Toeplitz operators — stationary kernels on regular grids
A Toeplitz matrix has constant diagonals: . Storage is — just the first column . Matvec is via FFT and the convolution theorem. For stationary kernels (RBF, Matérn, periodic) evaluated on a regular 1-D grid, the resulting covariance matrix is exactly Toeplitz, so we get a billion-fold storage saving and a thousand-fold compute saving over the dense form, with no approximation.
This is one of the cleanest “structure for free” wins in the catalog. Almost every classical signal-processing and time-series technique — autoregressive estimation, Wiener filtering, spectral analysis, fast power-spectrum density estimation — is, at heart, a Toeplitz solve.
0. Where Toeplitz structure shows up in geoscience¶
Wherever you sample a stationary process on a regular grid, you get a Toeplitz covariance. The relevant geoscience domains:
- Time series of a single station / mooring / buoy: tide gauges, pCO2 moorings, soil-moisture probes, GPS displacements. Sampling cadence is regular (hourly / daily / monthly), and the stationary kernel matrix is Toeplitz directly.
- 1-D transects on a regular spatial grid: along-track satellite-altimeter data (sea-surface height), ice-core depth profiles, vertical sounding profiles (radiosonde temperature vs altitude), DAS strain along a fibre. As long as the sampling spacing is uniform, the covariance is Toeplitz.
- 2-D stationary fields on a regular grid: this is the workhorse. Sea-surface temperature on a lat × lon grid, atmospheric reanalyses, gridded CO2 flux, surface displacements from InSAR — the covariance becomes
Kronecker(Toeplitz_x, Toeplitz_y), and 2-D matvec costs instead of . - Spectral analysis & autocovariance estimation: the periodogram, Welch’s method, Lomb–Scargle, AR(p) fits — all of these compute or invert Toeplitz matrices implicitly. The Wiener–Khinchin theorem (covariance = inverse Fourier transform of the spectrum) is the same FFT trick formalized at the population level.
- Stationary GP regression on equispaced data: the entire toolkit of [Wood 2017] / [Kaufman & Sain 2008] for GP regression on regular grids relies on Toeplitz solves to scale to time series points.
The catch: stationarity is required (kernel depends only on , not on absolute position) and regular sampling is required (constant grid spacing). Drop either assumption — non-stationary process, irregular timestamps — and you fall back to dense kernels (or 1.6 MaskedOperator for missing data on a regular grid).
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import einx
import jax
import jax.numpy as jnp
import lineax as lx
import matplotlib.pyplot as plt
import numpy as np
from gaussx import (
Kronecker,
Toeplitz,
logdet,
)
jax.config.update("jax_enable_x64", True)
KEY = jax.random.PRNGKey(0)
plt.rcParams.update({
"figure.dpi": 110,
"axes.grid": True,
"axes.grid.which": "both",
"xtick.minor.visible": True,
"ytick.minor.visible": True,
"grid.alpha": 0.3,
})1. The Toeplitz structure¶
A symmetric Toeplitz matrix is fully determined by its first column :
Storage: floats instead of . Symmetry is built in.
For a stationary kernel evaluated on a regular grid , the entries are — exactly Toeplitz with . The first column is the kernel function sampled at the grid spacings.
# Stationary RBF kernel on a regular grid → Toeplitz
n = 32
grid = jnp.linspace(0.0, 4.0, n)
ell = 0.5
col = jnp.exp(-0.5 * (grid - grid[0])**2 / ell**2)
T = Toeplitz(col)
T_dense = T.as_matrix()
# Visualize the constant-diagonal structure
fig, axes = plt.subplots(1, 2, figsize=(11, 4.5))
axes[0].plot(grid, col, "C0o-")
axes[0].set_xlabel(r"lag $\tau = i\Delta x$")
axes[0].set_ylabel(r"$c_i = k(\tau)$")
axes[0].set_title(r"First column = kernel sampled at grid lags")
im = axes[1].imshow(T_dense, cmap="viridis")
axes[1].set_title(r"Symmetric Toeplitz: constant diagonals $K_{ij}=c_{|i-j|}$")
axes[1].set_xlabel("$j$")
axes[1].set_ylabel("$i$")
plt.colorbar(im, ax=axes[1], shrink=0.8)
plt.tight_layout()
plt.show()
print(f"storage : {col.size} floats (vs dense {T_dense.size})")
print(f"saving : {T_dense.size / col.size:.0f}x")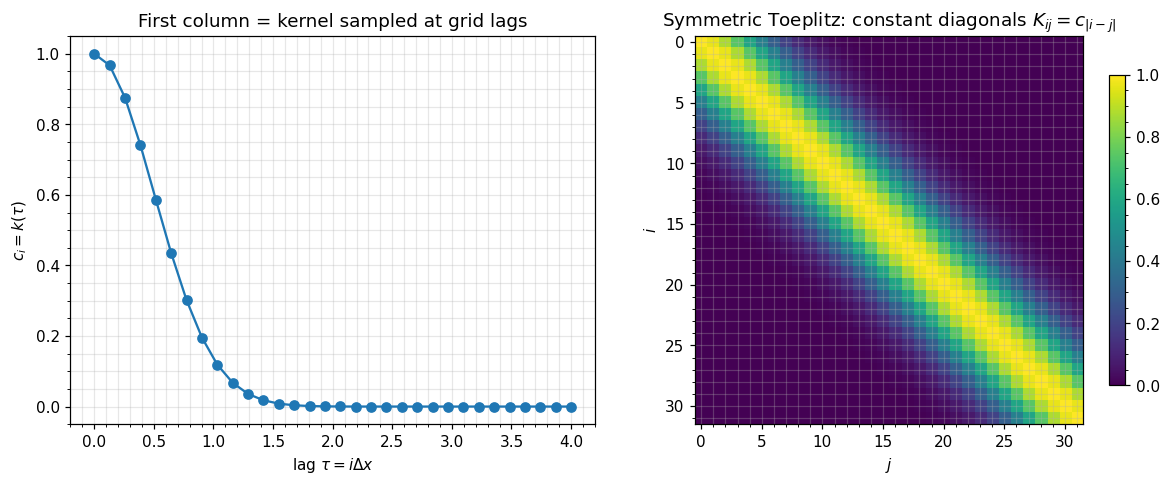
storage : 32 floats (vs dense 1024)
saving : 32x
2. Matvec via circulant embedding + FFT¶
The trick. Embed the Toeplitz matrix into a circulant matrix by appending the reflected column:
A circulant matrix is diagonalized by the Fourier basis, so multiplying it is element-wise multiplication in the frequency domain (the convolution theorem). The matvec on the original Toeplitz vector is:
where is zero-padded to length . Cost: three FFTs of length , total . Compared to dense matvec at this is a thousand-fold speedup at .
gaussx.Toeplitz.mv implements exactly this — let’s verify.
v = jax.random.normal(jax.random.PRNGKey(1), (n,))
y_fft = T.mv(v)
y_dense = T_dense @ v
err = jnp.linalg.norm(y_fft - y_dense)
print(f"FFT matvec vs dense : err = {err:.2e}")
# logdet still works (gaussx falls back to dense Cholesky for now;
# Levinson-style O(n^2) algorithms are forthcoming).
T_pd = Toeplitz(col.at[0].add(1e-3), tags=lx.positive_semidefinite_tag)
ld_struct = float(logdet(T_pd))
ld_dense = float(jnp.linalg.slogdet(T_pd.as_matrix())[1])
print(f"logdet (gaussx) : {ld_struct:.6f}")
print(f"logdet (dense ref) : {ld_dense:.6f}")FFT matvec vs dense : err = 3.91e-15
logdet (gaussx) : -143.237465
logdet (dense ref) : -143.237465
3. Cost table¶
| Operation | Dense | Toeplitz |
|---|---|---|
| Storage | ||
| Matvec | via FFT | |
| Solve (exact) | Cholesky | Levinson–Durbin (fallback in gaussx: dense) |
| Solve (iterative) | per CG iter | per CG iter |
| logdet (exact) | Cholesky | via Szegő-Levinson (fallback: dense) |
| logdet (asymptotic) | — | via FFT spectrum |
The current gaussx implementation gives you the matvec asymptotic for free. Exact Toeplitz solve via Levinson is on the roadmap; until then, the practical pattern is FFT-Toeplitz-matvec inside CG (or any iterative Krylov solver), which is the right pattern at large anyway because Levinson grows quadratically while CG with FFT matvec stays per iteration.
4. Asymptotic cost: dense vs Toeplitz¶
Theoretical FLOPs sweep, from 32 to 220. Note the point: dense matvec is 1012 flops (infeasible without distributed memory); Toeplitz matvec is flops (microseconds on a laptop). This is the gap that makes million-point regular-grid GP regression possible.
ns = np.array([2**k for k in range(5, 21)]) # 32 .. 1M
dense_mv = 2 * ns**2
toep_mv = 5 * ns * np.log2(ns) # ~3 FFTs of length 2n
dense_solve = (1/3) * ns**3
toep_solve_levinson = ns**2
fig, ax = plt.subplots(figsize=(7.5, 5.5))
ax.loglog(ns, dense_mv, "C3-", marker="o", label="dense matvec $O(n^2)$")
ax.loglog(ns, toep_mv, "C2-", marker="s", label="Toeplitz matvec $O(n\\log n)$")
ax.loglog(ns, dense_solve, "C3--", marker="o", label="dense Cholesky solve $O(n^3)$")
ax.loglog(ns, toep_solve_levinson, "C2--", marker="s", label="Toeplitz Levinson solve $O(n^2)$")
ax.axvspan(1e5, 2e6, color="grey", alpha=0.1)
ax.text(3e5, 1e3, "regime where\ndense is infeasible", ha="center", fontsize=9, color="0.3")
ax.set_xlabel(r"$n$ (grid points)")
ax.set_ylabel("FLOPs (theoretical)")
ax.set_title("Toeplitz vs dense — matvec and solve scaling")
ax.legend(loc="upper left")
plt.tight_layout()
plt.show()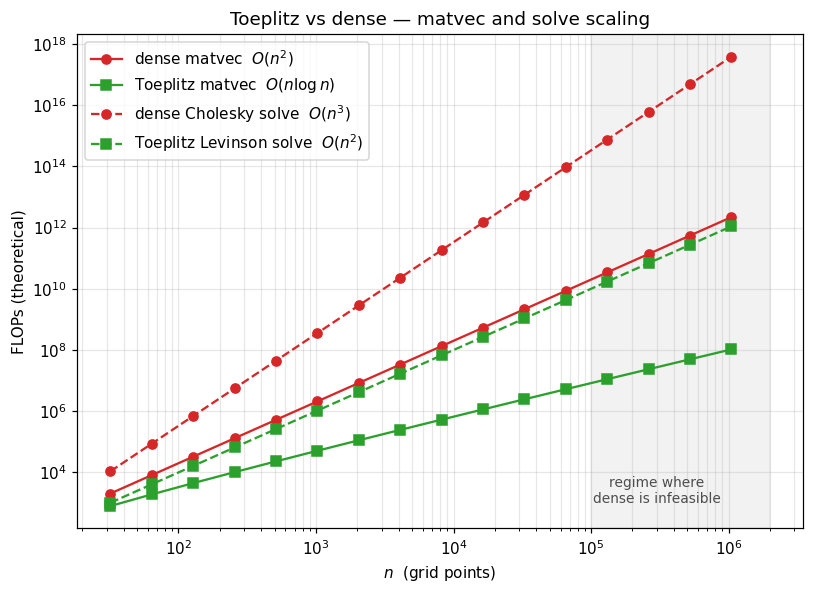
5. 2-D stationary kernels: Kronecker(Toeplitz, Toeplitz)¶
The composition is the headline geophysics application. A stationary kernel on a regular grid factorizes as
Storage drops from to . Matvec costs — i.e. — via the Kronecker vec-trick wrapping two FFT-Toeplitz matvecs.
This is the structure of essentially every 2-D gridded reanalysis covariance (SST, surface temperature, atmospheric pressure, soil moisture, gridded CO2 flux). When you see a paper claiming “we scale to 106 spatial points without approximation”, the operator is almost always Kronecker(Toeplitz_x, Toeplitz_y).
nx, ny = 32, 24
gx = jnp.linspace(0, 4, nx)
gy = jnp.linspace(0, 3, ny)
ell_x, ell_y = 0.5, 0.4
col_x = jnp.exp(-0.5 * (gx - gx[0])**2 / ell_x**2)
col_y = jnp.exp(-0.5 * (gy - gy[0])**2 / ell_y**2)
T_x = Toeplitz(col_x, tags=lx.positive_semidefinite_tag)
T_y = Toeplitz(col_y, tags=lx.positive_semidefinite_tag)
K_2d = Kronecker(T_x, T_y)
N = nx * ny
# Verify against the dense reference
K_dense = jnp.kron(T_x.as_matrix(), T_y.as_matrix())
v = jax.random.normal(jax.random.PRNGKey(2), (N,))
err = jnp.linalg.norm(K_2d.mv(v) - K_dense @ v)
print(f"grid : {nx} x {ny} = {N} points")
print(f"matvec error : {err:.2e}")
print(f"Kron-Toeplitz storage : {col_x.size + col_y.size} floats")
print(f"Dense storage : {N*N} floats ({N*N / (col_x.size + col_y.size):.0f}x more)")grid : 32 x 24 = 768 points
matvec error : 8.60e-14
Kron-Toeplitz storage : 56 floats
Dense storage : 589824 floats (10533x more)
# Visualize: a sample drawn from the 2-D stationary GP using the Kron-Toeplitz operator.
# Sample by drawing in eigenbasis: f = Q diag(sqrt(eigs)) z, where Q is Q_x ⊗ Q_y.
la, Qx = jnp.linalg.eigh(T_x.as_matrix())
mu, Qy = jnp.linalg.eigh(T_y.as_matrix())
eigs_2d = (la[:, None] * mu[None, :]) # outer product of factor spectra
z = jax.random.normal(jax.random.PRNGKey(3), (nx, ny))
f = Qx @ (jnp.sqrt(jnp.clip(eigs_2d, 0)) * (Qx.T @ z @ Qy)) @ Qy.T
fig, ax = plt.subplots(figsize=(7, 4.5))
im = ax.imshow(f, extent=[gy[0], gy[-1], gx[0], gx[-1]], origin="lower",
cmap="RdBu_r", aspect="auto")
ax.set_xlabel("$y$ (e.g. longitude)")
ax.set_ylabel("$x$ (e.g. latitude)")
ax.set_title("Sample from a 2-D stationary GP\n"
r"covariance = $K_x \otimes K_y$ with both factors Toeplitz")
plt.colorbar(im, ax=ax, label="field value")
plt.tight_layout()
plt.show()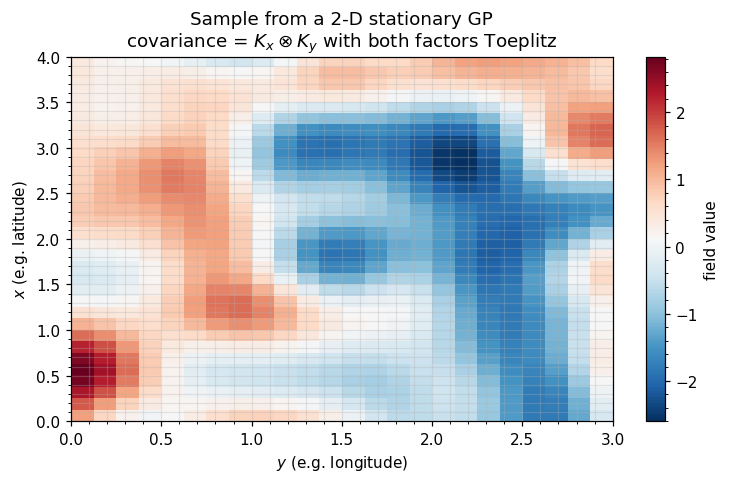
6. Caveats — when Toeplitz fails¶
Three sharp edges to watch for in real geoscience workflows:
Non-stationarity. If the kernel depends on absolute position rather than just lag , the matrix is not Toeplitz. Examples: input-dependent lengthscales (heteroscedastic kernels), spatially varying anisotropy, locally adaptive RBF. Workaround: locally stationary patches with a
BlockDiag(Toeplitz, Toeplitz, ...), or warp the input space before applying a stationary kernel.Irregular sampling. Toeplitz requires constant grid spacing. A tide-gauge series with missing observations is no longer Toeplitz — but it is a Toeplitz matrix on the full regular grid restricted to the observed indices, which is exactly what
MaskedOperator(notebook 1.6) handles. For genuinely irregular timestamps (event-driven data, asynchronous moorings), Toeplitz is gone and you fall back to dense or to inducing-point approximations.2-D irregular grids. The Kronecker × Toeplitz trick requires both axes to be regular. An ocean reanalysis on a regular lat/lon grid is fine; an unstructured triangular mesh from a finite-element ocean model (e.g. FESOM, MPAS) is not. For unstructured meshes use the GMRF / sparse-precision route instead (1.5 BlockTriDiag, future 4.C).
The next notebook (1.5 BlockTriDiag) covers the temporal-Markov alternative to Toeplitz: instead of an matvec for stationary covariances, you get an solve for sparse-precision (Markov) covariances. The two structures are dual and often combined.