Woodbury Solve Step by Step
The Woodbury matrix identity turns an solve into a cheap solve when the matrix has low-rank structure:
where is the capacitance matrix.
This notebook walks through each step and shows that gaussx.solve
does it automatically.
Background¶
The Woodbury identity (also known as the matrix inversion lemma or Sherman-Morrison-Woodbury formula) is one of the most important identities in computational linear algebra. It appears in:
- Sparse GP inference — the Nystrom approximation yields a low-rank + diagonal covariance, and the Woodbury identity turns the inversion into a problem where .
- Kalman filtering — sequential rank-1 updates to the state covariance are applied via the matrix inversion lemma at each time step.
- Online learning — rank-1 covariance updates (e.g. recursive least squares) use the Sherman-Morrison special case.
- Ridge regression with feature space formulations — the kernel trick relates the and solutions through Woodbury.
The identity was first published by Woodbury (1950) and independently by Sherman & Morrison (1950) for the rank-1 case.
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import jax
import jax.numpy as jnp
import matplotlib.pyplot as plt
import gaussx
jax.config.update("jax_enable_x64", True)Setup¶
We construct where and (rank 3).
key = jax.random.PRNGKey(0)
k1, k2, k3 = jax.random.split(key, 3)
n, rank = 100, 3
d = jnp.abs(jax.random.normal(k1, (n,))) + 0.5 # positive diagonal
U = jax.random.normal(k2, (n, rank)) * 0.5
b = jax.random.normal(k3, (n,))
# Build operator
sigma = gaussx.low_rank_plus_diag(d, U)
print(f"Operator: diag({n}) + rank-{rank} update")
print(f" Full matrix size: {n}x{n} = {n**2:,} entries")
print(f" Low-rank storage: {n} + {n}x{rank} = {n + n * rank:,} entries")Operator: diag(100) + rank-3 update
Full matrix size: 100x100 = 10,000 entries
Low-rank storage: 100 + 100x3 = 400 entries
Step-by-step Woodbury¶
# Step 1: L^{-1} b (cheap — diagonal solve)
Linv_b = b / d
print("Step 1: L^{-1} b — diagonal solve, O(n)")
# Step 2: L^{-1} U (k diagonal solves)
Linv_U = U / d[:, None]
print(f"Step 2: L^{{-1}} U — {rank} diagonal solves, O(nk)")
# Step 3: Capacitance matrix C = I + U^T L^{-1} U (k x k)
C = jnp.eye(rank) + U.T @ Linv_U
print(f"Step 3: C = I + U^T L^{{-1}} U — {rank}x{rank} matrix")
# Step 4: Solve C z = U^T L^{-1} b (k x k dense solve)
z = jnp.linalg.solve(C, U.T @ Linv_b)
print(f"Step 4: C^{{-1}} (V^T L^{{-1}} b) — {rank}x{rank} solve")
# Step 5: x = L^{-1} b - L^{-1} U z
x_woodbury = Linv_b - Linv_U @ z
print("Step 5: x = L^{-1} b - L^{-1} U z — final answer")Step 1: L^{-1} b — diagonal solve, O(n)
Step 2: L^{-1} U — 3 diagonal solves, O(nk)
Step 3: C = I + U^T L^{-1} U — 3x3 matrix
Step 4: C^{-1} (V^T L^{-1} b) — 3x3 solve
Step 5: x = L^{-1} b - L^{-1} U z — final answer
Verify against gaussx.solve¶
x_gaussx = gaussx.solve(sigma, b)
x_dense = jnp.linalg.solve(sigma.as_matrix(), b)
print(f"Woodbury vs dense: max|diff| = {jnp.max(jnp.abs(x_woodbury - x_dense)):.2e}")
print(f"gaussx vs dense: max|diff| = {jnp.max(jnp.abs(x_gaussx - x_dense)):.2e}")
print(f"Woodbury vs gaussx: max|diff| = {jnp.max(jnp.abs(x_woodbury - x_gaussx)):.2e}")Woodbury vs dense: max|diff| = 5.55e-15
gaussx vs dense: max|diff| = 5.55e-15
Woodbury vs gaussx: max|diff| = 0.00e+00
Matrix determinant lemma¶
Similarly, the logdet decomposes. This follows from the identity
which is a direct consequence of the Schur complement determinant identity. Taking logarithms:
where is the capacitance matrix. In our case (, ):
ld_gaussx = gaussx.logdet(sigma)
ld_dense = jnp.linalg.slogdet(sigma.as_matrix())[1]
print(f"gaussx logdet: {ld_gaussx:.6f}")
print(f"dense logdet: {ld_dense:.6f}")
print(f"match: {jnp.allclose(ld_gaussx, ld_dense, rtol=1e-10)}")gaussx logdet: 27.039277
dense logdet: 27.039277
match: True
Visualizing the decomposition¶
fig, axes = plt.subplots(1, 4, figsize=(14, 3))
axes[0].imshow(jnp.diag(d[:20]), cmap="Blues", interpolation="nearest")
axes[0].set_title("L = diag(d)\n(first 20x20)")
axes[1].imshow(U[:20], cmap="RdBu", interpolation="nearest", aspect="auto")
axes[1].set_title(f"U\n({n}x{rank}, first 20 rows)")
axes[2].imshow(C, cmap="Blues", interpolation="nearest")
axes[2].set_title(f"C (capacitance)\n({rank}x{rank})")
mat_20 = jnp.abs(sigma.as_matrix()[:20, :20])
axes[3].imshow(mat_20, cmap="Blues", interpolation="nearest")
axes[3].set_title("A = L + UU^T\n(first 20x20)")
for ax in axes:
ax.set_xticks([])
ax.set_yticks([])
plt.tight_layout()
plt.show()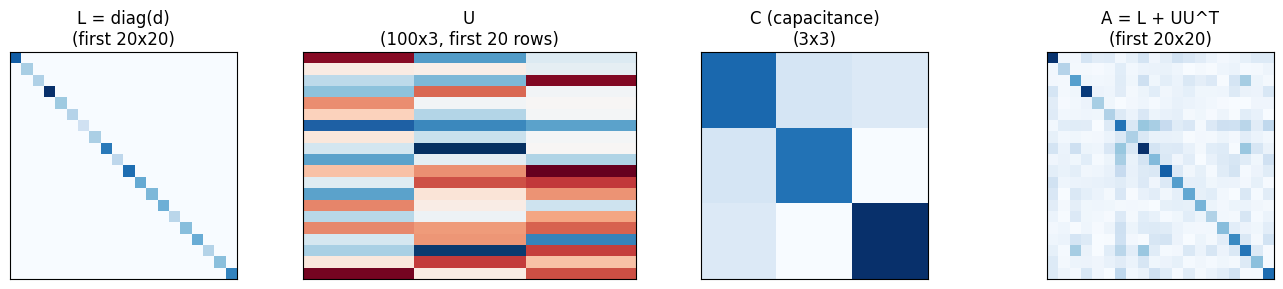
Cost comparison¶
| Operation | Dense | Woodbury |
|---|---|---|
| Solve | O(n^3) | O(nk^2 + k^3) |
| Logdet | O(n^3) | O(nk^2 + k^3) |
| Storage | O(n^2) | O(nk) |
For n=100, k=3: dense does ~10^6 ops, Woodbury does ~900.
References¶
- Hager, W. W. (1989). Updating the inverse of a matrix. SIAM Review, 31(2), 221--239.
- Henderson, H. V. & Searle, S. R. (1981). On deriving the inverse of a sum of matrices. SIAM Review, 23(1), 53--60.
- Woodbury, M. A. (1950). Inverting Modified Matrices. Statistical Research Group Memo Report 42, Princeton University.