Bayesian updates from scratch — sequential conjugate inference
Notebook 0.4 — three parameterizations showed that combining two Gaussian factors is addition in natural-parameter space. This notebook follows that thread to its conclusion: an inference loop that absorbs Gaussian observations one at a time, equivalent to processing them all at once, in any order, with the same answer to floating-point precision.
That single idea — exponential-family inference reduces to summing natural parameters — underlies the Kalman filter, GP regression, EP, and most of approximate inference. We make it concrete by
- writing the linear-Gaussian Bayes rule once, in natural form, leaning on ((5));
- running a sequential loop over noisy observations of an unknown 2D location;
- confirming batch = sequential = any-order to ;
- closing with the punch line: GP regression with a Gaussian likelihood is a single application of the same rule.
Prerequisites: 0.2 — MultivariateNormal API, 0.4 — three parameterizations, 0.5 — Joseph-form covariance update.
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import einx
import jax
import jax.numpy as jnp
import lineax as lx
import matplotlib.pyplot as plt
import numpy as np
import gaussx
from gaussx import GaussianExpFam, log_marginal_likelihood, solve, to_expectation, to_natural
jax.config.update("jax_enable_x64", True)
KEY = jax.random.PRNGKey(42)
plt.rcParams.update({
"figure.dpi": 110,
"axes.grid": True,
"axes.grid.which": "both",
"xtick.minor.visible": True,
"ytick.minor.visible": True,
"grid.alpha": 0.3,
})
def psd_op(M):
return lx.MatrixLinearOperator(M, lx.symmetric_tag)
def ellipse_points(mu, Sigma, n_std=2.0, n=200):
# Standard 2D confidence-ellipse construction via eigendecomposition.
theta = jnp.linspace(0.0, 2.0 * jnp.pi, n)
circle = jnp.stack([jnp.cos(theta), jnp.sin(theta)], axis=0)
eigvals, eigvecs = jnp.linalg.eigh(Sigma)
sqrt_l = jnp.sqrt(eigvals) * n_std
stretched = einx.multiply("j, j t -> j t", sqrt_l, circle)
rotated = einx.dot("i j, j t -> i t", eigvecs, stretched)
return einx.add("i, i t -> i t", mu, rotated)Linear-Gaussian Bayes rule, in one breath¶
The linear-Gaussian observation model from ((1)) gives a Gaussian posterior with three equivalent updates:
In natural parameters, recalling and from ((2)), the rule is just ((5)) applied to the prior and the likelihood factor:
That’s the entire algorithm. No matrix inverses, no Kalman gains. The mean-cov form ((2)) and Joseph form ((6)) are equivalent recipes for getting back to when you need to plot or sample.
A 2D location-estimation problem¶
We’re trying to estimate an unknown 2D location from a sequence of noisy linear measurements. Start with a vague Gaussian prior and convert it to natural form via gaussx.to_natural.
x_true = jnp.array([3.0, -1.0])
mu0 = jnp.zeros(2)
Sigma0 = 4.0 * jnp.eye(2)
S0_op = psd_op(Sigma0)
eta1_prior, eta2_prior_op = to_natural(mu0, S0_op)
eta2_prior_mat = eta2_prior_op.as_matrix()
print("Prior mu :", np.asarray(mu0))
print("Prior Sigma :\n", np.asarray(Sigma0))
print("\nPrior eta1 :", np.asarray(eta1_prior))
print("Prior eta2 :\n", np.asarray(eta2_prior_mat))Prior mu : [0. 0.]
Prior Sigma :
[[4. 0.]
[0. 4.]]
Prior eta1 : [0. 0.]
Prior eta2 :
[[-0.125 -0. ]
[-0. -0.125]]
A single observation: natural form vs covariance form¶
We observe the -coordinate, with and .
Two paths to the posterior — both implementations of the same equation.
| Path | Recipe | Cost |
|---|---|---|
| Natural | Add likelihood naturals via ((2)). | One matrix add per observation |
| Mean-cov / Kalman | Compute Kalman gain , then via ((2)) | One innovation solve, one Joseph rebuild |
We do both and confirm agreement.
H1 = jnp.array([[1.0, 0.0]])
R1 = jnp.array([[1.0]])
key, sub = jax.random.split(KEY)
y1 = H1 @ x_true + jax.random.normal(sub, (1,)) * jnp.sqrt(R1[0, 0])
print("Observed y1 =", float(y1[0]))
# --- Natural update via [](#eq:bayes-natural) ---
R1_inv = jnp.linalg.inv(R1)
eta1_lik = einx.dot("k i, k l, l -> i", H1, R1_inv, y1)
eta2_lik_mat = -0.5 * einx.dot("k i, k l, l j -> i j", H1, R1_inv, H1)
eta1_post1 = eta1_prior + eta1_lik
eta2_post1_mat = eta2_prior_mat + eta2_lik_mat
post1 = GaussianExpFam(eta1=eta1_post1, eta2=psd_op(eta2_post1_mat))
mu1_nat, S1_nat_op = to_expectation(post1)
S1_nat = S1_nat_op.as_matrix()
# --- Kalman / mean-cov update ---
S_innov = einx.dot("k i, i j, l j -> k l", H1, Sigma0, H1) + R1
K_gain = einx.dot("i j, k j, k l -> i l", Sigma0, H1, jnp.linalg.inv(S_innov))
mu1_cov = mu0 + einx.dot("i k, k -> i", K_gain, y1 - einx.dot("k i, i -> k", H1, mu0))
S1_cov = Sigma0 - einx.dot("i k, k l, j l -> i j", K_gain, S_innov, K_gain)
print("\nNatural posterior mu :", np.asarray(mu1_nat))
print("Mean-cov posterior mu:", np.asarray(mu1_cov))
print(f"\nmax |mu_nat - mu_cov | = {float(jnp.max(jnp.abs(mu1_nat - mu1_cov))):.2e}")
print(f"max |Sig_nat - Sig_cov| = {float(jnp.max(jnp.abs(S1_nat - S1_cov))):.2e}")Observed y1 = 3.6498920118351403
Natural posterior mu : [2.91991361 0. ]
Mean-cov posterior mu: [2.91991361 0. ]
max |mu_nat - mu_cov | = 0.00e+00
max |Sig_nat - Sig_cov| = 2.22e-16
Same posterior to . The natural-form path replaced two matrix solves (innovation + Joseph rebuild) with two matrix adds — the difference compounds as we chain updates.
fig, ax = plt.subplots(figsize=(5.5, 5.0))
prior_pts = ellipse_points(mu0, Sigma0)
post_pts = ellipse_points(mu1_nat, S1_nat)
ax.plot(np.asarray(prior_pts[0]), np.asarray(prior_pts[1]),
color="0.4", lw=2, label=r"prior $2\sigma$")
ax.plot(np.asarray(post_pts[0]), np.asarray(post_pts[1]),
color="crimson", lw=2, label=r"posterior $2\sigma$")
ax.scatter(*np.asarray(x_true), marker="*", s=180, color="k",
label=r"$x_\star$", zorder=5)
ax.axvline(float(y1[0]), color="steelblue", lw=1.4, ls="--",
label=fr"$y_1 = {float(y1[0]):.2f}$ (vertical likelihood ridge)")
ax.set_xlabel(r"$x_1$"); ax.set_ylabel(r"$x_2$")
ax.set_title(r"Single-observation update: prior $\to$ posterior")
ax.set_aspect("equal"); ax.legend(frameon=False, fontsize=8)
plt.tight_layout(); plt.show()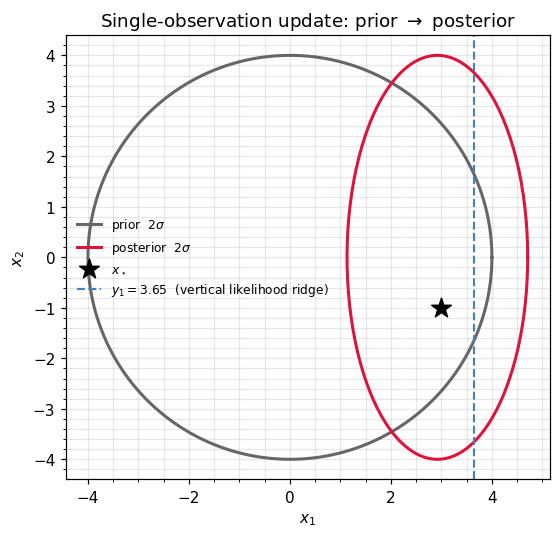
Sequential updates: factors, one addition at a time¶
Five additional measurements, each through a different (single coordinates and diagonal projections), each with . The update rule is just ((2)) applied repeatedly:
Each step is an array sum — no matrix inverses during the recursion. We only invert (via to_expectation) when we want to plot the running posterior.
H_list = [
jnp.array([[0.0, 1.0]]), # y-coordinate
jnp.array([[1.0, 1.0]]) / jnp.sqrt(2.0), # diagonal
jnp.array([[1.0, 0.0]]), # x-coordinate again
jnp.array([[0.0, 1.0]]), # y-coordinate again
jnp.array([[1.0, -1.0]]) / jnp.sqrt(2.0), # anti-diagonal
]
R_obs = jnp.array([[0.5]])
R_obs_inv = jnp.linalg.inv(R_obs)
key = sub
ys = []
for H_i in H_list:
key, k_obs = jax.random.split(key)
y_i = einx.dot("k i, i -> k", H_i, x_true) \
+ jax.random.normal(k_obs, (1,)) * jnp.sqrt(R_obs[0, 0])
ys.append(y_i)
# Recursion starting from the post-1 state.
eta1 = eta1_post1
eta2 = eta2_post1_mat
mus = [mu0, mu1_nat]
Sigs = [Sigma0, S1_nat]
for n, (H_i, y_i) in enumerate(zip(H_list, ys, strict=True), start=2):
eta1 = eta1 + einx.dot("k i, k l, l -> i", H_i, R_obs_inv, y_i)
eta2 = eta2 + (-0.5) * einx.dot("k i, k l, l j -> i j", H_i, R_obs_inv, H_i)
post_n = GaussianExpFam(eta1=eta1, eta2=psd_op(eta2))
mu_n, S_n_op = to_expectation(post_n)
mus.append(mu_n); Sigs.append(S_n_op.as_matrix())
print(f"after obs {n}: mu = [{float(mu_n[0]):+.3f}, {float(mu_n[1]):+.3f}] "
f"sqrt(tr Sigma) = {float(jnp.sqrt(jnp.trace(S_n_op.as_matrix()))):.3f}")
print(f"\nx_true = [{float(x_true[0]):.3f}, {float(x_true[1]):.3f}]")after obs 2: mu = [+2.920, -0.159] sqrt(tr Sigma) = 1.116
after obs 3: mu = [+2.622, -0.324] sqrt(tr Sigma) = 0.933
after obs 4: mu = [+2.988, -0.437] sqrt(tr Sigma) = 0.765
after obs 5: mu = [+3.130, -1.041] sqrt(tr Sigma) = 0.668
after obs 6: mu = [+2.816, -0.777] sqrt(tr Sigma) = 0.592
x_true = [3.000, -1.000]
fig, ax = plt.subplots(figsize=(6.6, 6.0))
n_steps = len(mus)
colors = plt.cm.viridis(np.linspace(0.1, 0.9, n_steps))
for k in range(n_steps):
pts = ellipse_points(mus[k], Sigs[k])
label = "prior" if k == 0 else f"after obs {k}"
ax.plot(np.asarray(pts[0]), np.asarray(pts[1]),
color=colors[k], lw=1.8, label=label)
ax.scatter(float(mus[k][0]), float(mus[k][1]),
s=22, color=colors[k], zorder=4)
ax.scatter(*np.asarray(x_true), marker="*", s=240, color="crimson",
label=r"$x_\star$", zorder=5)
ax.set_xlabel(r"$x_1$"); ax.set_ylabel(r"$x_2$")
ax.set_title(r"Sequential Bayesian updates — $\eta$ accumulates by addition")
ax.set_aspect("equal"); ax.legend(frameon=False, fontsize=8, loc="upper left")
plt.tight_layout(); plt.show()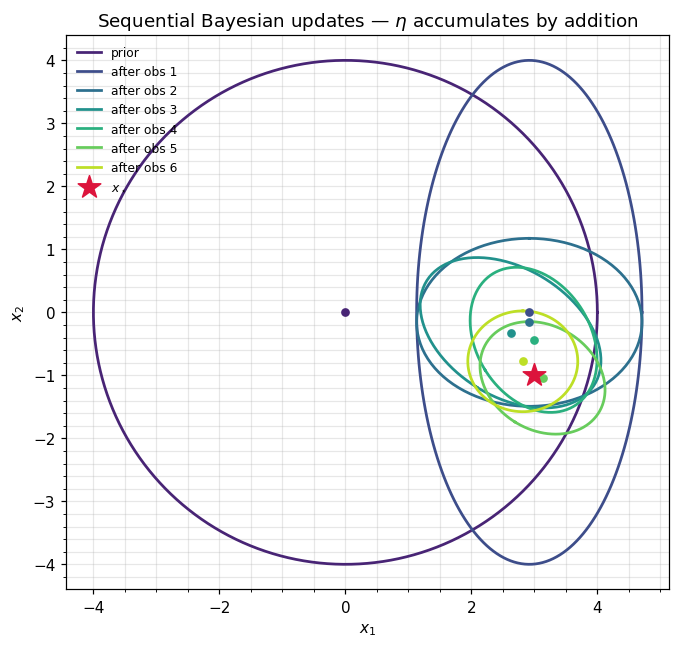
Batch = sequential = any order¶
Vector addition is commutative and associative, so the recursion ((3)) doesn’t care in what order we apply the likelihood factors:
This is not a coincidence of Gaussian arithmetic — it’s the defining property of exponential-family inference. Concretely:
# Stack ALL observations (including y1) and compare three computations.
H_all = [H1, *H_list]
y_all = [y1, *ys]
R_all = [R1, *([R_obs] * len(ys))]
def batch_natural(perm):
eta1 = eta1_prior
eta2 = eta2_prior_mat
for j in perm:
H, y, R = H_all[j], y_all[j], R_all[j]
Rinv = jnp.linalg.inv(R)
eta1 = eta1 + einx.dot("k i, k l, l -> i", H, Rinv, y)
eta2 = eta2 + (-0.5) * einx.dot("k i, k l, l j -> i j", H, Rinv, H)
return eta1, eta2
# (a) sequential = the order in which obs arrived
eta1_seq, eta2_seq = batch_natural(list(range(len(H_all))))
# (b) reversed
eta1_rev, eta2_rev = batch_natural(list(range(len(H_all)))[::-1])
# (c) random permutation
rng = np.random.default_rng(0)
perm = rng.permutation(len(H_all)).tolist()
eta1_rng, eta2_rng = batch_natural(perm)
ord_err_seq_rev = float(jnp.max(jnp.abs(eta1_seq - eta1_rev))) \
+ float(jnp.max(jnp.abs(eta2_seq - eta2_rev)))
ord_err_seq_rng = float(jnp.max(jnp.abs(eta1_seq - eta1_rng))) \
+ float(jnp.max(jnp.abs(eta2_seq - eta2_rng)))
# Compare to the running sequential trajectory's last entry.
last_post = GaussianExpFam(eta1=eta1_seq, eta2=psd_op(eta2_seq))
mu_batch, S_batch_op = to_expectation(last_post)
S_batch = S_batch_op.as_matrix()
trajectory_last_mu = mus[-1]
trajectory_last_Sig = Sigs[-1]
print(f"max ||eta(seq) - eta(reversed)|| = {ord_err_seq_rev:.2e}")
print(f"max ||eta(seq) - eta(random {perm})|| = {ord_err_seq_rng:.2e}")
print(f"max ||mu(batch) - mu(traj last)|| = "
f"{float(jnp.max(jnp.abs(mu_batch - trajectory_last_mu))):.2e}")
print(f"max ||Sig(batch) - Sig(traj last)|| = "
f"{float(jnp.max(jnp.abs(S_batch - trajectory_last_Sig))):.2e}")max ||eta(seq) - eta(reversed)|| = 1.78e-15
max ||eta(seq) - eta(random [3, 2, 5, 4, 0, 1])|| = 1.78e-15
max ||mu(batch) - mu(traj last)|| = 0.00e+00
max ||Sig(batch) - Sig(traj last)|| = 0.00e+00
All differences sit at — the recursion really is permutation-invariant up to round-off.
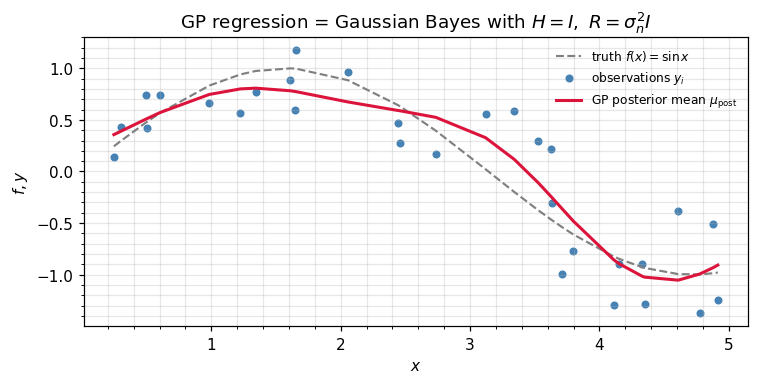
GP regression is this update¶
Gaussian-process regression with a Gaussian likelihood is precisely ((2)) applied with (we observe the function directly) and :
The standard GP posterior-mean formula on the right is ((5)) rearranged via the matrix-inversion identity. Different recipe, same equation.
Operationally, gaussx covers the whole loop:
N_gp = 30
key, k_x, k_y = jax.random.split(KEY, 3)
xs = jnp.sort(jax.random.uniform(k_x, (N_gp,), minval=0.0, maxval=5.0))
# RBF kernel and noisy observations of f(x) = sin(x)
ls, sigma2 = 1.0, 0.1
diff = xs[:, None] - xs[None, :]
K_gp = jnp.exp(-0.5 * diff ** 2 / ls ** 2)
f_true = jnp.sin(xs)
y_gp = f_true + jax.random.normal(k_y, (N_gp,)) * jnp.sqrt(sigma2)
# Posterior mean: K (K + sigma^2 I)^{-1} y -- one structured solve.
K_op = psd_op(K_gp + sigma2 * jnp.eye(N_gp))
alpha = solve(K_op, y_gp)
mu_gp = einx.dot("i j, j -> i", K_gp, alpha)
# Log marginal likelihood — used downstream for hyperparameter learning.
lml = log_marginal_likelihood(loc=jnp.zeros(N_gp), cov_operator=K_op, y=y_gp)
rmse = float(jnp.sqrt(jnp.mean((mu_gp - f_true) ** 2)))
print(f"GP log marginal likelihood : {float(lml):.3f}")
print(f"posterior-mean RMSE vs sin : {rmse:.4f}")
fig, ax = plt.subplots(figsize=(7.0, 3.6))
ax.plot(np.asarray(xs), np.asarray(f_true), "--", color="0.5",
lw=1.4, label=r"truth $f(x) = \sin x$")
ax.scatter(np.asarray(xs), np.asarray(y_gp), s=18, color="steelblue",
label=r"observations $y_i$")
ax.plot(np.asarray(xs), np.asarray(mu_gp), color="crimson", lw=2,
label=r"GP posterior mean $\mu_{\rm post}$")
ax.set_xlabel(r"$x$"); ax.set_ylabel(r"$f, y$")
ax.set_title(r"GP regression = Gaussian Bayes with $H = I,\ R = \sigma_n^2 I$")
ax.legend(frameon=False, fontsize=8)
plt.tight_layout(); plt.show()GP log marginal likelihood : -19.557
posterior-mean RMSE vs sin : 0.1561
Where this update lives in practice¶
| Setting | What “factor” means | Where in the curriculum |
|---|---|---|
| Kalman filter (part 7) | Each observation step = one Gaussian factor folded in via ((2)). The information form is the running natural-parameter sum; the standard form is its mean-cov twin. | Information-form filter / smoother. |
| GP regression (part 3) | All training observations are one big factor with . Posterior reduces to a single solve ((5)). | Exact, sparse, and approximate GP recipes. |
| Ensemble Kalman / data assimilation (part 7.x) | Each ensemble member or observation batch contributes a likelihood factor. Order-independence ((4)) lets you parallelise across sensors. | EnKF, particle Kalman, sequential 3D-Var. |
| EP for non-Gaussian likelihoods (part 6) | Each site approximation is a Gaussian factor in natural form. The cavity-tilted-update cycle is ((2)) with cavity = posterior − site, site = projection − cavity. | Classification GPs, robust regression. |
| Conjugate Bayesian linear regression | is the design matrix, . Sequential additions handle streaming data. | Recursive least squares as a degenerate Kalman filter. |
| Distributed inference | Each worker computes on its shard; the parameter server sums them. | Federated GP / Bayesian linear models. |
Recap¶
- Linear-Gaussian Bayes rule = one application of ((5)) with the likelihood factor read off ((1)). No matrix inverses during the recursion.
- observations = additions: ((3)). Mean-cov / Joseph forms are equivalent recipes if you need at any point.
- Order-independence ((4)) is exact (modulo round-off) and is the reason exponential-family inference parallelises so cleanly.
- GP regression with a Gaussian likelihood is one shot of this update ((5)), executed via
gaussx.solveon the structured operator .
References¶
- Bernardo, J. M. & Smith, A. F. M. (2000). Bayesian Theory. Wiley.
- Bishop, C. M. (2006). Pattern Recognition and Machine Learning. Springer. §2.3.
- Rasmussen, C. E. & Williams, C. K. I. (2006). Gaussian Processes for Machine Learning. MIT Press. §2.2.
- Minka, T. P. (2001). A family of algorithms for approximate Bayesian inference. PhD thesis, MIT.