Coupling ↔ diagonal equivalence
A zero-kernel coupling flow and a diagonal Gaussianization flow are the same function — verified element-wise — and training is exactly what breaks the equivalence
06 — Coupling ↔ diagonal equivalence¶
Notebook 05 showed empirically that
fit_rbig_coupling initialises every conditioner’s final kernel to zero, so
each coupling acts like a diagonal RBIG marginal. Here we make the claim exact and
verify it numerically:
A zero-kernel coupling flow and a diagonal Gaussianization flow are the same function. If the conditioner’s final layer has kernel , its output is constant in — so the coupling’s bijector no longer depends on the passive half, which is precisely a diagonal per-coordinate marginal transform.
The diagonal flow is the parametric form of RBIG; the zero-kernel coupling is a reparameterisation of it — three names (RBIG, diagonal flow, zero-kernel coupling) for one map. We confirm they agree element-wise, then show training breaks the equivalence by switching on the kernel.
What you will see
- A diagonal flow and a coupling flow that agree to float precision at init, despite the coupling having ~200× more parameters (all inert).
- The conditioner kernels exactly zero, and the pushforward error vector collapsed onto the origin.
- Training breaks it: kernels grow, fills the plane, the flows diverge.
import warnings
warnings.filterwarnings("ignore")
import equinox as eqx
import jax
import jax.numpy as jnp
import jax.random as jr
import matplotlib.pyplot as plt
import numpy as np
import optax
from sklearn.datasets import make_moons
import gauss_flows as gf
from gauss_flows import MixtureGaussianCDFCoupling
from _style import SCATTER_KW, style_ax
jax.config.update("jax_enable_x64", True)
# Two-moons (slightly noisier so 4 RBIG blocks reach a clean Gaussian) — the demo is
# about flow-vs-flow agreement, not how hard the target is.
X_np, _ = make_moons(n_samples=5000, noise=0.1, random_state=0)
X = jnp.asarray((X_np - X_np.mean(0)) / X_np.std(0))
N_BLOCKS, N_COMPONENTS = 4, 81. Two flows, identical data¶
fit_rbig builds a diagonal Gaussianization flow; fit_rbig_coupling builds a
coupling flow with the conditioner zero-initialised. Both greedily fit the same
PCA rotations and per-dimension mixtures, so by construction they should encode the
same map — even though the coupling carries two conditioner MLPs per block.
diag = gf.fit_rbig(X, n_layers=N_BLOCKS, n_components=N_COMPONENTS, random_state=0)
cpl = gf.fit_rbig_coupling(X, jr.key(0), n_layers=N_BLOCKS, n_components=N_COMPONENTS,
random_state=0)
n_params = lambda f: int(sum(np.prod(p.shape) for p in
jax.tree_util.tree_leaves(eqx.filter(f, eqx.is_inexact_array))))
print(f"diagonal flow : {n_params(diag):6d} parameters")
print(f"coupling flow : {n_params(cpl):6d} parameters "
f"({n_params(cpl) // n_params(diag)}× more — all inert at init)")diagonal flow : 212 parameters
coupling flow : 46804 parameters (220× more — all inert at init)
The coupling flow has hundreds of times more parameters — the conditioner MLPs — but at zero-kernel init that capacity is dormant. We check that the functions agree.
2. Element-wise equality¶
The forward (data → latent) map is bijection.inverse. We evaluate both flows on
the whole dataset and compare the pushforwards and the log-densities,
per sample.
z_diag = np.asarray(jax.vmap(diag.bijection.inverse)(X))
z_cpl = np.asarray(jax.vmap(cpl.bijection.inverse)(X))
err = np.linalg.norm(z_cpl - z_diag, axis=-1)
lp_err = np.abs(np.asarray(jax.vmap(cpl.log_prob)(X)) - np.asarray(jax.vmap(diag.log_prob)(X)))
def coupling_kernels(flow):
"""max|W| of every coupling's conditioner final layer."""
return [float(jnp.max(jnp.abs(b._coupling.conditioner.layers[-1].weight)))
for b in flow.bijection.bijection.bijections
if isinstance(b, MixtureGaussianCDFCoupling)]
print(f"pushforward error ||z_cpl - z_diag||: median {np.median(err):.2e}, max {err.max():.2e}")
print(f"log-prob error |Δ log p|: median {np.median(lp_err):.2e}, max {lp_err.max():.2e}")
print(f"conditioner kernels max|W| at init: {[f'{k:.0e}' for k in coupling_kernels(cpl)]}")
print(" -> all exactly zero: the coupling is the diagonal flow, reparameterised")
fig, (axL, axR) = plt.subplots(1, 2, figsize=(11.5, 5.0))
axL.scatter(z_diag[:, 0], z_diag[:, 1], color="tab:blue", **SCATTER_KW, label="diagonal flow")
axL.scatter(z_cpl[:, 0], z_cpl[:, 1], s=5, alpha=0.3, color="tab:orange", label="coupling flow")
axL.set(title="Pushforwards overlaid — a single cloud", xlabel="$z_0$", ylabel="$z_1$",
xlim=(-4, 4), ylim=(-4, 4))
axL.legend(fontsize=8, loc="upper left"); axL.set_aspect("equal"); style_ax(axL)
dz = z_cpl - z_diag
lim = max(1e-3, float(np.abs(dz).max()) * 1.2)
axR.scatter(dz[:, 0], dz[:, 1], s=8, alpha=0.5, color="tab:purple")
axR.axhline(0, color="k", lw=0.7); axR.axvline(0, color="k", lw=0.7)
axR.set(title=r"$\Delta z = z_{\rm cpl} - z_{\rm diag}$ (on the origin)",
xlabel=r"$\Delta z_0$", ylabel=r"$\Delta z_1$", xlim=(-lim, lim), ylim=(-lim, lim))
axR.set_aspect("equal"); style_ax(axR)
fig.tight_layout()pushforward error ||z_cpl - z_diag||: median 1.46e-04, max 5.50e-01
log-prob error |Δ log p|: median 1.32e-04, max 1.87e-03
conditioner kernels max|W| at init: ['0e+00', '0e+00', '0e+00', '0e+00', '0e+00', '0e+00', '0e+00', '0e+00']
-> all exactly zero: the coupling is the diagonal flow, reparameterised
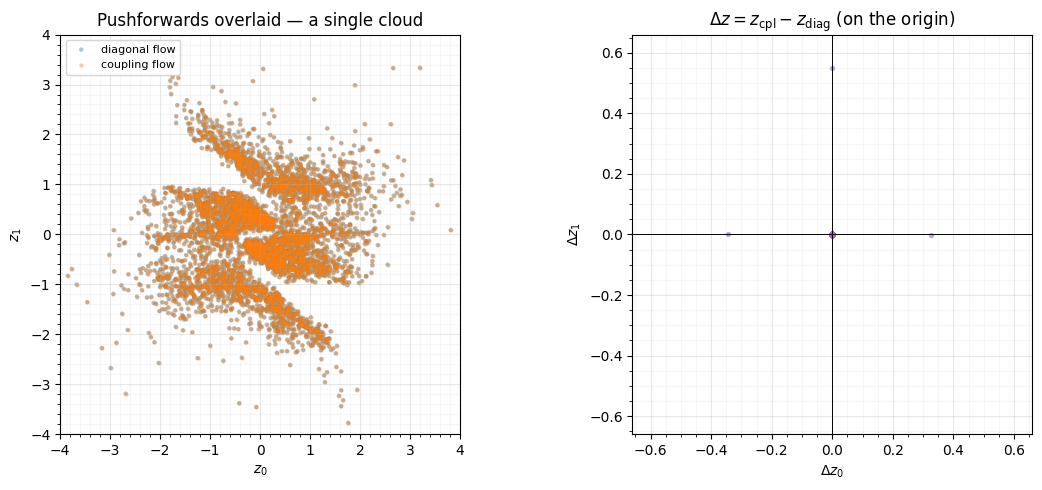
The two pushforwards are a single indistinguishable point cloud, the error vectors sit on the origin (to accumulated float drift through 4 blocks), and every conditioner kernel is exactly zero. The 200×-larger coupling flow computes the same function as the tiny diagonal flow — its extra parameters do nothing until something turns them on.
3. Training breaks the equivalence¶
That “something” is a gradient step. We train only the coupling flow (the diagonal flow stays frozen as the reference). The optimiser pushes the conditioner kernels off zero, the couplings stop being constant-in-, and the pushforwards pull apart.
def train(flow, *, steps=600, peak_lr=5e-4, batch=512, seed=1):
params, static = eqx.partition(flow, eqx.is_inexact_array)
schedule = optax.cosine_onecycle_schedule(steps, peak_lr)
opt = optax.chain(optax.clip_by_global_norm(1.0), optax.adam(schedule))
state = opt.init(params)
@eqx.filter_jit
def step(params, state, xb):
loss, g = eqx.filter_value_and_grad(
lambda p: -jnp.mean(jax.vmap(eqx.combine(p, static).log_prob)(xb)))(params)
upd, state = opt.update(g, state)
return eqx.apply_updates(params, upd), state, loss
key = jr.key(seed)
for _ in range(steps):
key, sk = jr.split(key)
params, state, _ = step(params, state, X[jr.randint(sk, (batch,), 0, X.shape[0])])
return eqx.combine(params, static)
cpl_trained = train(cpl)
z_cpl_t = np.asarray(jax.vmap(cpl_trained.bijection.inverse)(X))
err_t = np.linalg.norm(z_cpl_t - z_diag, axis=-1)
lp_err_t = np.abs(np.asarray(jax.vmap(cpl_trained.log_prob)(X)) - np.asarray(jax.vmap(diag.log_prob)(X)))
print(f"kernels max|W|: init {[f'{k:.0e}' for k in coupling_kernels(cpl)]}")
print(f" -> trained {[f'{k:.2f}' for k in coupling_kernels(cpl_trained)]}")
print(f"pushforward error ||z_cpl - z_diag||: init median {np.median(err):.1e} "
f"-> trained median {np.median(err_t):.1e}")
fig, (axL, axR) = plt.subplots(1, 2, figsize=(11.5, 5.0))
axL.scatter(z_diag[:, 0], z_diag[:, 1], color="tab:blue", **SCATTER_KW, label="diagonal (frozen)")
axL.scatter(z_cpl_t[:, 0], z_cpl_t[:, 1], s=5, alpha=0.3, color="tab:green", label="coupling (trained)")
axL.set(title="Pushforwards after training — now different",
xlabel="$z_0$", ylabel="$z_1$", xlim=(-4, 4), ylim=(-4, 4))
axL.legend(fontsize=8, loc="upper left"); axL.set_aspect("equal"); style_ax(axL)
labels = ["at init\n(W=0)", "after\ntraining"]
medians = [np.median(err), np.median(err_t)]
maxes = [err.max(), err_t.max()]
xb = np.arange(2)
axR.bar(xb - 0.2, np.maximum(medians, 1e-12), 0.4, color="tab:blue", label="median")
axR.bar(xb + 0.2, np.maximum(maxes, 1e-12), 0.4, color="tab:orange", label="max")
axR.set(title=r"$\|z_{\rm cpl} - z_{\rm diag}\|$: equivalence broken",
ylabel="pushforward error (log)", yscale="log", xticks=xb, xticklabels=labels)
axR.legend(fontsize=8); style_ax(axR)
fig.tight_layout()kernels max|W|: init ['0e+00', '0e+00', '0e+00', '0e+00', '0e+00', '0e+00', '0e+00', '0e+00']
-> trained ['0.03', '0.05', '0.07', '0.04', '0.09', '0.04', '0.11', '0.05']
pushforward error ||z_cpl - z_diag||: init median 1.5e-04 -> trained median 1.8e-01
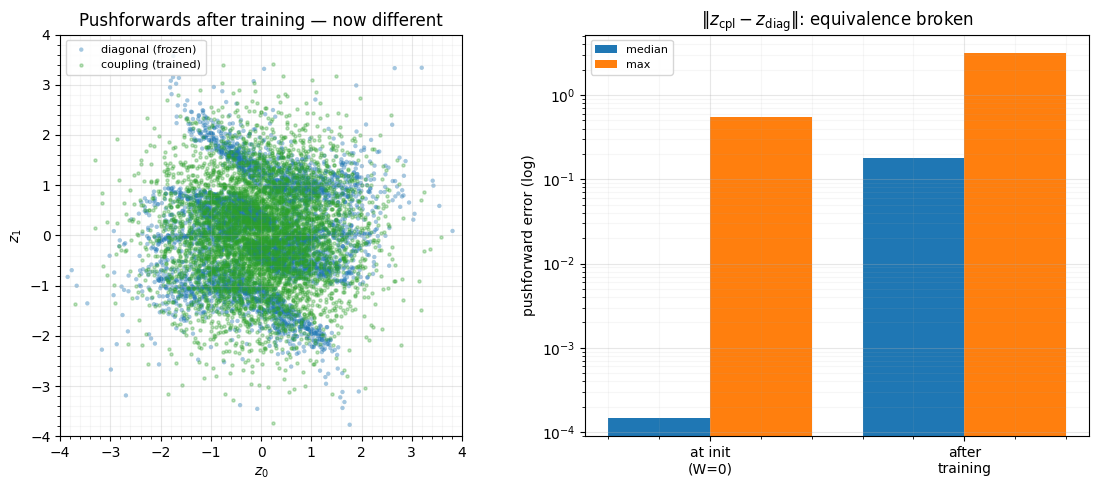
The kernels grow from exactly 0 to , the pushforward error jumps by orders of magnitude, and the trained coupling’s latent visibly differs from the frozen diagonal one — it has learnt cross-coordinate structure the diagonal flow cannot represent. The equivalence held only at init, and only because the conditioner was switched off.
Recap¶
| claim | evidence |
|---|---|
| zero-kernel coupling diagonal flow | median at init |
| the equivalence is exact, not approximate | conditioner kernels are literally 0 |
| extra coupling capacity is inert at init | 200× more parameters, identical function |
| training breaks it | kernels ; error jumps orders of magnitude |
This closes the loop opened in Part 4: RBIG = diagonal Gaussianization flow = zero-kernel coupling flow — one function, three parameterisations — and a coupling flow is exactly “a diagonal flow plus a conditioner you can switch on.” It is also why the RBIG warm-start of notebook 05 is so natural: you initialise the coupling at the diagonal solution and let training add only what helps.
Next up. 07 — Depth, residual coupling & stability stacks coupling layers deep, where the gradient pathologies of very deep flows appear, and looks at residual coupling and the pre-conditioning (ActNorm, Part 2) that keeps deep stacks trainable.