Kronecker Eigendecomposition
For a Kronecker product , all eigenvalues and eigenvectors decompose per-factor:
This means we can compute the full spectrum of an
matrix () by only decomposing two small matrices.
gaussx exploits this for cholesky, sqrt, logdet, and inv.
Context¶
The Kronecker eigendecomposition is a special case of the tensor product spectral theorem. If and , then
This result is foundational for:
- GP inference on grids -- computing solves and log-determinants without forming the full matrix (Saatci, 2012).
- Solving Kronecker-structured linear systems -- reducing an -dimensional problem to two smaller per-factor problems.
- Stochastic PDEs with separable operators -- exploiting product structure in spatial and temporal discretizations.
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import jax
import jax.numpy as jnp
import lineax as lx
import matplotlib.pyplot as plt
import gaussx
jax.config.update("jax_enable_x64", True)Setup: PSD Kronecker product¶
key = jax.random.PRNGKey(42)
k1, k2 = jax.random.split(key)
n1, n2 = 8, 10
N = n1 * n2
# Random PSD matrices
M1 = jax.random.normal(k1, (n1, n1))
A = M1 @ M1.T + 0.1 * jnp.eye(n1)
M2 = jax.random.normal(k2, (n2, n2))
B = M2 @ M2.T + 0.1 * jnp.eye(n2)
A_op = lx.MatrixLinearOperator(A, lx.positive_semidefinite_tag)
B_op = lx.MatrixLinearOperator(B, lx.positive_semidefinite_tag)
K = gaussx.Kronecker(A_op, B_op)
print(f"A: {n1}x{n1}, B: {n2}x{n2}")
print(f"A kron B: {N}x{N} = {N**2:,} entries")A: 8x8, B: 10x10
A kron B: 80x80 = 6,400 entries
Per-factor eigenvalues¶
eigs_A = jnp.linalg.eigvalsh(A)
eigs_B = jnp.linalg.eigvalsh(B)
# Kronecker eigenvalues = outer product of per-factor eigenvalues
eigs_kron = jnp.sort(jnp.outer(eigs_A, eigs_B).ravel())
# Dense eigenvalues (for verification)
eigs_dense = jnp.linalg.eigvalsh(K.as_matrix())
print(f"Per-factor eigenvalues: {n1} + {n2} = {n1 + n2} eigh calls")
print(f"Dense eigenvalues: one {N}x{N} eigh call")
print(f"Max eigenvalue error: {jnp.max(jnp.abs(eigs_kron - eigs_dense)):.2e}")Per-factor eigenvalues: 8 + 10 = 18 eigh calls
Dense eigenvalues: one 80x80 eigh call
Max eigenvalue error: 2.84e-13
fig, axes = plt.subplots(1, 3, figsize=(14, 3.5))
axes[0].stem(range(n1), eigs_A, linefmt="C0-", markerfmt="C0o", basefmt="k-")
axes[0].set_title(f"Eigenvalues of A ({n1}x{n1})")
axes[0].set_xlabel("Index")
axes[0].set_ylabel("$\\lambda$")
axes[0].grid(True, which="major", alpha=0.3)
axes[0].grid(True, which="minor", alpha=0.1)
axes[0].minorticks_on()
axes[1].stem(range(n2), eigs_B, linefmt="C1-", markerfmt="C1o", basefmt="k-")
axes[1].set_title(f"Eigenvalues of B ({n2}x{n2})")
axes[1].set_xlabel("Index")
axes[1].grid(True, which="major", alpha=0.3)
axes[1].grid(True, which="minor", alpha=0.1)
axes[1].minorticks_on()
axes[2].semilogy(eigs_kron, "C2-", lw=2, label="Kronecker (per-factor)")
axes[2].semilogy(eigs_dense, "k--", lw=1.5, label="Dense (verification)")
axes[2].set_title(f"Eigenvalues of A$\\otimes$B ({N}x{N})")
axes[2].set_xlabel("Index")
axes[2].legend(fontsize=9)
axes[2].grid(True, which="major", alpha=0.3)
axes[2].grid(True, which="minor", alpha=0.1)
axes[2].minorticks_on()
plt.tight_layout()
plt.show()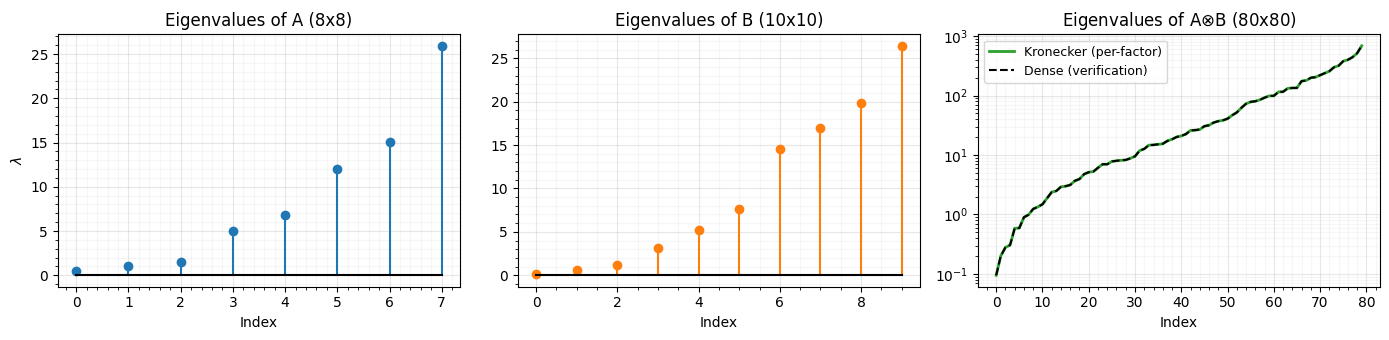
Structured Cholesky¶
cholesky(A kron B) = cholesky(A) kron cholesky(B)
L = gaussx.cholesky(K)
print(f"cholesky type: {type(L).__name__}")
# Reconstruction error
recon = L.as_matrix() @ L.as_matrix().T
print(f"||L L^T - K||_max: {jnp.max(jnp.abs(recon - K.as_matrix())):.2e}")cholesky type: Kronecker
||L L^T - K||_max: 5.68e-14
Structured sqrt¶
sqrt(A kron B) = sqrt(A) kron sqrt(B)
S = gaussx.sqrt(K)
print(f"sqrt type: {type(S).__name__}")
# Verify S @ S = K
recon_sqrt = S.as_matrix() @ S.as_matrix()
print(f"||S S - K||_max: {jnp.max(jnp.abs(recon_sqrt - K.as_matrix())):.2e}")sqrt type: Kronecker
||S S - K||_max: 4.41e-13
ld_structured = gaussx.logdet(K)
ld_dense = jnp.linalg.slogdet(K.as_matrix())[1]
ld_from_eigs = jnp.sum(jnp.log(eigs_kron))
print(f"Structured logdet: {ld_structured:.6f}")
print(f"Dense logdet: {ld_dense:.6f}")
print(f"From eigenvalues: {ld_from_eigs:.6f}")Structured logdet: 233.876851
Dense logdet: 233.876851
From eigenvalues: 233.876851
Summary¶
| Operation | Dense cost | Kronecker cost | Speedup |
|---|---|---|---|
| Eigenvalues | O(N^3) | O(n1^3 + n2^3) | ~N/n_max |
| Cholesky | O(N^3) | O(n1^3 + n2^3) | ~N/n_max |
| Logdet | O(N^3) | O(n1^3 + n2^3) | ~N/n_max |
| Solve | O(N^3) | O(n1^3 + n2^3) | ~N/n_max |
References¶
- Van Loan, C. F. (2000). The ubiquitous Kronecker product. Journal of Computational and Applied Mathematics, 123, 85--100.
- Saatci, Y. (2012). Scalable Inference for Structured Gaussian Process Models. PhD thesis, University of Cambridge.
- Steeb, W.-H. (2011). Matrix Calculus and Kronecker Product. 2nd edition, World Scientific.