KroneckerSum vs SumKronecker — two operators, one confusing name
Two of the most useful structured operators in gaussx have nearly identical names — and very different jobs. Sorting them out is worth a notebook of its own.
| Operator | Definition | Where it appears |
|---|---|---|
KroneckerSum | separable PDEs, spacetime GMRF priors, additive Gaussian fields | |
SumKronecker | multi-output GPs with correlated tasks, signal + iid noise on a grid |
Both produce matrices. Both look like “a sum of Kroneckers”. But the structure is wildly different: KroneckerSum is the additive separation of two operators acting on independent axes — eigendecomposable in closed form via the eigendecomposition of each factor alone. SumKronecker is the superposition of two unrelated Kronecker products — generally not jointly diagonalizable, and requiring an inner solve.
What you will see by the end:
- The vec-form identity that makes
KroneckerSummatvec cost . - The eigendecomposition (sum of spectra), and the resulting solve / logdet — same scaling as a single Kronecker.
- Why
SumKroneckerhas no analogous closed form, and the joint-eigenbasis trickgaussxuses to handle it. - A side-by-side cost table showing the gap: cheap factor-wise vs dense-after-a-rotation.
- The “which one do I want” decision rule.
By the end you should be able to spot which of the two is hiding inside any spacetime / multi-output covariance you encounter.
0. Where these structures show up in geoscience¶
These two operators are the workhorses of spacetime and multi-variable covariances — exactly the setting where geoscience problems live.
KroneckerSum: the additive separation. Used whenever a process is governed by independent action along two axes that add rather than multiply:- 2-D Laplacian for elliptic PDE inversion (gravity, electromagnetics): , the separable second-difference.
- Spacetime Gaussian Markov random fields (GMRFs): a precision encodes “AR(1) in time and CAR in space, independently”. Common in INLA-style spatial epidemiology, air-quality monitoring, climate-station analysis.
- Advection-diffusion priors where the diffusion operator splits across axes.
- Any separable covariance that is parameterized additively (e.g. squared-exponential length scales added in the precision form for stability).
SumKronecker: the superposition of two Kronecker layers. The shape behind every multi-output / multi-fidelity geoscience workflow:- Multi-output spacetime fields: — joint emulation of (temperature, salinity, velocity) on a gridded ocean reanalysis.
- Coregionalized geostatistics (LMC, ICM): a low-rank correlation matrix between outputs combined with a spatial kernel , plus iid noise. Land-cover, mineral-resource, and soil-property mapping all live here.
- Multi-fidelity emulation: low-fidelity covariance (cheap simulator) plus a discrepancy (correction term).
- Hierarchical Bayesian downscaling: a coarse-grid GP plus a fine-grid GP, each with its own Kronecker factorization.
The mental shortcut: KroneckerSum is for single-quantity processes that decouple across axes; SumKronecker is for multi-quantity / multi-fidelity processes superposed on the same grid. Spacetime climate emulators routinely contain both — a SumKronecker outer layer (signal + noise) wrapping a KroneckerSum inner layer (spacetime separable precision).
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import einx
import jax
import jax.numpy as jnp
import lineax as lx
import matplotlib.pyplot as plt
import numpy as np
from gaussx import (
Kronecker,
KroneckerSum,
SumKronecker,
logdet,
solve,
)
jax.config.update("jax_enable_x64", True)
KEY = jax.random.PRNGKey(0)
plt.rcParams.update({
"figure.dpi": 110,
"axes.grid": True,
"axes.grid.which": "both",
"xtick.minor.visible": True,
"ytick.minor.visible": True,
"grid.alpha": 0.3,
})
def psd_op(M):
return lx.MatrixLinearOperator(M, lx.positive_semidefinite_tag)
def random_psd(key, n, jitter=1e-1):
A = jax.random.normal(key, (n, n))
return A @ A.T + jitter * jnp.eye(n)1. KroneckerSum — additive separation¶
The Kronecker sum (note: not a sum of Kroneckers) is
Picture it as a 2-D tensor-product grid where one axis is governed by and the other by , and the two contributions add. The classic example is the 2-D Laplacian: a 1-D second-difference acting on plus a 1-D second-difference acting on becomes on the joint grid.
Because the action splits across axes, the matvec admits a vec-form analogous to Roth’s lemma for :
Two matrix multiplies of size — total cost , identical scaling to a Kronecker product matvec. (gaussx implements exactly this with einops.rearrange and jax.vmap.)
kA, kB = jax.random.split(jax.random.PRNGKey(1))
A_mat = random_psd(kA, 4)
B_mat = random_psd(kB, 3)
A_op = psd_op(A_mat)
B_op = psd_op(B_mat)
KS = KroneckerSum(A_op, B_op)
# Reference: build the dense Kronecker sum
KS_dense = jnp.kron(A_mat, jnp.eye(3)) + jnp.kron(jnp.eye(4), B_mat)
# matvec correctness
v = jax.random.normal(jax.random.PRNGKey(2), (12,))
err = jnp.linalg.norm(KS.mv(v) - KS_dense @ v)
print(f"matvec error : {err:.2e}")
print(f"shape : {KS.in_size()} x {KS.out_size()} (= {A_op.in_size()} * {B_op.in_size()})")matvec error : 2.26e-15
shape : 12 x 12 (= 4 * 3)
2. The eigendecomposition: sum-of-spectra¶
The headline result. If and are symmetric eigendecompositions, then has eigenvectors and eigenvalues :
The proof is one line: , and is diagonal with the entries .
Practically: solve and logdet need only the per-factor eigendecompositions (), not a joint pass:
The next cell verifies eq. ((3)) numerically.
# Per-factor eigendecompositions
lambda_A, Q_A = jnp.linalg.eigh(A_mat)
lambda_B, Q_B = jnp.linalg.eigh(B_mat)
# Predicted spectrum: outer-sum of factor spectra
predicted = (lambda_A[:, None] + lambda_B[None, :]).flatten()
predicted = jnp.sort(predicted)
# Direct eigendecomposition of the dense KroneckerSum
direct = jnp.linalg.eigvalsh(KS_dense)
direct = jnp.sort(direct)
print(f"max eigenvalue mismatch : {jnp.max(jnp.abs(predicted - direct)):.2e}")
print(f"first 6 sum-eigs : {predicted[:6]}")
print(f"first 6 direct eigs : {direct[:6]}")
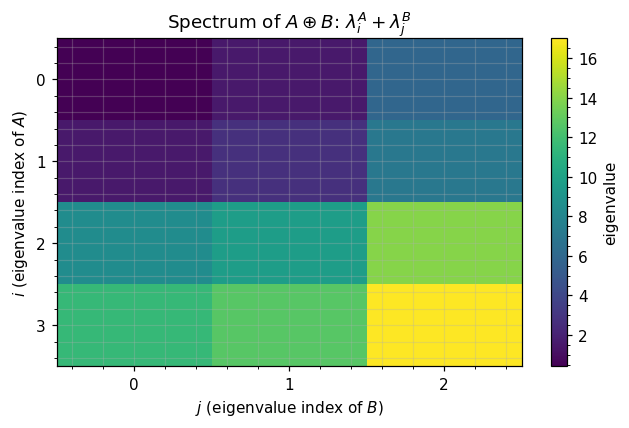
# Visualize spectrum as outer-sum heatmap
fig, ax = plt.subplots(figsize=(6, 4))
heat = lambda_A[:, None] + lambda_B[None, :]
im = ax.imshow(heat, cmap="viridis", aspect="auto")
ax.set_xticks(range(len(lambda_B)))
ax.set_yticks(range(len(lambda_A)))
ax.set_xlabel(r"$j$ (eigenvalue index of $B$)")
ax.set_ylabel(r"$i$ (eigenvalue index of $A$)")
ax.set_title(r"Spectrum of $A\oplus B$: $\lambda^A_i + \lambda^B_j$")
plt.colorbar(im, ax=ax, label="eigenvalue")
plt.tight_layout()
plt.show()max eigenvalue mismatch : 1.07e-14
first 6 sum-eigs : [0.42669084 1.56075562 1.58988744 2.72395222 5.91979167 7.05385645]
first 6 direct eigs : [0.42669084 1.56075562 1.58988744 2.72395222 5.91979167 7.05385645]
3. KroneckerSum solve in ¶
Per eq. ((4)), the solve is three steps:
- Project into the joint eigenbasis: .
- Element-wise divide by .
- Project back: .
Steps 1 and 3 are Kronecker-matvecs at . Step 2 is element-wise. The dominant cost is the two factor eigh calls.
This is the same scaling as a Kronecker product solve — additive separation costs no more than multiplicative.
# Manual solve through the eigenbasis (mirrors what a structured solver would do)
def kron_sum_solve(A_mat, B_mat, b):
la, QA = jnp.linalg.eigh(A_mat)
lb, QB = jnp.linalg.eigh(B_mat)
n_a, n_b = la.shape[0], lb.shape[0]
# rotate b into joint eigenbasis: vec(QB^T X QA), where X = mat(b)
X = einx.id("(a b) -> b a", b, a=n_a, b=n_b)
X_rot = QB.T @ X @ QA
# divide by sum of spectra
denom = la[None, :] + lb[:, None]
Y_rot = X_rot / denom
# rotate back
Y = QB @ Y_rot @ QA.T
return einx.id("b a -> (a b)", Y)
b = jax.random.normal(jax.random.PRNGKey(3), (12,))
x_struct = kron_sum_solve(A_mat, B_mat, b)
x_dense = jnp.linalg.solve(KS_dense, b)
print(f"solve error : {jnp.linalg.norm(x_struct - x_dense):.2e}")
# logdet via the spectrum (one log-sum, no Cholesky)
logdet_struct = jnp.sum(jnp.log(lambda_A[:, None] + lambda_B[None, :]))
logdet_dense = jnp.linalg.slogdet(KS_dense)[1]
print(f"logdet error: {abs(logdet_struct - logdet_dense):.2e}")solve error : 8.99e-15
logdet error: 7.11e-15
4. SumKronecker — superposition of two Kronecker products¶
A different beast. Now we have two Kronecker products and we add them:
The canonical case is multi-output GPs with correlated tasks: . Or signal-plus-noise on a tensor grid. Or any covariance where two distinct Kronecker layers superpose.
Matvec is the obvious sum of Kronecker matvecs — cost , twice that of a single Kronecker. Solve and logdet are the hard part: in general do not share eigenvectors, so we cannot simultaneously diagonalize and the spectrum is not a tidy sum or outer-product.
ks_keys = jax.random.split(jax.random.PRNGKey(10), 4)
A1 = psd_op(random_psd(ks_keys[0], 4))
B1 = psd_op(random_psd(ks_keys[1], 3))
A2 = psd_op(random_psd(ks_keys[2], 4))
B2 = psd_op(random_psd(ks_keys[3], 3))
SK = SumKronecker(Kronecker(A1, B1), Kronecker(A2, B2))
SK_dense = jnp.kron(A1.as_matrix(), B1.as_matrix()) + jnp.kron(A2.as_matrix(), B2.as_matrix())
v = jax.random.normal(jax.random.PRNGKey(11), (12,))
print(f"matvec error : {jnp.linalg.norm(SK.mv(v) - SK_dense @ v):.2e}")
# Are A1 and A2 jointly diagonalizable? Compute their commutator.
A1m, A2m = A1.as_matrix(), A2.as_matrix()
comm = A1m @ A2m - A2m @ A1m
print(f"||[A1, A2]|| : {jnp.linalg.norm(comm):.4f} (non-zero → no shared eigenbasis)")matvec error : 3.02e-14
||[A1, A2]|| : 35.6242 (non-zero → no shared eigenbasis)
5. The joint-eigenbasis-of-pair-2 trick¶
gaussx’s SumKronecker.eigendecompose() uses a hybrid approach:
- Eigendecompose the second pair: , . Cost: .
- Rotate the first pair into that basis: , . Cost: .
- Form the rotated full matrix — an matrix — and call
eigh. Cost: .
The third step is the bottleneck. Unlike KroneckerSum, we cannot avoid a dense pass on the full matrix. The trick only buys us:
- A symmetric form for
eigh(the rotation makes purely diagonal so the second pair contributes only to the diagonal). - Cheaper structural inheritance (the eigendecomposition is computed once and reused for solve, logdet, and matrix square root).
SumKronecker is intended for moderate factor sizes — typical multi-output GPs with task dimension . Past that, you need iterative Krylov solvers (1.D) or alternative parameterizations.
# Verify gaussx's SumKronecker.eigendecompose
evals, Q = SK.eigendecompose()
recon = Q @ jnp.diag(evals) @ Q.T
print(f"eigendecomp residual : {jnp.linalg.norm(recon - SK_dense):.2e}")
print(f"logdet via eigs : {float(jnp.sum(jnp.log(evals))):.6f}")
print(f"logdet ref : {float(jnp.linalg.slogdet(SK_dense)[1]):.6f}")
# Solve through the eigendecomposition
b = jax.random.normal(jax.random.PRNGKey(12), (12,))
x_eig = Q @ ((Q.T @ b) / evals)
x_ref = jnp.linalg.solve(SK_dense, b)
print(f"solve error via eigs : {jnp.linalg.norm(x_eig - x_ref):.2e}")eigendecomp residual : 1.07e-13
logdet via eigs : 27.528890
logdet ref : 27.528890
solve error via eigs : 1.97e-14
6. Cost & structure comparison¶
Side-by-side, with :
| Aspect | Kronecker | KroneckerSum | SumKronecker |
|---|---|---|---|
| Storage | |||
| Matvec | |||
| Spectrum | no closed form | ||
| Solve / logdet | via eigh on | ||
| Constructor | Kronecker(A, B) | KroneckerSum(A, B) | SumKronecker(Kronecker(A1,B1), Kronecker(A2,B2)) |
The two takeaways:
KroneckerSumis essentially free — same asymptotic cost as a single Kronecker product, including a clean closed-form spectrum.SumKroneckeris moderate-only — the cubic-in- inner eigh is the price of not having a shared eigenbasis. Use it when a few thousand (multi-output GP with small task dimension); reach for iterative methods otherwise.
7. Solve cost: KroneckerSum vs SumKronecker¶
Theoretical FLOP curves for the eigendecomposition-based solve, sweeping (square layout ). KroneckerSum scales as ; SumKronecker as . Plot is log–log so the slopes are exactly the asymptotic exponents.
ns = np.array([4, 8, 16, 32, 64])
Ns = ns**2
kron_sum_solve_flops = 2 * (1/3) * ns**3 # two eigh calls of size n
sum_kron_eig_flops = 2 * (1/3) * ns**3 + (1/3) * Ns**3 # two factor eighs + dense (n^2)^3 eigh
dense_solve_flops = (1/3) * Ns**3 # full Cholesky on N=n^2
fig, ax = plt.subplots(figsize=(7, 5))
ax.loglog(Ns, kron_sum_solve_flops, "C2-", marker="^", label="KroneckerSum solve")
ax.loglog(Ns, sum_kron_eig_flops, "C0--", marker="s", label="SumKronecker eigendecomp + solve")
ax.loglog(Ns, dense_solve_flops, "C3:", marker="o", label="dense Cholesky baseline")
ax.set_xlabel(r"$N = n_a n_b$ (square layout)")
ax.set_ylabel("FLOPs (theoretical)")
ax.set_title("Solve cost: separable vs superposed Kronecker structure")
ax.legend(loc="upper left")
plt.tight_layout()
plt.show()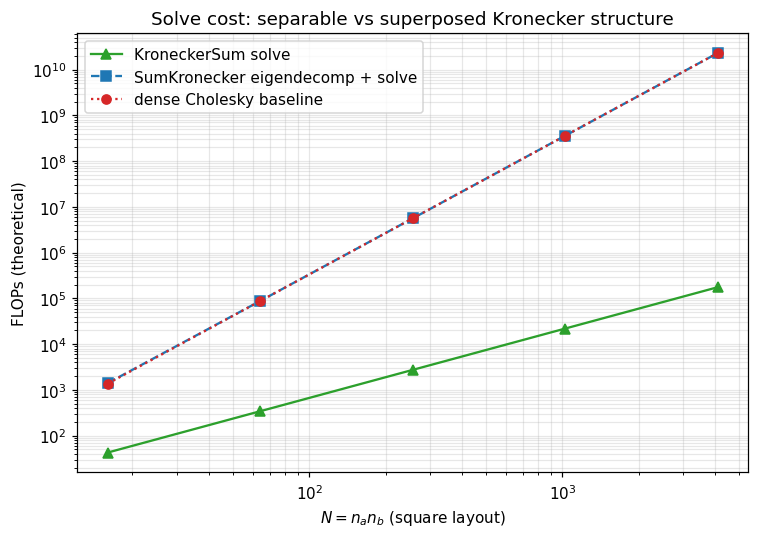
8. Which one do I want?¶
A simple decision rule when you build a covariance with Kronecker-shaped pieces:
| Pattern in your covariance | Reach for |
|---|---|
| “ acts on rows, acts on columns, contributions add” (separable PDE / GMRF) | KroneckerSum(A, B) |
| “Kernel on a tensor grid times a coregionalization matrix” | Kronecker(K_task, K_space) |
| “Kernel times correlation plus iid noise on a tensor grid” | SumKronecker(Kronecker(K_task, K_space), σ²·Kronecker(I, I)) |
| “Two independent Kronecker layers superposed” (multi-output, multi-fidelity) | SumKronecker(...) |
| “Three or more Kronecker layers” | iterative methods — see 1.E (SLQ logdet, BBMM) |
The naming is admittedly cruel — “Kronecker sum” means in linear-algebra texts, but a “sum of Kroneckers” reads like the same thing in plain English. In gaussx, the operator names track the linear-algebra convention: KroneckerSum = , SumKronecker = . When in doubt, check the eigendecomposition: if the spectrum is a tidy outer-sum of two factor spectra, you have a KroneckerSum. If not, you have a SumKronecker.
The rest of Part 1 fills in the remaining structured leaves (Toeplitz, BlockTriDiag, MaskedOperator) and then 1.7 explains how the kronecker_sum_tag attached on construction lets gaussx’s primitives dispatch through any of these forms automatically.