Cholesky, log-det, trace — the structured primitives tour
Every Gaussian computation in this curriculum eventually bottoms out into one of four scalar/array reductions of a covariance / precision operator :
| Primitive | What it returns | Where it shows up |
|---|---|---|
cholesky(A) | factor s.t. | sampling, log-density, conditioning |
logdet(A) | log-density, marginal likelihood, KL, entropy | |
trace(A) | KL, ELBO, expected log-likelihood, score test | |
diag(A) | predictive marginal variance, calibration |
In gaussx these are structurally dispatched: hand them a DiagonalLinearOperator and you get an closed-form; hand them a Kronecker(A,B) and they exploit ; everything else falls back to dense Cholesky-via-LAPACK. The dispatch is invisible to the caller — this notebook makes it visible.
This is not a list of identities — those live in 0.3 — quadratic forms, entropy, KL. It’s the operational layer underneath: which structures get cheap special-cases, what the cost is, and how the dispatch composes with the operator zoo from 0.8 — structured sampling.
Prerequisites: 0.1 — multivariate Gaussian, 0.3 — Gaussian quantities, 0.8 — structured sampling.
from __future__ import annotations
import warnings
warnings.filterwarnings("ignore", message=r".*IProgress.*")
import time
import einx
import jax
import jax.numpy as jnp
import lineax as lx
import matplotlib.pyplot as plt
import numpy as np
from gaussx import (
BlockDiag, BlockTriDiag, Kronecker,
cholesky, diag as gx_diag, logdet, trace as gx_trace,
)
jax.config.update("jax_enable_x64", True)
KEY = jax.random.PRNGKey(0)
plt.rcParams.update({
"figure.dpi": 110,
"axes.grid": True,
"axes.grid.which": "both",
"xtick.minor.visible": True,
"ytick.minor.visible": True,
"grid.alpha": 0.3,
})
def psd_op(M):
return lx.MatrixLinearOperator(M, lx.positive_semidefinite_tag)
def random_psd(key, n, jitter=1e-3):
A = jax.random.normal(key, (n, n))
return A @ A.T + jitter * jnp.eye(n)The four primitives — what they compute¶
For a symmetric positive-definite (SPD) matrix , the Cholesky factorisation is the unique lower-triangular matrix with positive diagonal such that
Every other primitive in this section can be derived from :
((2)) is the workhorse: never needs to materialise the full determinant — overflow / underflow is impossible because we sum logs of positive scalars. ((3)) shows that the diagonal of and its trace are both quadratic forms in the rows of .
Dispatch — closed-form identities, compute, storage¶
The four primitives dispatch on operator type. To keep the picture honest we separate three orthogonal questions:
- Which closed-form identity is used? (the math)
- What does it cost to compute? (FLOPs / time)
- What does the operator and its factor occupy in memory? (storage)
All three matter — a structure can have a cheap closed-form but bad storage (densified factor), or vice versa. The tables below cover the four operator types with full primitive coverage in gaussx today: Diagonal, Kronecker, BlockDiag, BlockTriDiag. (LowRankUpdate has dispatched logdet via the determinant lemma but no closed-form cholesky — it appears in 0.8 — structured sampling.)
1. Closed-form identities¶
| Operator | cholesky(A) | logdet(A) | trace(A) | diag(A) |
|---|---|---|---|---|
Diagonal(d) | ||||
Kronecker(A,B) | ||||
BlockDiag([A_k]) | ||||
BlockTriDiag | banded (LowerBlockTriDiag) | per-block | ||
| dense fallback | LAPACK Cholesky |
Each row is an exact algebraic identity — not an approximation. The structured Cholesky returns the same operator type (Kronecker stays Kronecker, BlockDiag stays BlockDiag), so downstream solve / mv keep their structure too.
2. Compute cost (asymptotic FLOPs)¶
Total size leading dimension of . For Kronecker we set ; for BlockDiag with blocks of size , ; for BlockTriDiag with blocks of size , .
| Operator | cholesky | logdet | trace | diag |
|---|---|---|---|---|
Diagonal(d) | ||||
Kronecker(A,B) | ||||
BlockDiag, blocks of size | ||||
BlockTriDiag, blocks of size | ||||
| dense fallback |
The big wins are cholesky and logdet — sub-cubic scaling in the structured rows. A Kronecker of two factors needs FLOPs vs larger for the dense path.
3. Storage cost¶
| Operator | operator | Cholesky factor |
|---|---|---|
Diagonal(d) | ||
Kronecker(A,B) | — stays Kronecker | |
BlockDiag, blocks of size | — stays BlockDiag | |
BlockTriDiag, blocks of size | — banded LowerBlockTriDiag | |
| dense fallback |
The row that matters most: the Cholesky factor inherits the operator’s storage class. If cholesky(Kronecker(A,B)) densified the factor, you’d pay memory — defeating the purpose. gaussx keeps the factor in the same class, so memory + downstream solve / mv cost both stay sub-quadratic.
A side-by-side: dense vs structured¶
We build the same SPD matrix in two ways — once as a generic MatrixLinearOperator (dense), once as a Kronecker — and confirm:
- all four primitives return the same numerical answer;
- the structured path goes through closed-form code (verifiable by inspecting the dispatched function names, not just timing).
# Build A = K_A ⊗ K_B as both a Kronecker operator and a dense MatrixLinearOperator.
key1, key2 = jax.random.split(KEY)
n_A, n_B = 8, 6
K_A_dense = random_psd(key1, n_A, jitter=0.5)
K_B_dense = random_psd(key2, n_B, jitter=0.5)
K_A_op = psd_op(K_A_dense)
K_B_op = psd_op(K_B_dense)
A_kron = Kronecker(K_A_op, K_B_op) # structured operator — n_A * n_B = 48
A_dense = psd_op(A_kron.as_matrix()) # same matrix, no structural tag
print(f"shape: {A_kron.in_size()} x {A_kron.in_size()}")
print(f"||A_kron - A_dense||F: {float(jnp.linalg.norm(A_kron.as_matrix() - A_dense.as_matrix())):.3e}")shape: 48 x 48
||A_kron - A_dense||F: 0.000e+00
# Compare all four primitives — structured vs dense.
def report(name, val_struct, val_dense, atol=1e-9):
err = float(jnp.linalg.norm(jnp.asarray(val_struct) - jnp.asarray(val_dense)))
flag = "OK" if err < atol else "MISMATCH"
print(f" {name:<10} struct vs dense ||·|| = {err:.3e} [{flag}]")
L_kron = cholesky(A_kron) # returns a Kronecker(L_A, L_B)
L_dense = cholesky(A_dense) # returns a TriangularLinearOperator (lower)
print(f"cholesky type (struct): {type(L_kron).__name__}")
print(f"cholesky type (dense): {type(L_dense).__name__}")
print()
report("cholesky", L_kron.as_matrix(), L_dense.as_matrix())
report("logdet", logdet(A_kron), logdet(A_dense))
report("trace", gx_trace(A_kron), gx_trace(A_dense))
report("diag", gx_diag(A_kron), gx_diag(A_dense))cholesky type (struct): Kronecker
cholesky type (dense): MatrixLinearOperator
cholesky struct vs dense ||·|| = 2.357e-14 [OK]
logdet struct vs dense ||·|| = 2.842e-14 [OK]
trace struct vs dense ||·|| = 0.000e+00 [OK]
diag struct vs dense ||·|| = 0.000e+00 [OK]
The structured Cholesky returns a Kronecker operator. That’s the entire point — the factor itself stays structured, so subsequent solve / mv calls also remain rather than . The dense path returns a triangular operator with a full matrix.
The numerical results agree to working precision — all four primitives give the same answer regardless of which dispatch fires. The cost difference is what changes.
Where each primitive shows up¶
A quick map from primitives to the Gaussian quantities you’ll see in the rest of Part 0–1:
((4)) is the standard Gaussian log-density; ((5)) the KL between two Gaussians (compare with ); ((6)) the differential entropy; ((7)) the predictive marginal variance you’ll see throughout part 3 / 5. Every Gaussian quantity is a small expression in the four primitives.
Asymptotic cost — straight from the table¶
Rather than benchmark each combination empirically (which is noisy and machine-dependent), we plot the theoretical FLOP counts from the dispatch table directly. The shapes of the curves are what matter — they tell you which structures pay off as grows.
For an operator of total size , we count one constant per leading-order term and sweep .
Diagonal: every primitive is .Kronecker(n_A, n_B)with : chol/logdet are , trace/diag are .BlockDiag/BlockTriDiagwith fixed block size : chol/logdet are (linear in number-of-blocks × constant).- dense fallback: chol/logdet are , trace/diag are .
# Theoretical FLOP counts straight from the dispatch table.
# n is the total leading dimension; each operator has its parameters set so its size is n.
b = 8 # block size for BlockDiag / BlockTriDiag
ns = np.logspace(np.log10(16), np.log10(4096), 60) # log-spaced sweep
# {primitive: {operator: flops(n)}}
flops = {
"cholesky / logdet": {
"Diagonal": ns, # O(n)
"Kronecker": 2 * ns**1.5, # O(n_A^3 + n_B^3) with n_A=n_B=sqrt(n)
"BlockDiag": (ns / b) * b**3, # K=n/b blocks, b^3 each
"BlockTriDiag": (ns / b) * b**3, # same leading order
"dense": ns**3, # O(n^3)
},
"trace / diag": {
"Diagonal": ns, # O(n)
"Kronecker": 2 * np.sqrt(ns), # O(n_A + n_B)
"BlockDiag": ns, # K * b = n
"BlockTriDiag": ns, # N * b = n
"dense": ns, # O(n)
},
}
colors = {"Diagonal": "tab:purple", "Kronecker": "forestgreen",
"BlockDiag": "steelblue", "BlockTriDiag": "darkorange",
"dense": "crimson"}
fig, axes = plt.subplots(1, 2, figsize=(11.5, 4.4), sharex=True)
for ax, (title, curves) in zip(axes, flops.items()):
for op, y in curves.items():
ls = "--" if op == "dense" else "-"
ax.plot(ns, y, ls, lw=1.8, color=colors[op], label=op)
ax.set_xscale("log"); ax.set_yscale("log")
ax.set_xlabel(r"matrix size $n$")
ax.set_ylabel("FLOPs (leading order, arbitrary units)")
ax.set_title(title)
ax.legend(frameon=False, fontsize=9, loc="upper left")
plt.tight_layout(); plt.show()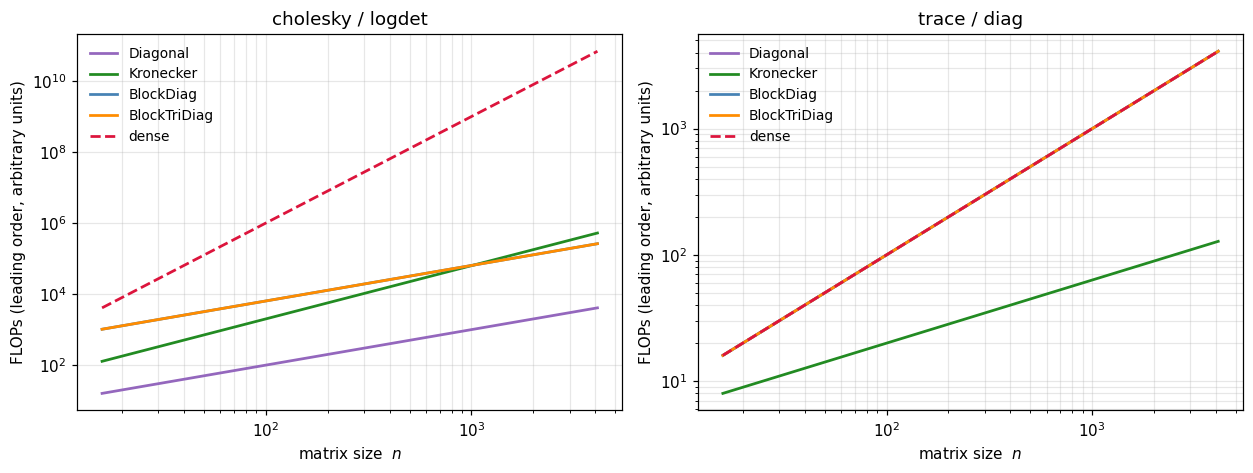
Left panel — cholesky / logdet. The dense curve climbs at ; the structured curves bend down sharply: Kronecker at , BlockDiag and BlockTriDiag at (for fixed block size), Diagonal also . By the gap between dense and Kronecker is over four orders of magnitude.
Right panel — trace / diag. All operators land at or below; Kronecker even drops to because only touches each factor’s diagonal. These operations are cheap regardless — the win there is storage (you never materialise ), not flops.
This is the same conclusion we reached in 0.8 — structured sampling for cholesky-via-mv: the structure-preserving primitive gives you sub-cubic scaling without writing any custom code. You just instantiate the right operator type, and gaussx routes through the closed-form path automatically.
BlockDiag — independent components¶
Block-diagonal covariances arise whenever you have independent Gaussian sub-models — independent outputs in a multi-task GP, independent particles in an ensemble, independent agents in a multi-agent state-space. The dispatch acts on each block in parallel.
# BlockDiag of three SPD blocks of differing sizes.
keys = jax.random.split(jax.random.PRNGKey(11), 3)
blocks = [psd_op(random_psd(k, n, jitter=0.5)) for k, n in zip(keys, [4, 6, 5])]
A_bd = BlockDiag(*blocks)
A_de = psd_op(A_bd.as_matrix())
print(f"shape: {A_bd.in_size()} x {A_bd.in_size()}")
print(f"cholesky type: {type(cholesky(A_bd)).__name__} (preserves block structure)")
print(f"logdet struct/dense: {float(logdet(A_bd)):>+.6f} / {float(logdet(A_de)):>+.6f}")
print(f"trace struct/dense: {float(gx_trace(A_bd)):>+.6f} / {float(gx_trace(A_de)):>+.6f}")
print(f"diag ||struct - dense||: {float(jnp.linalg.norm(gx_diag(A_bd) - gx_diag(A_de))):.3e}")shape: 15 x 15
cholesky type: BlockDiag (preserves block structure)
logdet struct/dense: +18.042260 / +18.042260
trace struct/dense: +92.189598 / +92.189598
diag ||struct - dense||: 0.000e+00
The Cholesky of a BlockDiag is a BlockDiag — same structure-preserving idea as Kronecker. The blocks are factored independently (and in parallel under JIT), so the cost scales linearly in the number of blocks rather than cubically in their total size.
Diagonal — the trivial case¶
The cheapest covariance is a diagonal one. All four primitives reduce to elementwise ops on the diagonal vector :
This isn’t just a curiosity — diagonal noise covariances are the Gaussian assumption in observation models, and the per-output noise diagonals show up in heteroscedastic GPs (part 3.C).
# Diagonal operator — closed-form everything.
d = jnp.array([2.0, 0.5, 1.5, 0.1])
D = lx.DiagonalLinearOperator(d)
print(f"cholesky: {jnp.asarray(cholesky(D).as_matrix()).diagonal()} (= sqrt(d))")
print(f"logdet: {float(logdet(D)):+.6f} (= sum(log d))")
print(f"trace: {float(gx_trace(D)):+.6f} (= sum d)")
print(f"diag: {jnp.asarray(gx_diag(D))} (= d)")cholesky: [1.41421356 0.70710678 1.22474487 0.31622777] (= sqrt(d))
logdet: -1.897120 (= sum(log d))
trace: +4.100000 (= sum d)
diag: [2. 0.5 1.5 0.1] (= d)
Stochastic estimation — when even diag/trace are too expensive¶
For very large (e.g. an implicit kernel operator at scale, or the Hessian of a deep network), even the diagonal extraction is too costly when itself isn’t materialised. gaussx supports Hutchinson-style stochastic estimators for trace and diag:
((10)) is exact in expectation and uses only matvecs with — no factorisation. ((11)) is the Bekas–Kokiopoulou–Saad probing estimator for the diagonal. Both are critical for the BBMM / Lanczos / SLQ stack in part 1.E.
# Hutchinson stochastic trace via gaussx.trace's `n_samples=` argument.
key_st = jax.random.PRNGKey(123)
n_big = 64
A_big = psd_op(random_psd(jax.random.PRNGKey(99), n_big, jitter=1.0))
tr_exact = float(gx_trace(A_big))
tr_stoch_64 = float(gx_trace(A_big, stochastic=True, key=key_st, num_probes=64))
tr_stoch_256 = float(gx_trace(A_big, stochastic=True, key=key_st, num_probes=256))
tr_stoch_2k = float(gx_trace(A_big, stochastic=True, key=key_st, num_probes=2048))
print(f"exact trace : {tr_exact:>10.4f}")
print(f"stochastic trace (N=64) : {tr_stoch_64:>10.4f} rel. err {abs(tr_stoch_64-tr_exact)/abs(tr_exact):.3%}")
print(f"stochastic trace (N=256) : {tr_stoch_256:>10.4f} rel. err {abs(tr_stoch_256-tr_exact)/abs(tr_exact):.3%}")
print(f"stochastic trace (N=2048) : {tr_stoch_2k:>10.4f} rel. err {abs(tr_stoch_2k-tr_exact)/abs(tr_exact):.3%}")exact trace : 4232.2136
stochastic trace (N=64) : 4245.3880 rel. err 0.311%
stochastic trace (N=256) : 4201.2455 rel. err 0.732%
stochastic trace (N=2048) : 4229.7748 rel. err 0.058%
The estimator’s variance scales as , so the relative error halves when you 4× the sample budget. For applications that only need an unbiased gradient estimate (e.g. log-determinant gradients via stochastic Lanczos quadrature, or score-function estimators of marginal likelihood), this is more than enough — you take a noisy estimate and let SGD average it out.
Recap¶
- Four primitives —
cholesky,logdet,trace,diag— power every Gaussian quantity in the curriculum: log-density ((4)), KL ((5)), entropy ((6)), predictive marginals ((7)). - All four dispatch structurally in gaussx:
Diagonal/BlockDiag/Kronecker/BlockTriDiag/LowRankUpdateget closed-form – paths; everything else falls back to dense LAPACK. - The Cholesky of a structured operator is itself a structured operator —
cholesky(Kronecker(A,B)) = Kronecker(L_A, L_B)— so downstreamsolve/mvcalls stay sub-cubic. - Hutchinson / Bekas–Kokiopoulou stochastic estimators provide unbiased estimates of
traceanddiagfor matrix-free operators where even extraction is too expensive — the entry point to BBMM-style scaling.