Matrix-Free GP with ImplicitKernelOperator
For large , forming the kernel matrix requires
memory and time for dense factorizations.
ImplicitKernelOperator avoids materializing entirely: it computes
on-the-fly by evaluating the kernel function row-by-row, using
only memory. Combined with CG-based solvers, this enables GP
inference on problems that are too large for dense methods.
What you’ll learn:
- Building a matrix-free kernel operator with
ImplicitKernelOperator - Solving kernel systems with
CGSolver,PreconditionedCGSolver, andBBMMSolver - Using structural tags for metadata
- Exploiting orthonormal low-rank structure with
LowRankUpdate - Full GP prediction: implicit vs dense
1. Setup¶
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import jax
import jax.numpy as jnp
import lineax as lx
import matplotlib.pyplot as plt
import gaussx
jax.config.update("jax_enable_x64", True)
# Register missing lineax dispatches for ImplicitKernelOperator so
# that lineax CG (used internally by gaussx strategies) can work.
lx.is_negative_semidefinite.register(gaussx.ImplicitKernelOperator)(lambda _: False)
lx.linearise.register(gaussx.ImplicitKernelOperator)(lambda op: op)
lx.is_negative_semidefinite.register(gaussx.LowRankUpdate)(lambda _: False)
lx.linearise.register(gaussx.LowRankUpdate)(lambda op: op)<function __main__.<lambda>(op)>We generate a 1D GP regression problem with noisy observations of a smooth function.
key = jax.random.PRNGKey(42)
N = 2000
# Training data: noisy sine
key_x, key_noise = jax.random.split(key)
X_train = jnp.sort(jax.random.uniform(key_x, (N, 1), minval=-5.0, maxval=5.0), axis=0)
f_true = jnp.sin(X_train[:, 0]) + 0.2 * jnp.cos(3.0 * X_train[:, 0])
noise_var = 0.1
y_train = f_true + jnp.sqrt(noise_var) * jax.random.normal(key_noise, (N,))
print(f"Training set: N = {N}, D = {X_train.shape[1]}")
print(f"Noise variance: {noise_var}")Training set: N = 2000, D = 1
Noise variance: 0.1
2. Build ImplicitKernelOperator¶
We define a scalar RBF kernel
where are 1D arrays of shape (D,). The kernel function is
passed to ImplicitKernelOperator, which wraps it into a lineax
operator that computes without ever forming .
# Kernel hyperparameters
length_scale = 1.0
signal_var = 1.0
def rbf_kernel(x: jnp.ndarray, xp: jnp.ndarray) -> jnp.ndarray:
"""Scalar RBF kernel: k(x, x') -> scalar."""
sq_dist = jnp.sum((x - xp) ** 2)
return signal_var * jnp.exp(-0.5 * sq_dist / length_scale**2)
op = gaussx.ImplicitKernelOperator(
rbf_kernel,
X_train,
noise_var=noise_var,
tags=frozenset({lx.positive_semidefinite_tag, lx.symmetric_tag}),
)
print(f"Operator size: {op.in_size()} x {op.out_size()}")Operator size: 2000 x 2000
The operator supports mv (matrix-vector product) for CG-based
methods. It also has as_matrix() for debugging, but that
materializes the full matrix -- exactly what we want to
avoid at scale.
# Verify matvec matches dense
v = jax.random.normal(jax.random.PRNGKey(0), (N,))
mv_implicit = op.mv(v)
# Dense reference (expensive -- only for validation)
K_dense = op.as_matrix()
mv_dense = K_dense @ v
err = jnp.max(jnp.abs(mv_implicit - mv_dense))
print(f"Matvec error (implicit vs dense): {err:.2e}")Matvec error (implicit vs dense): 5.68e-14
3. Solve with CGSolver¶
CGSolver uses lineax CG for the linear solve and matfree’s stochastic Lanczos quadrature (SLQ) for the
log-determinant. It only needs matvec access -- no matrix
materialization.
cg = gaussx.CGSolver(rtol=1e-6, atol=1e-6, max_steps=200, num_probes=30)
alpha_cg = cg.solve(op, y_train)
residual_cg = jnp.max(jnp.abs(op.mv(alpha_cg) - y_train))
print(f"CG solve residual: {residual_cg:.2e}")CG solve residual: 9.80e-07
logdet_cg = cg.logdet(op, key=jax.random.PRNGKey(0))
print(f"CG stochastic logdet: {logdet_cg:.4f}")
# Dense reference logdet
logdet_dense = jnp.linalg.slogdet(K_dense)[1]
print(f"Dense logdet: {logdet_dense:.4f}")
print(f"Logdet error: {jnp.abs(logdet_cg - logdet_dense):.2e}")CG stochastic logdet: -4522.7308
Dense logdet: -4522.1695
Logdet error: 5.61e-01
4. Solve with PreconditionedCGSolver¶
PreconditionedCGSolver builds a rank- partial Cholesky
preconditioner that dramatically reduces the number of CG iterations
for ill-conditioned kernel systems. The shift parameter should match
the noise variance.
pcg = gaussx.PreconditionedCGSolver(
preconditioner_rank=30,
shift=noise_var,
rtol=1e-6,
atol=1e-6,
)
alpha_pcg = pcg.solve(op, y_train)
residual_pcg = jnp.max(jnp.abs(op.mv(alpha_pcg) - y_train))
print(f"Preconditioned CG solve residual: {residual_pcg:.2e}")
# Compare with plain CG
diff_pcg_cg = jnp.max(jnp.abs(alpha_pcg - alpha_cg))
print(f"Max |alpha_pcg - alpha_cg|: {diff_pcg_cg:.2e}")Preconditioned CG solve residual: 9.76e-08
Max |alpha_pcg - alpha_cg|: 2.02e-06
5. Solve with BBMMSolver¶
BBMMSolver (Gardner et al. 2018) jointly computes the solve and
log-determinant via solve_and_logdet, amortizing the matvec cost.
bbmm = gaussx.BBMMSolver(
cg_max_iter=200,
cg_tolerance=1e-6,
lanczos_iter=50,
num_probes=20,
)
alpha_bbmm, ld_bbmm = bbmm.solve_and_logdet(op, y_train)
residual_bbmm = jnp.max(jnp.abs(op.mv(alpha_bbmm) - y_train))
print(f"BBMM solve residual: {residual_bbmm:.2e}")
print(f"BBMM logdet: {ld_bbmm:.4f}")
print(f"Dense logdet: {logdet_dense:.4f}")BBMM solve residual: 9.80e-07
BBMM logdet: -4527.8656
Dense logdet: -4522.1695
Solver comparison¶
All three iterative solvers converge to the same solution.
print("Max difference between solvers:")
print(f" CG vs PCG: {jnp.max(jnp.abs(alpha_cg - alpha_pcg)):.2e}")
print(f" CG vs BBMM: {jnp.max(jnp.abs(alpha_cg - alpha_bbmm)):.2e}")
print(f" PCG vs BBMM: {jnp.max(jnp.abs(alpha_pcg - alpha_bbmm)):.2e}")Max difference between solvers:
CG vs PCG: 2.02e-06
CG vs BBMM: 0.00e+00
PCG vs BBMM: 2.02e-06
6. Tags for structural metadata¶
ImplicitKernelOperator accepts lineax tags to declare structural
properties. Kernel matrices from stationary kernels are symmetric
and positive semi-definite; declaring this lets lineax and gaussx
exploit the structure.
op_tagged = gaussx.ImplicitKernelOperator(
rbf_kernel,
X_train,
noise_var=noise_var,
tags=frozenset({lx.symmetric_tag, lx.positive_semidefinite_tag}),
)
print(f"is_symmetric: {gaussx.is_symmetric(op_tagged)}")
print(f"is_positive_semidefinite: {gaussx.is_positive_semidefinite(op_tagged)}")is_symmetric: True
is_positive_semidefinite: True
7. Orthonormal low-rank updates via LowRankUpdate¶
When a truncated SVD approximation of the kernel matrix is available
(e.g., from a Nystrom approximation), LowRankUpdate can represent
the operator as . Setting
orthonormal=True preserves the SVD-specific symmetry inference
without a separate operator class.
Here we compute a dense truncated SVD for demonstration. In practice, you would obtain from a randomized or Nystrom method without forming the full matrix.
rank = 50
# Truncated SVD of the kernel matrix (dense, for demonstration only)
K_no_noise = K_dense - noise_var * jnp.eye(N)
U_full, S_full, Vt_full = jnp.linalg.svd(K_no_noise, full_matrices=False)
U_k = U_full[:, :rank]
S_k = S_full[:rank]
V_k = Vt_full[:rank, :].T # (N, rank)
# Base operator: noise_var * I
base_diag = lx.DiagonalLinearOperator(noise_var * jnp.ones(N))
svd_op = gaussx.LowRankUpdate(
base_diag,
U_k,
S_k,
V_k,
tags=frozenset({lx.positive_semidefinite_tag, lx.symmetric_tag}),
orthonormal=True,
)
print(f"LowRankUpdate: rank-{rank} approximation")
print(f"Operator size: {svd_op.in_size()} x {svd_op.out_size()}")LowRankUpdate: rank-50 approximation
Operator size: 2000 x 2000
# Solve with the SVD low-rank operator via CG
alpha_svd = cg.solve(svd_op, y_train)
residual_svd = jnp.max(jnp.abs(svd_op.mv(alpha_svd) - y_train))
print(f"SVD solve residual: {residual_svd:.2e}")
# Logdet via SLQ
logdet_svd = cg.logdet(svd_op, key=jax.random.PRNGKey(1))
print(f"SVD logdet: {logdet_svd:.4f}")
print(f"Dense logdet: {logdet_dense:.4f}")SVD solve residual: 1.05e-06
SVD logdet: -4528.3541
Dense logdet: -4522.1695
8. GP prediction¶
With in hand, the posterior mean at test points is
We compare predictions from the implicit (CG) solve with the dense Cholesky baseline.
# Test points
X_test = jnp.linspace(-5.5, 5.5, 300)[:, None]
# Cross-covariance: K(X_test, X_train) via vmap
K_star = jax.vmap(
lambda x_star: jax.vmap(lambda x_tr: rbf_kernel(x_star, x_tr))(X_train)
)(X_test)
# Predictions from implicit CG solve
mu_cg = K_star @ alpha_cg
# Dense baseline
alpha_dense = jnp.linalg.solve(K_dense, y_train)
mu_dense = K_star @ alpha_dense
pred_err = jnp.max(jnp.abs(mu_cg - mu_dense))
print(f"Max prediction error (CG vs dense): {pred_err:.2e}")Max prediction error (CG vs dense): 8.86e-07
9. Plot: implicit vs dense GP predictions¶
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
# Left: predictions
ax = axes[0]
ax.scatter(
X_train[:, 0],
y_train,
s=30,
alpha=0.3,
color="gray",
edgecolors="k",
linewidths=0.5,
label="Train data",
zorder=5,
)
ax.plot(X_test[:, 0], mu_cg, color="C0", lw=2, label="CG (implicit)", zorder=3)
ax.plot(
X_test[:, 0],
mu_dense,
"--",
color="C1",
lw=2,
label="Dense (Cholesky)",
zorder=3,
)
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.set_title("GP posterior mean")
ax.legend(fontsize=9)
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
# Right: absolute difference
ax = axes[1]
ax.plot(X_test[:, 0], jnp.abs(mu_cg - mu_dense), color="C2", lw=2)
ax.set_xlabel("x")
ax.set_ylabel("|CG - Dense|")
ax.set_title("Prediction difference")
ax.set_yscale("log")
ax.grid(True, which="major", alpha=0.3)
ax.grid(True, which="minor", alpha=0.1)
ax.minorticks_on()
plt.tight_layout()
plt.show()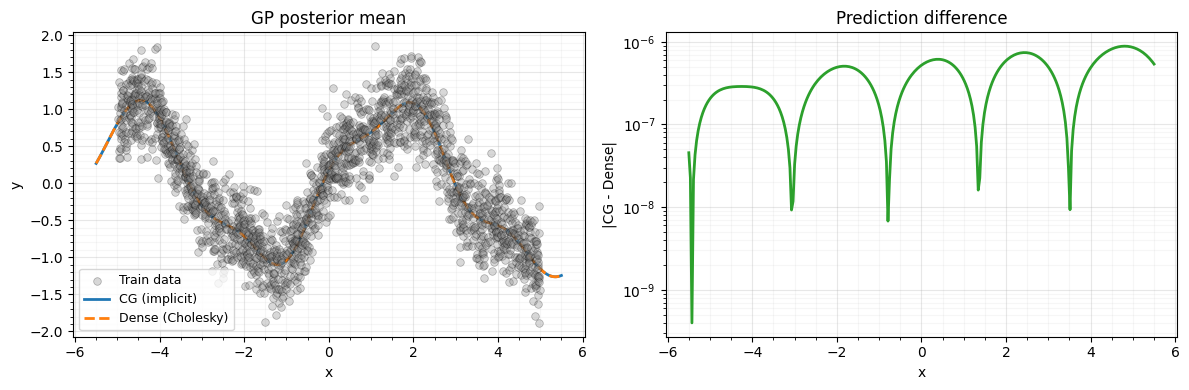
Summary¶
ImplicitKernelOperator-- Matrix-free via on-the-fly kernel evaluation. memory.CGSolver-- Iterative CG solve + SLQ logdet. Only needs matvec access.PreconditionedCGSolver-- Partial Cholesky preconditioner for faster CG convergence.BBMMSolver-- Joint solve + logdet via batched CG (Gardner et al. 2018).LowRankUpdate(..., orthonormal=True)-- SVD-parameterized low-rank structure for cheap solves and logdets.
When to use what:
- Small (): Dense Cholesky is fastest and exact.
- Medium (5000--50000):
CGSolverorPreconditionedCGSolverwithImplicitKernelOperator. - Large ():
BBMMSolverwithImplicitKernelOperatoramortizes matvec costs across solve and logdet. UsePreconditionedCGSolverif the system is ill-conditioned. - Low-rank structure available:
LowRankUpdate(..., orthonormal=True)exploits the factored form directly.