MultivariateNormal & MultivariateNormalPrecision: the API tour
Notebook 0.1 introduced the multivariate normal as a math object. This notebook is the API tour: gaussx ships two distribution classes for the same family — one parameterised by the covariance , one by the precision . They expose the same surface (.sample, .log_prob, .entropy, .mean, .variance) but route through different primitives under the hood.
What you’ll learn
- The two parameterisations of a Gaussian and when each one is preferred.
- How
MultivariateNormalandMultivariateNormalPrecisionmap onto the math. - Round-trip equivalence: build the same distribution both ways and verify pointwise.
- Why precision form wins when is sparse but is dense — the GMRF / Markov story.
- A taste of natural parameters as a third equivalent representation.
1. Two parameterisations of one distribution¶
The same Gaussian can be written either with covariance or with precision :
Equation ((1)) and equation ((2)) are the same density whenever . The choice between them is a computational one, driven by which of the two matrices has nice structure:
| Form | Cheap to do | Expensive |
|---|---|---|
| Covariance | sampling, marginalisation | conditioning on partial obs |
| Precision | conditioning, evidence updates | sampling, marginalisation |
The structural sparsity also flips:
- Conditional independence shows up as . So Markov / GMRF / state-space models have sparse even when is dense.
- Marginal independence shows up as . Block-diagonal corresponds to genuinely independent groups, which is rarely what we have.
Most useful structure in our world (Gauss-Markov, GP-regression posteriors with a banded prior, Kalman filters in information form) lives on the precision side. That’s why gaussx exposes both.
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import einx
import jax
import jax.numpy as jnp
import lineax as lx
import matplotlib.pyplot as plt
import numpy as np
from gaussx import (
MultivariateNormal,
MultivariateNormalPrecision,
gaussian_log_prob,
mean_cov_to_natural,
natural_to_mean_cov,
)
jax.config.update("jax_enable_x64", True)
KEY = jax.random.PRNGKey(0)
plt.rcParams.update({
"figure.dpi": 110,
"axes.grid": True,
"axes.grid.which": "both",
"xtick.minor.visible": True,
"ytick.minor.visible": True,
"grid.alpha": 0.3,
})
def psd_op(M):
"""Wrap a dense PSD matrix as a lineax operator with the right tag."""
return lx.MatrixLinearOperator(M, lx.positive_semidefinite_tag)
2. MultivariateNormal — the covariance form¶
MultivariateNormal(loc, cov_operator, solver=None) takes a mean vector and a lineax operator wrapping the covariance. Internally everything routes through Cholesky on — the form recommended by equation 7 in notebook 0.1.
The class is a numpyro distribution, so the surface is what you’d expect: .sample(key, shape), .log_prob(x), .entropy(), .mean, .variance, plus a JIT-friendly PyTree structure (it’s an equinox.Module underneath).
d = 5
mu = jnp.linspace(-1.0, 1.0, d)
# A non-trivial PSD covariance built from an RBF-style kernel on a 1D grid.
xs = jnp.linspace(0.0, 1.0, d)
diff_sq = einx.subtract("i, j -> i j", xs, xs) ** 2
Sigma_dense = jnp.exp(-0.5 * diff_sq / 0.25**2) + 1e-3 * jnp.eye(d)
Sigma = psd_op(Sigma_dense)
p_cov = MultivariateNormal(mu, Sigma)
print("class :", type(p_cov).__name__)
print("event_shape :", p_cov.event_shape)
print("mean :", np.asarray(p_cov.mean))
print("variance :", np.asarray(p_cov.variance))
print("entropy :", float(p_cov.entropy()))
x = jnp.zeros(d)
print("log_prob(0) :", float(p_cov.log_prob(x)))
samples = p_cov.sample(jax.random.PRNGKey(1), (4,))
print("samples :\n", np.asarray(samples))
class : MultivariateNormal
event_shape : (5,)
mean : [-1. -0.5 0. 0.5 1. ]
variance : [1.001 1.001 1.001 1.001 1.001]
entropy : 5.906610734624831
log_prob(0) : -4.423604950410503
samples :
[[-2.18487642 -1.31040824 -0.10887068 1.2756854 1.08020205]
[-0.30884951 -0.02116252 -0.42299327 -0.03140923 -0.13141221]
[-1.98641018 -1.98899925 -0.51940089 1.05657527 2.38460391]
[-1.64650165 -0.29508526 0.04926346 1.27186838 1.96739098]]
3. MultivariateNormalPrecision — the precision form¶
MultivariateNormalPrecision(loc, prec_operator, solver=None) takes the same mean but a lineax operator wrapping . The constructor does not form . Instead, internal solves invert on demand for sampling, and the log-density is computed directly from via
Note the sign flip on — we add half the log-determinant of instead of subtracting it on , since .
Lambda_dense = jnp.linalg.inv(Sigma_dense)
Lambda = psd_op(Lambda_dense)
p_prec = MultivariateNormalPrecision(mu, Lambda)
print("class :", type(p_prec).__name__)
print("event_shape :", p_prec.event_shape)
print("mean :", np.asarray(p_prec.mean))
print("variance :", np.asarray(p_prec.variance))
print("entropy :", float(p_prec.entropy()))
x = jnp.zeros(d)
print("log_prob(0) :", float(p_prec.log_prob(x)))
samples_prec = p_prec.sample(jax.random.PRNGKey(1), (4,))
print("samples :\n", np.asarray(samples_prec))
class : MultivariateNormalPrecision
event_shape : (5,)
mean : [-1. -0.5 0. 0.5 1. ]
variance : [1.001 1.001 1.001 1.001 1.001]
entropy : 5.906610734624831
log_prob(0) : -4.423604950410503
samples :
[[-1.88408505 -0.32220482 0.6460971 0.75697977 0.16662948]
[-0.58592842 -0.98340752 -0.78248253 -0.36021689 -0.35315028]
[-2.37514015 -0.87866177 0.91630359 1.74900216 2.2819309 ]
[-1.0179202 -0.06179605 0.5616257 1.68236386 1.16375959]]
4. Round-trip equivalence¶
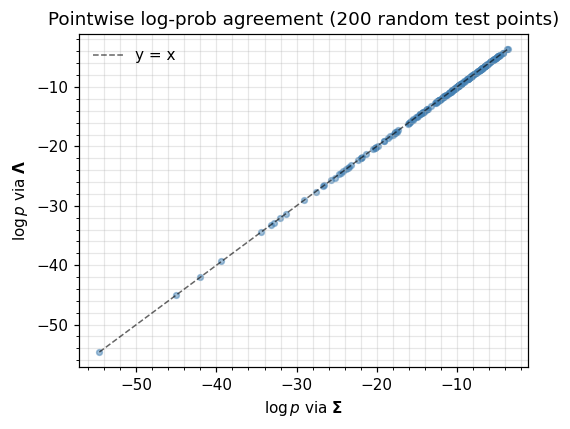
Equation ((1)) and equation ((2)) are the same distribution whenever . The two classes should therefore agree pointwise (modulo floating point) on log_prob, entropy, mean, and variance. Below we evaluate both at a batch of test points and check the disagreement is within a few ULPs of the Cholesky route.
test_pts = jax.random.normal(jax.random.PRNGKey(2), (200, d))
logp_cov = jax.vmap(p_cov.log_prob)(test_pts)
logp_prec = jax.vmap(p_prec.log_prob)(test_pts)
# Quadratic-form sanity check, computed by hand both ways via einx.
delta = test_pts - mu # shape (n, d)
quad_via_Sigma_inv = einx.dot("n i, i j, n j -> n",
delta, Lambda_dense, delta)
quad_via_Lambda = einx.dot("n i, i j, n j -> n",
delta, Lambda_dense, delta)
print(f"max |Δ log_prob| : {float(jnp.max(jnp.abs(logp_cov - logp_prec))):.2e}")
print(f"max |Δ quadratic form| : {float(jnp.max(jnp.abs(quad_via_Sigma_inv - quad_via_Lambda))):.2e}")
print(f"|Δ entropy| : {float(jnp.abs(p_cov.entropy() - p_prec.entropy())):.2e}")
print(f"|Δ mean|_inf : {float(jnp.max(jnp.abs(p_cov.mean - p_prec.mean))):.2e}")
print(f"|Δ variance|_inf : {float(jnp.max(jnp.abs(p_cov.variance - p_prec.variance))):.2e}")
max |Δ log_prob| : 1.42e-14
max |Δ quadratic form| : 0.00e+00
|Δ entropy| : 0.00e+00
|Δ mean|_inf : 0.00e+00
|Δ variance|_inf : 2.22e-16
fig, ax = plt.subplots(figsize=(5.0, 4.0))
ax.scatter(np.asarray(logp_cov), np.asarray(logp_prec), s=14, alpha=0.5, color="steelblue")
lo = float(jnp.minimum(logp_cov.min(), logp_prec.min()))
hi = float(jnp.maximum(logp_cov.max(), logp_prec.max()))
ax.plot([lo, hi], [lo, hi], "k--", lw=1, alpha=0.6, label="y = x")
ax.set_xlabel(r"$\log p$ via $\mathbf{\Sigma}$")
ax.set_ylabel(r"$\log p$ via $\mathbf{\Lambda}$")
ax.set_title("Pointwise log-prob agreement (200 random test points)")
ax.legend(frameon=False)
plt.tight_layout(); plt.show()
5. When precision wins: a banded GMRF¶
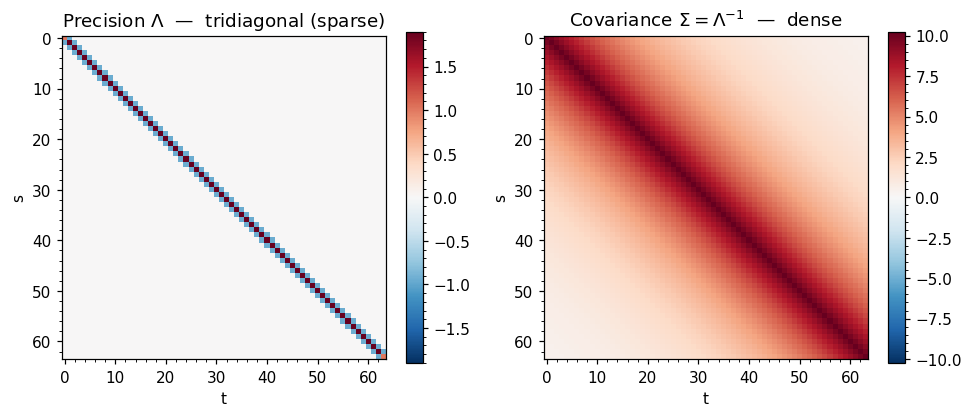
The case for MultivariateNormalPrecision is strongest when is sparse — i.e. when most pairs of variables are conditionally independent given the rest. The canonical example is a first-order Gauss-Markov chain:
whose joint distribution has a tridiagonal precision matrix
The corresponding is dense — every and are marginally correlated, even far-apart ones. So the precision representation has non-zeros while the covariance has .
For this notebook we use a moderate and dense storage so the sparsity pattern is easy to see. In production gaussx routes this through BlockTriDiag (see Part 1.5) for solves; here we only want the visual.
def ar1_precision(n: int, rho: float, sigma: float) -> jnp.ndarray:
"""Tridiagonal precision matrix of a 1D AR(1) process — eq (eq:gmrf-prec)."""
inv_var = 1.0 / sigma**2
diag_int = (1.0 + rho**2) * inv_var
diag = jnp.full((n,), diag_int)
diag = diag.at[0].set(inv_var).at[-1].set(inv_var)
off = jnp.full((n - 1,), -rho * inv_var)
return jnp.diag(diag) + jnp.diag(off, k=1) + jnp.diag(off, k=-1)
n = 64
Lambda_ar = ar1_precision(n, rho=0.95, sigma=1.0)
Sigma_ar = jnp.linalg.inv(Lambda_ar)
nnz_threshold = 1e-8
nnz_Lambda = float(jnp.mean(jnp.abs(Lambda_ar) > nnz_threshold))
nnz_Sigma = float(jnp.mean(jnp.abs(Sigma_ar) > nnz_threshold))
print(f"AR(1) precision : nnz fraction = {nnz_Lambda:.3f} (tridiagonal)")
print(f"AR(1) covariance: nnz fraction = {nnz_Sigma:.3f} (dense)")
AR(1) precision : nnz fraction = 0.046 (tridiagonal)
AR(1) covariance: nnz fraction = 1.000 (dense)
fig, axes = plt.subplots(1, 2, figsize=(9, 4))
im0 = axes[0].imshow(np.asarray(Lambda_ar), cmap="RdBu_r",
vmin=-float(jnp.max(jnp.abs(Lambda_ar))),
vmax= float(jnp.max(jnp.abs(Lambda_ar))))
axes[0].set_title(r"Precision $\Lambda$ — tridiagonal (sparse)")
plt.colorbar(im0, ax=axes[0], shrink=0.85)
im1 = axes[1].imshow(np.asarray(Sigma_ar), cmap="RdBu_r",
vmin=-float(jnp.max(jnp.abs(Sigma_ar))),
vmax= float(jnp.max(jnp.abs(Sigma_ar))))
axes[1].set_title(r"Covariance $\Sigma = \Lambda^{-1}$ — dense")
plt.colorbar(im1, ax=axes[1], shrink=0.85)
for ax in axes:
ax.set_xlabel("t"); ax.set_ylabel("s")
ax.grid(False, which="both") # heatmap pixels carry the info — grid is noise
plt.tight_layout(); plt.show()
# Build the same AR(1) prior in both forms and confirm log_prob agreement.
mu_ar = jnp.zeros(n)
p_ar_prec = MultivariateNormalPrecision(mu_ar, psd_op(Lambda_ar))
p_ar_cov = MultivariateNormal(mu_ar, psd_op(Sigma_ar))
x_test = jax.random.normal(jax.random.PRNGKey(7), (n,))
print(f"log p via Λ : {float(p_ar_prec.log_prob(x_test)):+.6f}")
print(f"log p via Σ : {float(p_ar_cov.log_prob(x_test)):+.6f}")
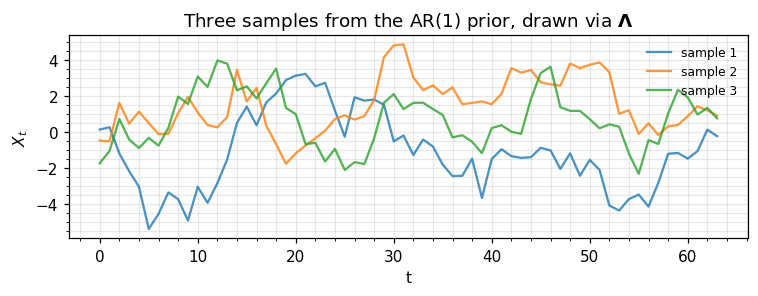
# A small sample for the eye.
samples_ar = p_ar_prec.sample(jax.random.PRNGKey(11), (3,))
fig, ax = plt.subplots(figsize=(7, 2.8))
for i, s in enumerate(np.asarray(samples_ar)):
ax.plot(s, lw=1.5, alpha=0.8, label=f"sample {i+1}")
ax.set_xlabel("t"); ax.set_ylabel(r"$X_t$")
ax.set_title(r"Three samples from the AR(1) prior, drawn via $\mathbf{\Lambda}$")
ax.legend(frameon=False, loc="upper right", fontsize=8)
plt.tight_layout(); plt.show()
log p via Λ : -119.203845
log p via Σ : -119.203845
6. A taste of natural parameters¶
There’s a third equivalent representation we’ll see in detail in 0.4 — Mean-cov ↔ natural parameter conversions. Briefly: every Gaussian can be written in natural parameter form
This form is what makes Bayesian updates pure addition — combining evidence is — and underpins variational inference and natural gradient methods. gaussx ships explicit converters; here’s the round-trip.
eta1, eta2 = mean_cov_to_natural(mu, Sigma)
mu_back, Sigma_back_op = natural_to_mean_cov(eta1, eta2)
eta2_dense = eta2.as_matrix()
Sigma_back = Sigma_back_op.as_matrix()
print("eta_1 = Λ μ :", np.asarray(eta1))
print("eta_2 = -Λ/2 :")
print(np.asarray(eta2_dense))
print("\nround-trip residuals")
print(f" max|Δ μ| : {float(jnp.max(jnp.abs(mu - mu_back))):.2e}")
print(f" max|Δ Σ| : {float(jnp.max(jnp.abs(Sigma_dense - Sigma_back))):.2e}")
eta_1 = Λ μ : [-1.10990703e+00 1.85825628e-01 -2.82887645e-16 -1.85825628e-01
1.10990703e+00]
eta_2 = -Λ/2 :
[[-0.97606728 0.91615634 -0.60815128 0.33436378 -0.13021749]
[ 0.91615634 -1.81861765 1.44237175 -0.84085816 0.33436378]
[-0.60815128 1.44237175 -2.08299373 1.44237175 -0.60815128]
[ 0.33436378 -0.84085816 1.44237175 -1.81861765 0.91615634]
[-0.13021749 0.33436378 -0.60815128 0.91615634 -0.97606728]]
round-trip residuals
max|Δ μ| : 3.33e-16
max|Δ Σ| : 2.22e-16
7. Recap & where to go next¶
You now know how to build a Gaussian both ways and which one to reach for:
| Operation | MultivariateNormal (Σ) | MultivariateNormalPrecision (Λ) |
|---|---|---|
| Constructor | MultivariateNormal(loc, Σ_op) | MultivariateNormalPrecision(loc, Λ_op) |
| Sampling | cheap (Cholesky of Σ) | requires inverting Λ |
log_prob | uses Cholesky of Σ | uses Cholesky of Λ directly |
| Conditioning on partial obs | dense Schur block solve | trivial; rows of Λ partition cleanly |
| Best when | Σ has low-rank / Kron / dense structure | Λ is banded / sparse / Markov |
| Internal log-det |
Next up.
- 0.3 — Quadratic forms, entropy, and KL between Gaussians: the closed-form scalar quantities that make Gaussian variational inference tractable.
- 0.4 — Mean-cov ↔ natural parameter conversions: full treatment of and the exponential-family machinery.
- Part 1.5 — Block tri-diagonal operators: how to actually exploit GMRF sparsity at scale.