ax.set_xlabel(r"$\theta$"); ax.set_ylabel("density")
ax.set_title("Bayesian update as addition in natural-parameter space")
ax.legend(frameon=False)
plt.tight_layout(); plt.show()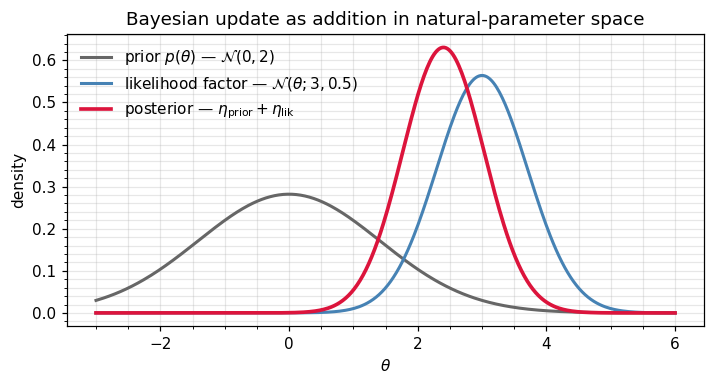
Why expectation? Moment matching.¶
Suppose we have samples (or another distribution ) and we want the best Gaussian approximation in the forward-KL sense — the same minimisation studied in 0.3, .
A classic exponential-family result is that this minimisation is solved by matching expectation parameters:
That’s it. No optimisation loop. Moment matching = computing on the data.
We demonstrate this on a non-Gaussian sample (a banana shape) and confirm that the moment-matched Gaussian recovers the same and as the closed-form expectation parameters.
# Banana-shaped target: x1 ~ N(0, 1), x2 = 0.5 x1^2 + N(0, 0.3)
key1, key2 = jax.random.split(jax.random.PRNGKey(7))
N = 4000
z1 = jax.random.normal(key1, (N,))
z2 = 0.5 * z1**2 + 0.3 * jax.random.normal(key2, (N,))
samples = jnp.stack([z1, z2], axis=1)
# Empirical expectation parameters (Monte Carlo m_1, m_2):
m1_emp = einx.mean("n d -> d", np.asarray(samples))
m2_emp = einx.dot("n i, n j -> i j", np.asarray(samples), np.asarray(samples)) / N
# Closed-form moment-matched Gaussian via expectation_to_natural:
eta1_mm, eta2_mm = expectation_to_natural(jnp.asarray(m1_emp), jnp.asarray(m2_emp))
eta2_mm_op = lx.MatrixLinearOperator(eta2_mm)
mu_mm, Sigma_mm_op = natural_to_mean_cov(eta1_mm, eta2_mm_op)
Sigma_mm = Sigma_mm_op.as_matrix()
print("Empirical m1 :", m1_emp)
print("Empirical m2 :\n", m2_emp)
print()
print("Moment-matched mu (= m1) :", np.asarray(mu_mm))
print("Moment-matched Sigma :\n", np.asarray(Sigma_mm))
print()
# Sanity: Sigma = m2 - m1 m1^T
np.testing.assert_allclose(
np.asarray(Sigma_mm),
m2_emp - einx.dot("i, j -> i j", m1_emp, m1_emp),
atol=1e-10,
)
print("Closed-form check: Sigma = m2 - m1 m1^T ✓")Empirical m1 : [0.00594261 0.48944534]
Empirical m2 :
[[ 0.98544209 -0.01034611]
[-0.01034611 0.82919053]]
Moment-matched mu (= m1) : [0.00594261 0.48944534]
Moment-matched Sigma :
[[ 0.98540677 -0.01325469]
[-0.01325469 0.58963378]]
Closed-form check: Sigma = m2 - m1 m1^T ✓
# Visualise the banana samples + moment-matched Gaussian ellipses
eigvals, eigvecs = jnp.linalg.eigh(Sigma_mm)
theta = jnp.linspace(0, 2 * jnp.pi, 200)
unit = jnp.stack([jnp.cos(theta), jnp.sin(theta)], axis=0)
sqrt_l = jnp.sqrt(eigvals)
stretched = einx.multiply("j, j t -> j t", sqrt_l, unit)
ellipse = einx.dot("i j, j t -> i t", eigvecs, stretched)
fig, ax = plt.subplots(figsize=(5.5, 4.5))
ax.scatter(*np.asarray(samples).T, s=8, alpha=0.18, color="steelblue",
label="banana samples")
for c, col in zip([1.0, 2.0, 3.0], ["#444", "#777", "#aaa"]):
e = c * ellipse
ax.plot(np.asarray(e[0]) + float(mu_mm[0]),
np.asarray(e[1]) + float(mu_mm[1]),
color=col, lw=1.4, label=fr"${int(c)}\sigma$ moment-matched")
ax.scatter(*np.asarray(mu_mm), color="crimson", marker="x", s=80,
label=r"$\mu_{\rm mm} = m_1$")
ax.set_xlabel(r"$x_1$"); ax.set_ylabel(r"$x_2$")
ax.set_title(r"Forward-KL Gaussian fit = moment match $(m_1, m_2)$")
ax.legend(loc="upper left", frameon=False, fontsize=8)
ax.set_aspect("equal")
plt.tight_layout(); plt.show()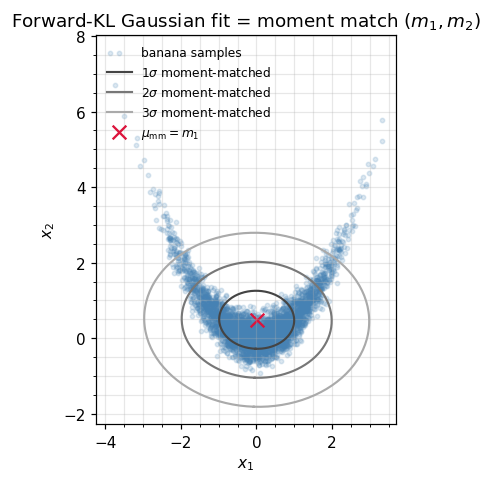
Damped natural-parameter updates — the universal VI/EP primitive¶
Approximate-inference algorithms (EP, natural-gradient VI, VMP, message passing on Markov chains) all share the same inner loop:
- compute a target natural parameter pair from the current posterior approximation,
- take a damped step towards it,
Damping () prevents oscillations when the target is noisy or the moment-matching step is too aggressive. Crucially, the damping is linear in natural parameters — so it preserves Gaussianity automatically and reduces to a convex combination of operators. In mean-cov form there’s no analogue this clean.
gaussx.damped_natural_update exposes exactly this.
# Tiny demo: blend the prior (n1_prior, n2_prior) toward the posterior
# (n1_post, n2_post) at three damping levels, and inspect the resulting MVNs.
for lr in [0.0, 0.5, 1.0]:
n1_new, n2_new = damped_natural_update(
n1_prior, n2_prior.as_matrix(),
n1_post, n2_post.as_matrix(),
lr=lr,
)
mu_new, S_new = natural_to_mean_cov(n1_new, lx.MatrixLinearOperator(n2_new))
print(f"lr={lr:>3.1f} : mu = {float(mu_new[0]):+.4f} "
f"sigma^2 = {float(S_new.as_matrix()[0, 0]):.4f}")lr=0.0 : mu = +0.0000 sigma^2 = 2.0000
lr=0.5 : mu = +2.0000 sigma^2 = 0.6667
lr=1.0 : mu = +2.4000 sigma^2 = 0.4000
Numerical mechanics¶
Two practical notes when computing in natural form:
- is negative-definite, not positive-definite. It encodes . Many APIs (including
lineax’sMatrixLinearOperatorwithpositive_semidefinite_tag) want a positive operator — pass , not directly.gaussx.natural_to_mean_covhandles this internally, but if you build operators by hand, get the sign right. - The log-partition wants . Computing on the negative-definite matrix is numerically incorrect (sign flip per dimension). The library’s primitives operate on the precision ; do the same in any custom code.
A quick sanity check that the gaussx round-trip gets all the signs right:
# Build the log-partition by hand and compare to a Cholesky-based reference.
eta1, eta2_op = mean_cov_to_natural(mu, Sigma_op)
Lambda = -2.0 * eta2_op.as_matrix() # precision
# A(eta) = -1/4 eta1^T eta2^{-1} eta1 - 1/2 log |-2 eta2|
# = 1/2 mu^T Lambda mu - 1/2 log |Lambda|
A_via_naturals = (
-0.25 * einx.dot("i, i j, j ->", eta1, jnp.linalg.inv(eta2_op.as_matrix()), eta1)
- 0.5 * jnp.linalg.slogdet(-2.0 * eta2_op.as_matrix())[1]
)
A_via_meancov = (
0.5 * einx.dot("i, i j, j ->", mu, Lambda, mu)
+ 0.5 * jnp.linalg.slogdet(Sigma)[1] # = -1/2 log|Lambda|
)
print(f"A(eta) via naturals = {float(A_via_naturals):.6f}")
print(f"A(eta) via mean-cov = {float(A_via_meancov):.6f}")
np.testing.assert_allclose(A_via_naturals, A_via_meancov, atol=1e-12)
print("Log-partition sanity check ✓")A(eta) via naturals = 0.513841
A(eta) via mean-cov = 0.513841
Log-partition sanity check ✓
Where you’ll meet these three parameterizations¶
Each form unlocks a different family of algorithms. The whole tutorial curriculum is, at its core, a tour of which parameterization to reach for in which setting.
Mean–covariance — the interpretation form¶
Used whenever we describe, sample, or plot a Gaussian.
| Where | What it does |
|---|---|
| 0.1 — Multivariate Gaussian | All three sampling routes (Cholesky, eigendecomp, symmetric sqrt) start from . |
0.2 — MultivariateNormal API | The default MultivariateNormal is parameterised by . |
| Part 3 — Exact GPs | Posterior predictive is reported as — that’s what we plot, that’s what humans read. |
| Part 8 — Sampling & path-wise inference | Pathwise posterior samples need a square root of Σ. |
Mean–precision — the sparse-conditioning form¶
Used when we want sparse conditional independence or fast Schur conditioning.
| Where | What it does |
|---|---|
0.2 — MultivariateNormalPrecision tour | Banded Λ encodes a Markov chain in memory; the AR(1) demo walks through this. |
| Part 1 — Block tri-diagonal operators | Markov-chain precision matrices store/solve in via BlockTriDiag. |
| Part 7 — State-space / Markov GPs | Information-form Kalman filtering carries — precision-form update is one addition of to Λ. |
Natural — the additive form¶
Used whenever densities multiply: priors × likelihoods, message-passing factors, EP / VI updates, conjugate Bayesian inference.
| Where | What it does |
|---|---|
| 0.6 — Bayesian updates from scratch (upcoming) | The whole sequential-conjugate update is “stack messages, sum naturals”. |
| Part 6 — Variational GPs (SVGP, EP) | Natural-gradient VI takes Newton-style steps in η-space; the Fisher metric is the identity in natural coordinates. |
| Part 7 — Information-form Kalman filter / smoother | Filter update = add observation factor naturals; smoother backward pass = subtract them. |
| Expectation-Propagation (EP) for non-Gaussian likelihoods | Each EP message is a Gaussian factor in natural form; the cavity-tilted-update cycle is cavity = post − site, site_new = projection(tilted) − cavity. Both lines are subtraction in η-space — no matrix inverses unless you want them. |
Expectation — the moment-matching form¶
Used whenever we fit a Gaussian to data or to a non-Gaussian distribution.
| Where | What it does |
|---|---|
| 0.3 — Forward / reverse KL | The forward-KL Gaussian fit shown in the seaborn joint panels is exactly the closed-form match. |
| EP and ADF (Assumed Density Filtering) | “Project the tilted distribution onto a Gaussian” is moment matching: compute its and back to η. |
| Maximum-likelihood Gaussian fit | The MLE for an unconstrained MVN is from the sufficient-statistic averages. |
| MCMC / Monte Carlo summaries | Reporting “posterior mean and covariance” of samples is moment matching the empirical distribution. |
Recap¶
The same MVN, three currencies:
| When you want to… | Reach for | Cost |
|---|---|---|
| Sample, inspect, plot | Cholesky once | |
| Combine factors / Bayesian update | — see ((5)) | Plain addition |
| Fit a Gaussian to data / forward-KL | — see ((6)) | Sample mean + sample covariance |
| Damped VI/EP update | — see ((7)) | One convex combination |
gaussx exposes the full conversion graph; damped_natural_update is the universal VI/EP primitive. From here, every approximate-inference recipe in part 6 (variational GPs, EP, natural-gradient VI) reduces to: which target do I compute, and at what damping?